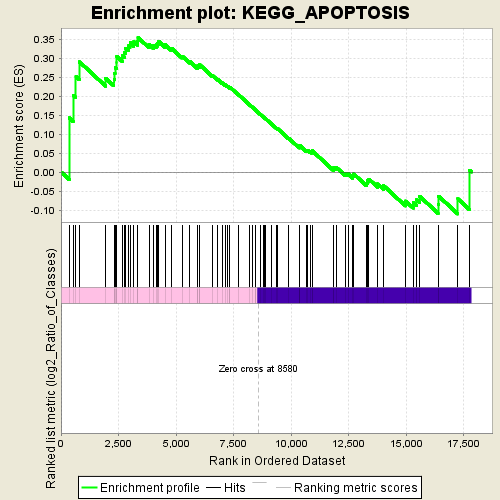
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA_1.class_1.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class_1.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | AHRR-overexpressed |
| GeneSet | KEGG_APOPTOSIS |
| Enrichment Score (ES) | 0.3543068 |
| Normalized Enrichment Score (NES) | 1.521201 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.5119045 |
| FWER p-Value | 0.752 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | IL1A | na | 357 | 5.223 | 0.1427 | Yes | ||
| 2 | NTRK1 | na | 526 | 2.210 | 0.2022 | Yes | ||
| 3 | IL1B | na | 647 | 1.784 | 0.2510 | Yes | ||
| 4 | PIK3R3 | na | 800 | 1.527 | 0.2901 | Yes | ||
| 5 | IL1R1 | na | 1939 | 0.669 | 0.2469 | Yes | ||
| 6 | CFLAR | na | 2300 | 0.561 | 0.2441 | Yes | ||
| 7 | MAP3K14 | na | 2331 | 0.553 | 0.2596 | Yes | ||
| 8 | PRKACA | na | 2348 | 0.546 | 0.2758 | Yes | ||
| 9 | TNFRSF10A | na | 2422 | 0.530 | 0.2882 | Yes | ||
| 10 | CASP6 | na | 2424 | 0.530 | 0.3046 | Yes | ||
| 11 | FAS | na | 2657 | 0.471 | 0.3062 | Yes | ||
| 12 | CASP9 | na | 2764 | 0.449 | 0.3143 | Yes | ||
| 13 | BID | na | 2802 | 0.440 | 0.3259 | Yes | ||
| 14 | TNFRSF10D | na | 2921 | 0.416 | 0.3322 | Yes | ||
| 15 | PRKAR2B | na | 2997 | 0.404 | 0.3406 | Yes | ||
| 16 | TNFRSF10B | na | 3166 | 0.372 | 0.3427 | Yes | ||
| 17 | MYD88 | na | 3321 | 0.345 | 0.3448 | Yes | ||
| 18 | AIFM1 | na | 3343 | 0.342 | 0.3543 | Yes | ||
| 19 | PIK3R2 | na | 3827 | 0.284 | 0.3360 | No | ||
| 20 | DFFB | na | 4026 | 0.264 | 0.3331 | No | ||
| 21 | PIK3CG | na | 4134 | 0.252 | 0.3349 | No | ||
| 22 | PIK3CB | na | 4195 | 0.247 | 0.3392 | No | ||
| 23 | TP53 | na | 4247 | 0.242 | 0.3439 | No | ||
| 24 | RIPK1 | na | 4519 | 0.219 | 0.3355 | No | ||
| 25 | TRADD | na | 4805 | 0.199 | 0.3256 | No | ||
| 26 | PRKAR1B | na | 5258 | 0.169 | 0.3054 | No | ||
| 27 | CYCS | na | 5589 | 0.149 | 0.2915 | No | ||
| 28 | BAD | na | 5928 | 0.127 | 0.2764 | No | ||
| 29 | CASP7 | na | 5945 | 0.126 | 0.2795 | No | ||
| 30 | PRKAR2A | na | 6029 | 0.121 | 0.2786 | No | ||
| 31 | BAX | na | 6030 | 0.121 | 0.2823 | No | ||
| 32 | AKT2 | na | 6591 | 0.090 | 0.2536 | No | ||
| 33 | RELA | na | 6799 | 0.080 | 0.2444 | No | ||
| 34 | TNFRSF1A | na | 7010 | 0.069 | 0.2348 | No | ||
| 35 | CAPN1 | na | 7138 | 0.063 | 0.2296 | No | ||
