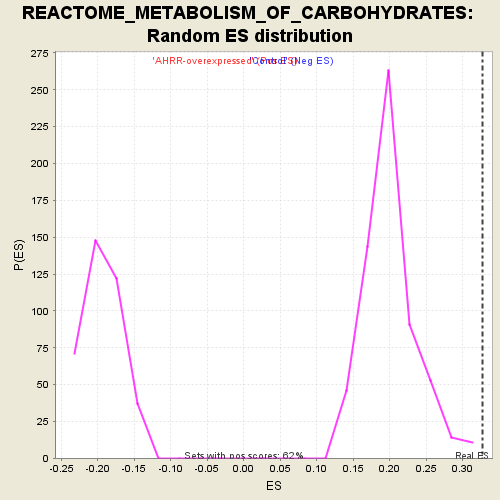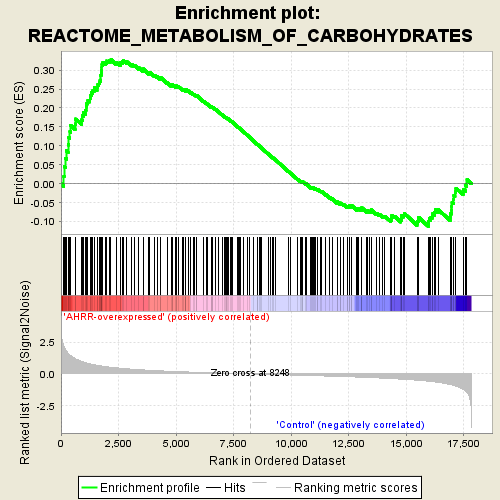
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA.class.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | AHRR-overexpressed |
| GeneSet | REACTOME_METABOLISM_OF_CARBOHYDRATES |
| Enrichment Score (ES) | 0.3275722 |
| Normalized Enrichment Score (NES) | 1.6495005 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.25246564 |
| FWER p-Value | 0.641 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SGSH | na | 121 | 2.179 | 0.0205 | Yes | ||
| 2 | MGAM | na | 135 | 2.102 | 0.0461 | Yes | ||
| 3 | SDC2 | na | 184 | 1.950 | 0.0679 | Yes | ||
| 4 | VCAN | na | 239 | 1.778 | 0.0871 | Yes | ||
| 5 | DSE | na | 308 | 1.619 | 0.1036 | Yes | ||
| 6 | NDST3 | na | 335 | 1.567 | 0.1218 | Yes | ||
| 7 | HEXB | na | 385 | 1.478 | 0.1375 | Yes | ||
| 8 | HK1 | na | 418 | 1.423 | 0.1536 | Yes | ||
| 9 | HS3ST3A1 | na | 610 | 1.209 | 0.1579 | Yes | ||
| 10 | PRKACA | na | 639 | 1.170 | 0.1710 | Yes | ||
| 11 | HS6ST3 | na | 903 | 0.983 | 0.1684 | Yes | ||
| 12 | GLB1 | na | 926 | 0.970 | 0.1794 | Yes | ||
| 13 | PGM1 | na | 971 | 0.930 | 0.1886 | Yes | ||
| 14 | PGAM2 | na | 1063 | 0.879 | 0.1944 | Yes | ||
| 15 | B4GALT4 | na | 1086 | 0.870 | 0.2041 | Yes | ||
| 16 | GYS2 | na | 1118 | 0.855 | 0.2131 | Yes | ||
| 17 | GPC5 | na | 1166 | 0.827 | 0.2208 | Yes | ||
| 18 | PFKFB3 | na | 1257 | 0.792 | 0.2256 | Yes | ||
| 19 | HEXA | na | 1289 | 0.781 | 0.2337 | Yes | ||
| 20 | NDST1 | na | 1325 | 0.766 | 0.2413 | Yes | ||
| 21 | B4GALT6 | na | 1383 | 0.741 | 0.2474 | Yes | ||
| 22 | PFKM | na | 1437 | 0.720 | 0.2534 | Yes | ||
| 23 | EXT1 | na | 1564 | 0.673 | 0.2547 | Yes | ||
| 24 | CHPF2 | na | 1600 | 0.663 | 0.2610 | Yes | ||
| 25 | GXYLT2 | na | 1648 | 0.648 | 0.2665 | Yes | ||
| 26 | HAS3 | na | 1690 | 0.635 | 0.2721 | Yes | ||
| 27 | HAS1 | na | 1720 | 0.627 | 0.2784 | Yes | ||
| 28 | PYGM | na | 1721 | 0.627 | 0.2862 | Yes | ||
| 29 | SDC3 | na | 1741 | 0.623 | 0.2930 | Yes | ||
| 30 | HS6ST1 | na | 1757 | 0.619 | 0.2999 | Yes | ||
| 31 | GOT1 | na | 1773 | 0.614 | 0.3067 | Yes | ||
| 32 | CHST15 | na | 1777 | 0.614 | 0.3143 | Yes | ||
| 33 | G6PD | na | 1816 | 0.602 | 0.3197 | Yes | ||
