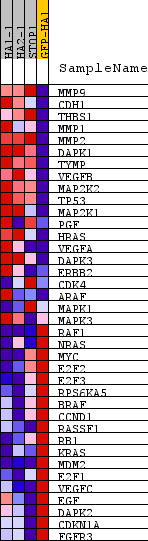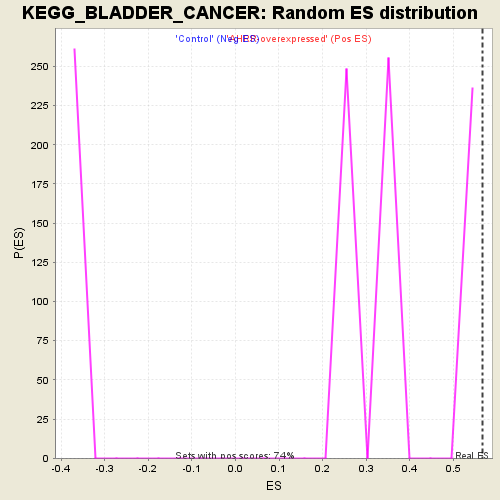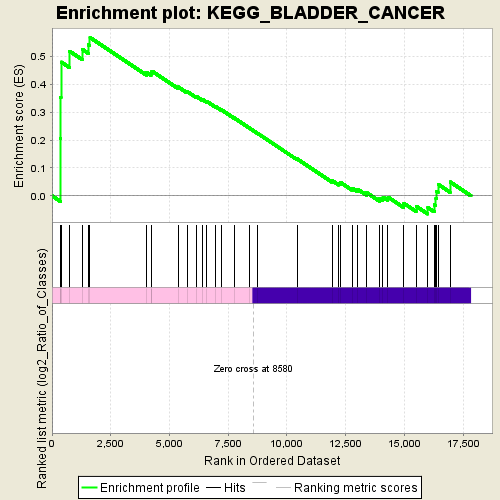
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA_1.class_1.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class_1.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | AHRR-overexpressed |
| GeneSet | KEGG_BLADDER_CANCER |
| Enrichment Score (ES) | 0.5676621 |
| Normalized Enrichment Score (NES) | 1.4701294 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.6252251 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MMP9 | na | 353 | 6.185 | 0.2069 | Yes | ||
| 2 | CDH1 | na | 365 | 4.030 | 0.3541 | Yes | ||
| 3 | THBS1 | na | 383 | 3.455 | 0.4798 | Yes | ||
| 4 | MMP1 | na | 741 | 1.611 | 0.5188 | Yes | ||
| 5 | MMP2 | na | 1295 | 1.005 | 0.5246 | Yes | ||
| 6 | DAPK1 | na | 1548 | 0.849 | 0.5416 | Yes | ||
| 7 | TYMP | na | 1614 | 0.811 | 0.5677 | Yes | ||
| 8 | VEGFB | na | 4023 | 0.264 | 0.4421 | No | ||
| 9 | MAP2K2 | na | 4215 | 0.245 | 0.4404 | No | ||
| 10 | TP53 | na | 4247 | 0.242 | 0.4475 | No | ||
| 11 | MAP2K1 | na | 5369 | 0.163 | 0.3905 | No | ||
| 12 | PGF | na | 5750 | 0.139 | 0.3742 | No | ||
| 13 | HRAS | na | 6149 | 0.115 | 0.3561 | No | ||
| 14 | VEGFA | na | 6386 | 0.101 | 0.3465 | No | ||
| 15 | DAPK3 | na | 6586 | 0.090 | 0.3387 | No | ||
| 16 | ERBB2 | na | 6960 | 0.072 | 0.3204 | No | ||
| 17 | CDK4 | na | 7209 | 0.060 | 0.3086 | No | ||
| 18 | ARAF | na | 7751 | 0.035 | 0.2795 | No | ||
| 19 | MAPK1 | na | 8426 | 0.004 | 0.2418 | No | ||
| 20 | MAPK3 | na | 8739 | -0.009 | 0.2246 | No | ||
| 21 | RAF1 | na | 10433 | -0.086 | 0.1327 | No | ||
| 22 | NRAS | na | 11941 | -0.163 | 0.0540 | No | ||
| 23 | MYC | na | 12214 | -0.178 | 0.0453 | No | ||
| 24 | E2F2 | na | 12275 | -0.182 | 0.0486 | No | ||
| 25 | E2F3 | na | 12784 | -0.215 | 0.0279 | No | ||
| 26 | RPS6KA5 | na | 13023 | -0.231 | 0.0230 | No | ||
| 27 | BRAF | na | 13396 | -0.254 | 0.0115 | No | ||
| 28 | CCND1 | na | 13947 | -0.303 | -0.0083 | No | ||
| 29 | RASSF1 | na | 14091 | -0.317 | -0.0048 | No | ||
| 30 | RB1 | na | 14304 | -0.340 | -0.0042 | No | ||
| 31 | KRAS | na | 14978 | -0.424 | -0.0265 | No | ||
| 32 | MDM2 | na | 15504 | -0.514 | -0.0371 | No | ||
| 33 | E2F1 | na | 16003 | -0.623 | -0.0422 | No | ||
| 34 | VEGFC | na | 16281 | -0.710 | -0.0318 | No | ||
| 35 | EGF | na | 16345 | -0.732 | -0.0085 | No | ||
| 36 | DAPK2 | na | 16378 | -0.741 | 0.0169 | No | ||
| 37 | CDKN1A | na | 16439 | -0.757 | 0.0413 | No | ||
| 38 | FGFR3 | na | 16951 | -1.020 | 0.0500 | No |