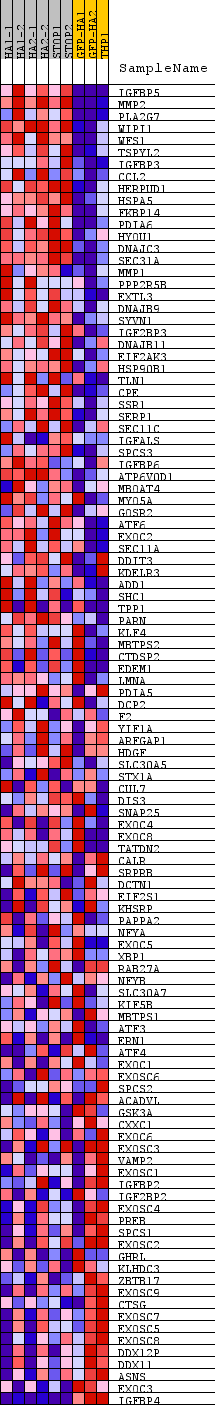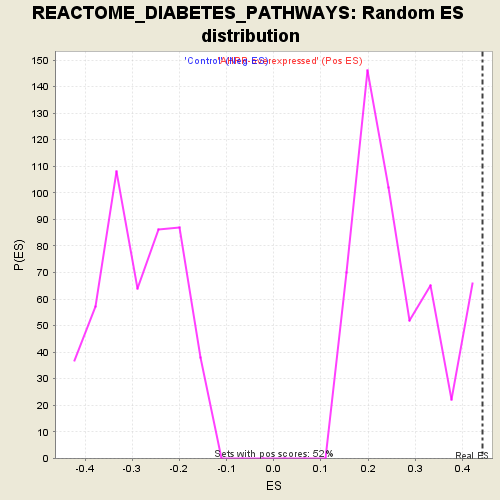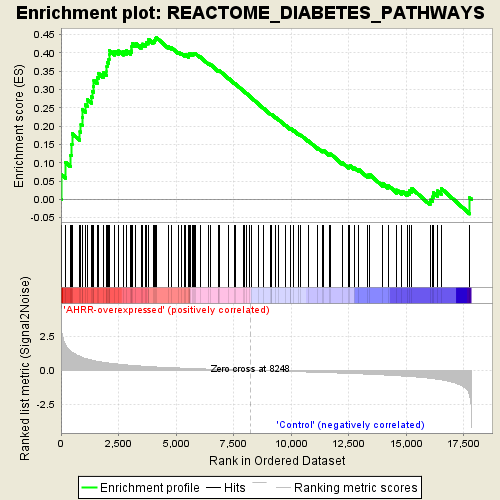
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA.class.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | AHRR-overexpressed |
| GeneSet | REACTOME_DIABETES_PATHWAYS |
| Enrichment Score (ES) | 0.44242266 |
| Normalized Enrichment Score (NES) | 1.680366 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.23661625 |
| FWER p-Value | 0.534 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | IGFBP5 | na | 25 | 3.002 | 0.0672 | Yes | ||
| 2 | MMP2 | na | 193 | 1.904 | 0.1013 | Yes | ||
| 3 | PLA2G7 | na | 422 | 1.420 | 0.1209 | Yes | ||
| 4 | WIPI1 | na | 459 | 1.371 | 0.1502 | Yes | ||
| 5 | WFS1 | na | 480 | 1.347 | 0.1798 | Yes | ||
| 6 | TSPYL2 | na | 820 | 1.036 | 0.1844 | Yes | ||
| 7 | IGFBP3 | na | 864 | 1.001 | 0.2048 | Yes | ||
| 8 | CCL2 | na | 922 | 0.973 | 0.2238 | Yes | ||
| 9 | HERPUD1 | na | 932 | 0.961 | 0.2453 | Yes | ||
| 10 | HSPA5 | na | 1060 | 0.881 | 0.2583 | Yes | ||
| 11 | FKBP14 | na | 1151 | 0.835 | 0.2723 | Yes | ||
| 12 | PDIA6 | na | 1319 | 0.768 | 0.2804 | Yes | ||
| 13 | HYOU1 | na | 1356 | 0.751 | 0.2955 | Yes | ||
| 14 | DNAJC3 | na | 1413 | 0.727 | 0.3090 | Yes | ||
| 15 | SEC31A | na | 1427 | 0.723 | 0.3248 | Yes | ||
| 16 | MMP1 | na | 1569 | 0.673 | 0.3322 | Yes | ||
| 17 | PPP2R5B | na | 1619 | 0.658 | 0.3445 | Yes | ||
| 18 | EXTL3 | na | 1835 | 0.596 | 0.3460 | Yes | ||
| 19 | DNAJB9 | na | 1989 | 0.562 | 0.3502 | Yes | ||
| 20 | SYVN1 | na | 1992 | 0.561 | 0.3629 | Yes | ||
| 21 | IGF2BP3 | na | 2004 | 0.558 | 0.3750 | Yes | ||
| 22 | DNAJB11 | na | 2076 | 0.544 | 0.3834 | Yes | ||
| 23 | EIF2AK3 | na | 2089 | 0.541 | 0.3951 | Yes | ||
| 24 | HSP90B1 | na | 2096 | 0.541 | 0.4071 | Yes | ||
| 25 | TLN1 | na | 2329 | 0.494 | 0.4053 | Yes | ||
| 26 | CPE | na | 2509 | 0.460 | 0.4058 | Yes | ||
| 27 | SSR1 | na | 2714 | 0.425 | 0.4040 | Yes | ||
| 28 | SERP1 | na | 2843 | 0.408 | 0.4061 | Yes | ||
| 29 | SEC11C | na | 3021 | 0.382 | 0.4048 | Yes | ||
| 30 | IGFALS | na | 3074 | 0.375 | 0.4105 | Yes | ||