| 34 | B3GNT7 | na | 1928 | 0.576 | 0.3206 | Yes | ||
| 35 | HK2 | na | 1993 | 0.561 | 0.3240 | Yes | ||
| 36 | ST3GAL2 | na | 2084 | 0.543 | 0.3257 | Yes | ||
| 37 | PFKFB1 | na | 2169 | 0.526 | 0.3276 | Yes | ||
| 38 | CALM1 | na | 2418 | 0.478 | 0.3195 | No | ||
| 39 | AGL | na | 2573 | 0.447 | 0.3164 | No | ||
| 40 | GYG1 | na | 2591 | 0.445 | 0.3210 | No | ||
| 41 | CHSY3 | na | 2673 | 0.433 | 0.3219 | No | ||
| 42 | GPI | na | 2713 | 0.425 | 0.3250 | No | ||
| 43 | SLC25A12 | na | 2856 | 0.406 | 0.3220 | No | ||
| 44 | PGD | na | 3058 | 0.378 | 0.3154 | No | ||
| 45 | CHPF | na | 3183 | 0.362 | 0.3129 | No | ||
| 46 | SLC2A4 | na | 3380 | 0.336 | 0.3060 | No | ||
| 47 | HK3 | na | 3567 | 0.315 | 0.2994 | No | ||
| 48 | GOT2 | na | 3571 | 0.315 | 0.3032 | No | ||
| 49 | GAPDH | na | 3821 | 0.289 | 0.2927 | No | ||
| 50 | HS3ST3B1 | na | 3854 | 0.285 | 0.2945 | No | ||
| 51 | SLC2A1 | na | 4080 | 0.263 | 0.2850 | No | ||
| 52 | TKT | na | 4180 | 0.252 | 0.2826 | No | ||
| 53 | RPE | na | 4301 | 0.241 | 0.2788 | No | ||
| 54 | B4GALT1 | na | 4312 | 0.241 | 0.2813 | No | ||
| 55 | ARSB | na | 4611 | 0.218 | 0.2671 | No | ||
| 56 | CALM2 | na | 4797 | 0.203 | 0.2592 | No | ||
| 57 | HS2ST1 | na | 4834 | 0.201 | 0.2597 | No | ||
| 58 | ST3GAL6 | na | 4838 | 0.200 | 0.2620 | No | ||
| 59 | PRPS2 | na | 4993 | 0.189 | 0.2557 | No | ||
| 60 | HYAL1 | na | 5009 | 0.188 | 0.2572 | No | ||
| 61 | PPP2R5D | na | 5030 | 0.186 | 0.2584 | No | ||
| 62 | RANBP2 | na | 5112 | 0.181 | 0.2560 | No | ||
| 63 | B4GALT3 | na | 5296 | 0.169 | 0.2478 | No | ||
| 64 | MDH2 | na | 5302 | 0.168 | 0.2496 | No | ||
| 65 | GYG2 | na | 5401 | 0.162 | 0.2461 | No | ||
| 66 | EXT2 | na | 5403 | 0.161 | 0.2481 | No | ||
| 67 | PPP2R1A | na | 5405 | 0.161 | 0.2500 | No | ||
| 68 | PGK1 | na | 5525 | 0.153 | 0.2452 | No | ||
| 69 | NUP214 | na | 5634 | 0.146 | 0.2409 | No | ||
| 70 | CHST3 | na | 5739 | 0.139 | 0.2368 | No | ||
| 71 | ALDOC | na | 5801 | 0.135 | 0.2350 | No | ||
| 72 | ENO1 | na | 5877 | 0.130 | 0.2324 | No | ||
| 73 | GNS | na | 5903 | 0.128 | 0.2326 | No | ||
| 74 | ABCC5 | na | 6180 | 0.111 | 0.2183 | No | ||
| 75 | SLC9A1 | na | 6317 | 0.103 | 0.2119 | No | ||
