Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA.class.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | AHRR-overexpressed |
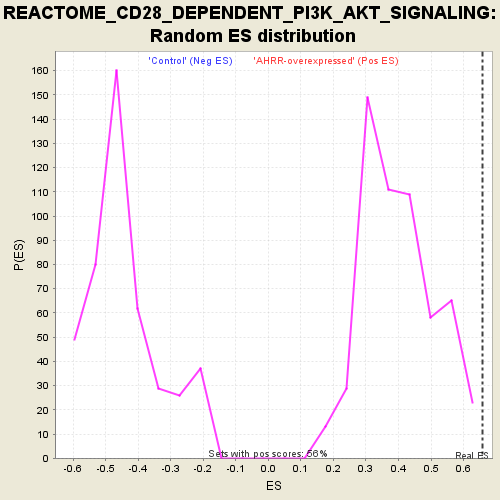
| GeneSet | REACTOME_CD28_DEPENDENT_PI3K_AKT_SIGNALING |
| Enrichment Score (ES) | 0.6586165 |
| Normalized Enrichment Score (NES) | 1.6336396 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.25563332 |
| FWER p-Value | 0.702 |

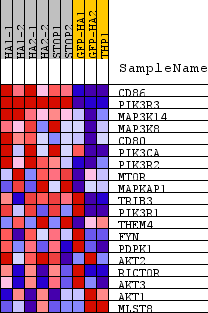
| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CD86 | na | 15 | 3.220 | 0.2894 | Yes | ||
| 2 | PIK3R3 | na | 163 | 2.009 | 0.4623 | Yes | ||
| 3 | MAP3K14 | na | 798 | 1.051 | 0.5214 | Yes | ||
| 4 | MAP3K8 | na | 1575 | 0.670 | 0.5383 | Yes | ||
| 5 | CD80 | na | 1737 | 0.623 | 0.5854 | Yes | ||
| 6 | PIK3CA | na | 2813 | 0.412 | 0.5623 | Yes | ||
| 7 | PIK3R2 | na | 2845 | 0.408 | 0.5973 | Yes | ||
| 8 | MTOR | na | 3631 | 0.308 | 0.5810 | Yes | ||
| 9 | MAPKAP1 | na | 3749 | 0.298 | 0.6013 | Yes | ||
| 10 | TRIB3 | na | 3841 | 0.287 | 0.6221 | Yes | ||
| 11 | PIK3R1 | na | 3998 | 0.271 | 0.6377 | Yes | ||
| 12 | THEM4 | na | 4053 | 0.265 | 0.6586 | Yes | ||
| 13 | FYN | na | 7370 | 0.046 | 0.4767 | No | ||
| 14 | PDPK1 | na | 7382 | 0.045 | 0.4801 | No | ||
| 15 | AKT2 | na | 7400 | 0.044 | 0.4832 | No | ||
| 16 | RICTOR | na | 10804 | -0.125 | 0.3035 | No | ||
| 17 | AKT3 | na | 11227 | -0.148 | 0.2932 | No | ||
| 18 | AKT1 | na | 13100 | -0.260 | 0.2115 | No | ||
| 19 | MLST8 | na | 16151 | -0.604 | 0.0948 | No |