| 31 | SPCS3 | na | 3076 | 0.375 | 0.4190 | Yes | ||
| 32 | IGFBP6 | na | 3099 | 0.373 | 0.4263 | Yes | ||
| 33 | ATP6V0D1 | na | 3256 | 0.352 | 0.4255 | Yes | ||
| 34 | MBOAT4 | na | 3480 | 0.325 | 0.4204 | Yes | ||
| 35 | MYO5A | na | 3535 | 0.320 | 0.4246 | Yes | ||
| 36 | GOSR2 | na | 3671 | 0.304 | 0.4240 | Yes | ||
| 37 | ATF6 | na | 3701 | 0.302 | 0.4292 | Yes | ||
| 38 | EXOC2 | na | 3785 | 0.293 | 0.4312 | Yes | ||
| 39 | SEC11A | na | 3794 | 0.292 | 0.4375 | Yes | ||
| 40 | DDIT3 | na | 3999 | 0.271 | 0.4321 | Yes | ||
| 41 | KDELR3 | na | 4065 | 0.264 | 0.4345 | Yes | ||
| 42 | ADD1 | na | 4090 | 0.261 | 0.4391 | Yes | ||
| 43 | SHC1 | na | 4136 | 0.256 | 0.4424 | Yes | ||
| 44 | TPP1 | na | 4665 | 0.213 | 0.4175 | No | ||
| 45 | PARN | na | 4791 | 0.204 | 0.4151 | No | ||
| 46 | KLF4 | na | 5097 | 0.182 | 0.4021 | No | ||
| 47 | MBTPS2 | na | 5225 | 0.174 | 0.3989 | No | ||
| 48 | CTDSP2 | na | 5359 | 0.164 | 0.3951 | No | ||
| 49 | EDEM1 | na | 5391 | 0.162 | 0.3971 | No | ||
| 50 | LMNA | na | 5550 | 0.151 | 0.3916 | No | ||
| 51 | PDIA5 | na | 5561 | 0.150 | 0.3945 | No | ||
| 52 | DCP2 | na | 5563 | 0.150 | 0.3978 | No | ||
| 53 | F2 | na | 5624 | 0.147 | 0.3978 | No | ||
| 54 | YIF1A | na | 5699 | 0.141 | 0.3969 | No | ||
| 55 | ARFGAP1 | na | 5743 | 0.138 | 0.3976 | No | ||
| 56 | HDGF | na | 5795 | 0.135 | 0.3978 | No | ||
| 57 | SLC30A5 | na | 5843 | 0.132 | 0.3982 | No | ||
| 58 | STX1A | na | 6048 | 0.119 | 0.3894 | No | ||
| 59 | CUL7 | na | 6418 | 0.097 | 0.3708 | No | ||
| 60 | DIS3 | na | 6509 | 0.092 | 0.3678 | No | ||
| 61 | SNAP25 | na | 6857 | 0.073 | 0.3499 | No | ||
| 62 | EXOC4 | na | 6864 | 0.073 | 0.3512 | No | ||
| 63 | EXOC8 | na | 6882 | 0.071 | 0.3519 | No | ||
| 64 | TATDN2 | na | 7292 | 0.050 | 0.3300 | No | ||
| 65 | CALR | na | 7531 | 0.038 | 0.3174 | No | ||
| 66 | SRPRB | na | 7573 | 0.035 | 0.3159 | No | ||
| 67 | DCTN1 | na | 7927 | 0.016 | 0.2964 | No | ||
| 68 | EIF2S1 | na | 7989 | 0.013 | 0.2932 | No | ||
| 69 | KHSRP | na | 8076 | 0.008 | 0.2886 | No | ||
| 70 | PAPPA2 | na | 8195 | 0.003 | 0.2820 | No | ||
| 71 | NFYA | na | 8285 | -0.001 | 0.2770 | No | ||