| 36 | PIK3CA | na | 7226 | 0.059 | 0.2265 | No | ||
| 37 | IRAK4 | na | 7331 | 0.054 | 0.2224 | No | ||
| 38 | IKBKG | na | 7342 | 0.054 | 0.2235 | No | ||
| 39 | IRAK1 | na | 7711 | 0.037 | 0.2039 | No | ||
| 40 | FADD | na | 8212 | 0.015 | 0.1762 | No | ||
| 41 | IL1RAP | na | 8306 | 0.010 | 0.1713 | No | ||
| 42 | DFFA | na | 8438 | 0.004 | 0.1641 | No | ||
| 43 | CHUK | na | 8441 | 0.004 | 0.1641 | No | ||
| 44 | BCL2L1 | na | 8656 | -0.006 | 0.1522 | No | ||
| 45 | CSF2RB | na | 8801 | -0.012 | 0.1444 | No | ||
| 46 | NFKB1 | na | 8854 | -0.014 | 0.1420 | No | ||
| 47 | EXOG | na | 8891 | -0.016 | 0.1404 | No | ||
| 48 | XIAP | na | 9135 | -0.025 | 0.1276 | No | ||
| 49 | PPP3CB | na | 9381 | -0.037 | 0.1149 | No | ||
| 50 | PRKAR1A | na | 9419 | -0.038 | 0.1140 | No | ||
| 51 | TRAF2 | na | 9894 | -0.061 | 0.0892 | No | ||
| 52 | APAF1 | na | 10357 | -0.082 | 0.0658 | No | ||
| 53 | ENDOG | na | 10365 | -0.083 | 0.0680 | No | ||
| 54 | PIK3R1 | na | 10367 | -0.083 | 0.0705 | No | ||
| 55 | AKT1 | na | 10681 | -0.098 | 0.0559 | No | ||
| 56 | CASP3 | na | 10710 | -0.099 | 0.0575 | No | ||
| 57 | TNFSF10 | na | 10836 | -0.106 | 0.0537 | No | ||
| 58 | CASP8 | na | 10908 | -0.110 | 0.0532 | No | ||
| 59 | CASP10 | na | 10914 | -0.110 | 0.0563 | No | ||
| 60 | NFKBIA | na | 11856 | -0.159 | 0.0083 | No | ||
| 61 | BIRC2 | na | 11857 | -0.159 | 0.0132 | No | ||
| 62 | PPP3CC | na | 11970 | -0.164 | 0.0120 | No | ||
| 63 | PPP3R1 | na | 12369 | -0.189 | -0.0045 | No | ||
| 64 | PPP3CA | na | 12475 | -0.195 | -0.0043 | No | ||
| 65 | PIK3R5 | na | 12688 | -0.209 | -0.0097 | No | ||
| 66 | IKBKB | na | 12722 | -0.211 | -0.0050 | No | ||
| 67 | BIRC3 | na | 13284 | -0.248 | -0.0288 | No | ||
| 68 | ATM | na | 13314 | -0.250 | -0.0227 | No | ||
| 69 | CAPN2 | na | 13378 | -0.253 | -0.0183 | No | ||
| 70 | PRKACB | na | 13764 | -0.285 | -0.0311 | No | ||
| 71 | PIK3CD | na | 14023 | -0.311 | -0.0360 | No | ||
| 72 | IRAK3 | na | 14973 | -0.424 | -0.0762 | No | ||
| 73 | PRKX | na | 15305 | -0.473 | -0.0801 | No | ||
| 74 | AKT3 | na | 15461 | -0.503 | -0.0731 | No | ||
| 75 | ENDOD1 | na | 15589 | -0.532 | -0.0637 | No | ||
| 76 | TNFRSF10C | na | 16391 | -0.745 | -0.0856 | No | ||
| 77 | IRAK2 | na | 16410 | -0.749 | -0.0632 | No | ||
| 78 | BCL2 | na | 17251 | -1.278 | -0.0707 | No | ||
| 79 | TNF | na | 17754 | -3.330 | 0.0049 | No |