| 76 | PFKL | na | 6362 | 0.100 | 0.2107 | No | ||
| 77 | B4GALT5 | na | 6523 | 0.091 | 0.2028 | No | ||
| 78 | ALDOA | na | 6537 | 0.090 | 0.2032 | No | ||
| 79 | AMY2B | na | 6577 | 0.087 | 0.2020 | No | ||
| 80 | PYGB | na | 6602 | 0.086 | 0.2018 | No | ||
| 81 | TPI1 | na | 6701 | 0.082 | 0.1972 | No | ||
| 82 | CHST2 | na | 6828 | 0.074 | 0.1910 | No | ||
| 83 | PCK2 | na | 7004 | 0.064 | 0.1819 | No | ||
| 84 | PPP2CB | na | 7109 | 0.058 | 0.1768 | No | ||
| 85 | PC | na | 7156 | 0.056 | 0.1749 | No | ||
| 86 | NUP50 | na | 7205 | 0.054 | 0.1728 | No | ||
| 87 | IDUA | na | 7211 | 0.054 | 0.1732 | No | ||
| 88 | ST3GAL1 | na | 7254 | 0.052 | 0.1715 | No | ||
| 89 | POM121 | na | 7259 | 0.051 | 0.1719 | No | ||
| 90 | PPP2R1B | na | 7381 | 0.045 | 0.1656 | No | ||
| 91 | NAGLU | na | 7406 | 0.044 | 0.1648 | No | ||
| 92 | NUPL2 | na | 7446 | 0.042 | 0.1631 | No | ||
| 93 | NDST2 | na | 7686 | 0.030 | 0.1499 | No | ||
| 94 | PPP2CA | na | 7712 | 0.028 | 0.1489 | No | ||
| 95 | B4GALT7 | na | 7756 | 0.025 | 0.1467 | No | ||
| 96 | GBE1 | na | 7816 | 0.022 | 0.1437 | No | ||
| 97 | PRKACB | na | 7911 | 0.017 | 0.1386 | No | ||
| 98 | NUP133 | na | 8111 | 0.006 | 0.1274 | No | ||
| 99 | PHKA2 | na | 8202 | 0.002 | 0.1223 | No | ||
| 100 | PFKFB4 | na | 8354 | -0.005 | 0.1138 | No | ||
| 101 | CHST11 | na | 8538 | -0.014 | 0.1036 | No | ||
| 102 | B3GAT3 | na | 8539 | -0.014 | 0.1038 | No | ||
| 103 | GXYLT1 | na | 8605 | -0.018 | 0.1003 | No | ||
| 104 | PHKA1 | na | 8669 | -0.020 | 0.0970 | No | ||
| 105 | UGP2 | na | 8693 | -0.021 | 0.0960 | No | ||
| 106 | PHKB | na | 9001 | -0.036 | 0.0790 | No | ||
| 107 | NUP153 | na | 9115 | -0.042 | 0.0731 | No | ||
| 108 | SLC25A1 | na | 9209 | -0.046 | 0.0684 | No | ||
| 109 | CHST6 | na | 9254 | -0.048 | 0.0665 | No | ||
| 110 | RAE1 | na | 9310 | -0.051 | 0.0641 | No | ||
| 111 | GYS1 | na | 9883 | -0.079 | 0.0326 | No | ||
| 112 | NUP188 | na | 9979 | -0.083 | 0.0283 | No | ||
| 113 | HS6ST2 | na | 10299 | -0.099 | 0.0115 | No | ||
| 114 | GLCE | na | 10398 | -0.105 | 0.0072 | No | ||
| 115 | PYGL | na | 10441 | -0.107 | 0.0062 | No | ||
| 116 | NUP210 | na | 10480 | -0.108 | 0.0054 | No | ||