| 72 | EXOC5 | na | 8576 | -0.016 | 0.2610 | No | ||
| 73 | XBP1 | na | 8800 | -0.026 | 0.2490 | No | ||
| 74 | RAB27A | na | 9100 | -0.041 | 0.2331 | No | ||
| 75 | NFYB | na | 9143 | -0.043 | 0.2317 | No | ||
| 76 | SLC30A7 | na | 9146 | -0.043 | 0.2325 | No | ||
| 77 | KIF5B | na | 9332 | -0.052 | 0.2233 | No | ||
| 78 | MBTPS1 | na | 9449 | -0.057 | 0.2181 | No | ||
| 79 | ATF3 | na | 9747 | -0.071 | 0.2030 | No | ||
| 80 | ERN1 | na | 9953 | -0.082 | 0.1933 | No | ||
| 81 | ATF4 | na | 9968 | -0.083 | 0.1944 | No | ||
| 82 | EXOC1 | na | 10123 | -0.091 | 0.1878 | No | ||
| 83 | EXOSC6 | na | 10327 | -0.100 | 0.1786 | No | ||
| 84 | SPCS2 | na | 10418 | -0.106 | 0.1759 | No | ||
| 85 | ACADVL | na | 10752 | -0.122 | 0.1600 | No | ||
| 86 | GSK3A | na | 11157 | -0.144 | 0.1405 | No | ||
| 87 | CXXC1 | na | 11356 | -0.156 | 0.1329 | No | ||
| 88 | EXOC6 | na | 11403 | -0.159 | 0.1339 | No | ||
| 89 | EXOSC3 | na | 11659 | -0.174 | 0.1235 | No | ||
| 90 | VAMP2 | na | 11720 | -0.177 | 0.1242 | No | ||
| 91 | EXOSC1 | na | 12232 | -0.205 | 0.1000 | No | ||
| 92 | IGFBP2 | na | 12497 | -0.220 | 0.0902 | No | ||
| 93 | IGF2BP2 | na | 12559 | -0.223 | 0.0918 | No | ||
| 94 | EXOSC4 | na | 12762 | -0.237 | 0.0859 | No | ||
| 95 | PREB | na | 12924 | -0.248 | 0.0824 | No | ||
| 96 | SPCS1 | na | 13319 | -0.278 | 0.0666 | No | ||
| 97 | EXOSC2 | na | 13422 | -0.286 | 0.0673 | No | ||
| 98 | GHRL | na | 13990 | -0.333 | 0.0430 | No | ||
| 99 | KLHDC3 | na | 14213 | -0.353 | 0.0385 | No | ||
| 100 | ZBTB17 | na | 14601 | -0.390 | 0.0256 | No | ||
| 101 | EXOSC9 | na | 14820 | -0.414 | 0.0228 | No | ||
| 102 | CTSG | na | 15047 | -0.438 | 0.0201 | No | ||
| 103 | EXOSC7 | na | 15161 | -0.451 | 0.0240 | No | ||
| 104 | EXOSC5 | na | 15249 | -0.462 | 0.0296 | No | ||
| 105 | EXOSC8 | na | 16043 | -0.587 | -0.0017 | No | ||
| 106 | DDX12P | na | 16142 | -0.603 | 0.0066 | No | ||
| 107 | DDX11 | na | 16189 | -0.611 | 0.0179 | No | ||
| 108 | ASNS | na | 16364 | -0.653 | 0.0230 | No | ||
| 109 | EXOC3 | na | 16527 | -0.697 | 0.0298 | No | ||
| 110 | IGFBP4 | na | 17768 | -1.935 | 0.0041 | No |