| 117 | NUP93 | na | 10496 | -0.109 | 0.0059 | No | ||
| 118 | GAPDHS | na | 10628 | -0.116 | -0.0000 | No | ||
| 119 | B3GALT6 | na | 10655 | -0.117 | -0.0001 | No | ||
| 120 | TPR | na | 10862 | -0.129 | -0.0101 | No | ||
| 121 | G6PC3 | na | 10900 | -0.131 | -0.0106 | No | ||
| 122 | ALDOB | na | 10925 | -0.132 | -0.0103 | No | ||
| 123 | SLC2A5 | na | 10956 | -0.133 | -0.0103 | No | ||
| 124 | GUSB | na | 11036 | -0.137 | -0.0131 | No | ||
| 125 | B4GALT2 | na | 11045 | -0.138 | -0.0118 | No | ||
| 126 | PFKFB2 | na | 11135 | -0.142 | -0.0150 | No | ||
| 127 | PGAM1 | na | 11159 | -0.144 | -0.0145 | No | ||
| 128 | NUP54 | na | 11283 | -0.152 | -0.0196 | No | ||
| 129 | NUP205 | na | 11325 | -0.154 | -0.0200 | No | ||
| 130 | SLC25A13 | na | 11510 | -0.165 | -0.0283 | No | ||
| 131 | NUP85 | na | 11680 | -0.175 | -0.0357 | No | ||
| 132 | CHST13 | na | 11786 | -0.180 | -0.0394 | No | ||
| 133 | HS3ST5 | na | 12011 | -0.192 | -0.0497 | No | ||
| 134 | RPIA | na | 12013 | -0.192 | -0.0473 | No | ||
| 135 | CALM3 | na | 12159 | -0.201 | -0.0530 | No | ||
| 136 | PRPS1 | na | 12162 | -0.201 | -0.0506 | No | ||
| 137 | NUP88 | na | 12287 | -0.209 | -0.0550 | No | ||
| 138 | CHST14 | na | 12440 | -0.217 | -0.0609 | No | ||
| 139 | OGN | na | 12467 | -0.218 | -0.0597 | No | ||
| 140 | GALT | na | 12518 | -0.221 | -0.0597 | No | ||
| 141 | NUP155 | na | 12540 | -0.222 | -0.0581 | No | ||
| 142 | AAAS | na | 12605 | -0.226 | -0.0589 | No | ||
| 143 | PGM2 | na | 12640 | -0.228 | -0.0580 | No | ||
| 144 | HMMR | na | 12853 | -0.243 | -0.0669 | No | ||
| 145 | GALE | na | 12899 | -0.246 | -0.0664 | No | ||
| 146 | CHSY1 | na | 12923 | -0.247 | -0.0646 | No | ||
| 147 | PGLS | na | 13049 | -0.256 | -0.0685 | No | ||
| 148 | NUP37 | na | 13056 | -0.256 | -0.0656 | No | ||
| 149 | SDC4 | na | 13077 | -0.258 | -0.0635 | No | ||
| 150 | MDH1 | na | 13285 | -0.274 | -0.0718 | No | ||
| 151 | PHKG2 | na | 13303 | -0.276 | -0.0693 | No | ||
| 152 | SEH1L | na | 13425 | -0.286 | -0.0725 | No | ||
| 153 | SLC2A3 | na | 13484 | -0.291 | -0.0722 | No | ||
| 154 | TALDO1 | na | 13487 | -0.291 | -0.0686 | No | ||
| 155 | NUP35 | na | 13716 | -0.310 | -0.0777 | No | ||
| 156 | B3GNT4 | na | 13833 | -0.320 | -0.0802 | No | ||
| 157 | NUP62 | na | 13993 | -0.334 | -0.0850 | No | ||
| 158 | IDS | na | 14080 | -0.340 | -0.0856 | No | ||
| 159 | UST | na | 14334 | -0.364 | -0.0954 | No | ||
| 160 | BCAN | na | 14356 | -0.366 | -0.0920 | No | ||
| 161 | FBP1 | na | 14369 | -0.367 | -0.0881 | No | ||
| 162 | ENO2 | na | 14379 | -0.368 | -0.0840 | No | ||
| 163 | SLC25A11 | na | 14488 | -0.380 | -0.0853 | No | ||
| 164 | HSPG2 | na | 14764 | -0.408 | -0.0958 | No | ||
| 165 | CD44 | na | 14797 | -0.411 | -0.0924 | No | ||
| 166 | GPC4 | na | 14801 | -0.412 | -0.0875 | No | ||
| 167 | CHST7 | na | 14809 | -0.412 | -0.0827 | No | ||
| 168 | HYAL2 | na | 14900 | -0.423 | -0.0825 | No | ||
| 169 | NUP43 | na | 14925 | -0.425 | -0.0785 | No | ||
| 170 | CHST5 | na | 15475 | -0.491 | -0.1034 | No | ||
| 171 | SLC37A4 | na | 15501 | -0.494 | -0.0986 | No | ||
| 172 | HPSE | na | 15533 | -0.502 | -0.0941 | No | ||
| 173 | NUP107 | na | 15547 | -0.503 | -0.0885 | No | ||
| 174 | GPC2 | na | 15964 | -0.571 | -0.1049 | No | ||
| 175 | PHKG1 | na | 15983 | -0.574 | -0.0988 | No | ||
| 176 | SLC25A10 | na | 15995 | -0.577 | -0.0921 | No | ||
| 177 | KHK | na | 16076 | -0.594 | -0.0892 | No | ||
| 178 | GPC6 | na | 16129 | -0.600 | -0.0846 | No | ||
| 179 | GPC1 | na | 16163 | -0.606 | -0.0789 | No | ||
| 180 | PFKP | na | 16236 | -0.622 | -0.0752 | No | ||
| 181 | CSGALNACT2 | na | 16260 | -0.629 | -0.0686 | No | ||
| 182 | PRELP | na | 16392 | -0.657 | -0.0678 | No | ||
| 183 | ST3GAL3 | na | 16916 | -0.829 | -0.0870 | No | ||
| 184 | GALK1 | na | 16940 | -0.840 | -0.0778 | No | ||
| 185 | ENO3 | na | 16977 | -0.859 | -0.0690 | No | ||
| 186 | CSPG4 | na | 16985 | -0.862 | -0.0586 | No | ||
| 187 | PRPS1L1 | na | 16994 | -0.867 | -0.0482 | No | ||
| 188 | AGRN | na | 17067 | -0.904 | -0.0409 | No | ||
| 189 | CSGALNACT1 | na | 17077 | -0.907 | -0.0301 | No | ||
| 190 | STAB2 | na | 17126 | -0.927 | -0.0211 | No | ||
| 191 | SDC1 | na | 17165 | -0.952 | -0.0114 | No | ||
| 192 | GPC3 | na | 17479 | -1.197 | -0.0141 | No | ||
| 193 | B3GNT2 | na | 17579 | -1.316 | -0.0032 | No | ||
| 194 | CHST12 | na | 17646 | -1.431 | 0.0110 | No |