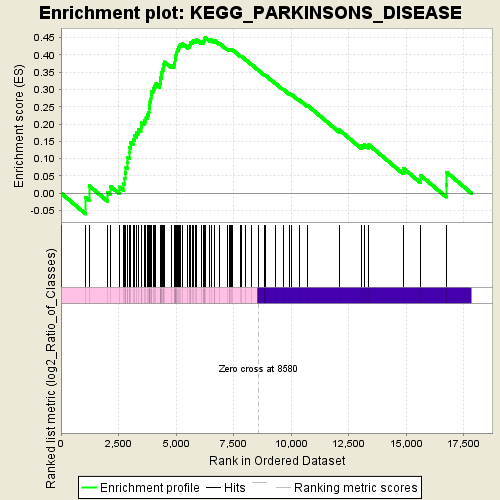
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA_1.class_1.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class_1.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | AHRR-overexpressed |
| GeneSet | KEGG_PARKINSONS_DISEASE |
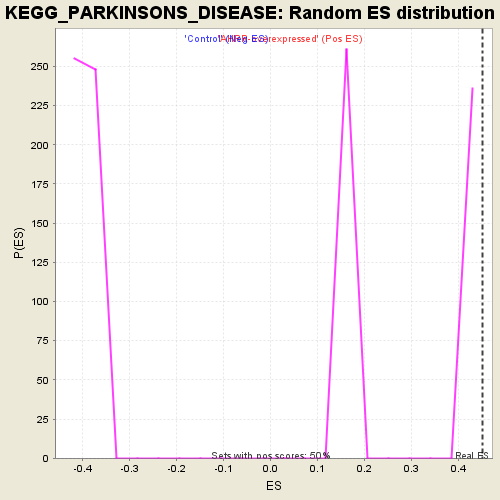
| Enrichment Score (ES) | 0.45049366 |
| Normalized Enrichment Score (NES) | 1.4902834 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.63404566 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | COX8C | na | 19 | 9223372036854775807.-1-1-1 | -0.0007 | Yes | ||
| 2 | LRRK2 | na | 1071 | 1.179 | -0.0124 | Yes | ||
| 3 | SLC25A4 | na | 1217 | 1.054 | 0.0220 | Yes | ||
| 4 | COX6B2 | na | 2010 | 0.648 | 0.0034 | Yes | ||
| 5 | NDUFS3 | na | 2132 | 0.605 | 0.0210 | Yes | ||
| 6 | NDUFA1 | na | 2541 | 0.500 | 0.0182 | Yes | ||
| 7 | SLC18A1 | na | 2708 | 0.461 | 0.0274 | Yes | ||
| 8 | CASP9 | na | 2764 | 0.449 | 0.0424 | Yes | ||
| 9 | NDUFA9 | na | 2783 | 0.444 | 0.0593 | Yes | ||
| 10 | COX6B1 | na | 2818 | 0.438 | 0.0751 | Yes | ||
| 11 | ATP5A1 | na | 2884 | 0.424 | 0.0885 | Yes | ||
| 12 | UBE2L6 | na | 2900 | 0.421 | 0.1046 | Yes | ||
| 13 | NDUFS7 | na | 2965 | 0.411 | 0.1176 | Yes | ||
| 14 | NDUFB2 | na | 2973 | 0.409 | 0.1337 | Yes | ||
| 15 | COX4I1 | na | 3028 | 0.398 | 0.1467 | Yes | ||
| 16 | ATP5G3 | na | 3153 | 0.374 | 0.1548 | Yes | ||
| 17 | UQCRFS1 | na | 3198 | 0.366 | 0.1671 | Yes | ||
| 18 | ATP5B | na | 3283 | 0.350 | 0.1765 | Yes | ||
| 19 | SDHB | na | 3375 | 0.338 | 0.1850 | Yes | ||
| 20 | NDUFC1 | na | 3475 | 0.326 | 0.1925 | Yes | ||
| 21 | SLC25A5 | na | 3501 | 0.323 | 0.2041 | Yes | ||
| 22 | UQCRQ | na | 3614 | 0.308 | 0.2102 | Yes | ||
| 23 | NDUFA4 | na | 3691 | 0.299 | 0.2180 | Yes | ||
| 24 | NDUFS6 | na | 3745 | 0.293 | 0.2269 | Yes | ||
| 25 | NDUFA2 | na | 3809 | 0.287 | 0.2349 | Yes | ||
| 26 | COX8A | na | 3825 | 0.284 | 0.2455 | Yes | ||
| 27 | UQCR10 | na | 3851 | 0.281 | 0.2554 | Yes | ||
| 28 | UQCR11 | na | 3864 | 0.281 | 0.2661 | Yes | ||
| 29 | COX7C | na | 3907 | 0.277 | 0.2749 | Yes | ||
| 30 | NDUFB8 | na | 3914 | 0.276 | 0.2856 | Yes | ||
| 31 | NDUFA3 | na | 3945 | 0.273 | 0.2949 | Yes | ||
| 32 | NDUFV1 | na | 3998 | 0.266 | 0.3027 | Yes | ||
| 33 | PARK7 | na | 4053 | 0.261 | 0.3102 | Yes | ||
| 34 | COX6A1 | na | 4124 | 0.253 | 0.3165 | Yes | ||
| 35 | UBB | na | 4300 | 0.237 | 0.3162 | Yes | ||
| 36 | PARK2 | na | 4306 | 0.237 | 0.3254 | Yes | ||
| 37 | ATP5F1 | na | 4320 | 0.234 | 0.3342 | Yes | ||
| 38 | CYC1 | na | 4356 | 0.232 | 0.3415 | Yes | ||
| 39 | COX7B | na | 4366 | 0.231 | 0.3503 | Yes | ||
| 40 | NDUFB7 | na | 4394 | 0.229 | 0.3580 | Yes | ||
| 41 | UQCRC1 | na | 4453 | 0.225 | 0.3638 | Yes | ||
| 42 | NDUFB1 | na | 4457 | 0.224 | 0.3727 | Yes | ||
| 43 | UQCRC2 | na | 4492 | 0.222 | 0.3797 | Yes | ||
| 44 | ATP5G2 | na | 4801 | 0.199 | 0.3704 | Yes | ||
| 45 | NDUFB10 | na | 4910 | 0.191 | 0.3720 | Yes | ||
| 46 | SDHD | na | 4941 | 0.189 | 0.3779 | Yes | ||
| 47 | COX7A2L | na | 4952 | 0.188 | 0.3850 | Yes | ||
| 48 | ATP5D | na | 4959 | 0.188 | 0.3922 | Yes | ||
| 49 | COX5A | na | 4986 | 0.186 | 0.3982 | Yes | ||
| 50 | NDUFS8 | na | 5004 | 0.185 | 0.4047 | Yes | ||
| 51 | UQCRB | na | 5038 | 0.183 | 0.4102 | Yes | ||
| 52 | NDUFA10 | na | 5053 | 0.182 | 0.4168 | Yes | ||
| 53 | NDUFS5 | na | 5108 | 0.178 | 0.4209 | Yes | ||
| 54 | COX5B | na | 5128 | 0.177 | 0.4270 | Yes | ||
| 55 | ATP5E | na | 5185 | 0.173 | 0.4308 | Yes | ||
| 56 | SDHC | na | 5276 | 0.168 | 0.4325 | Yes | ||
| 57 | UBE2J2 | na | 5514 | 0.153 | 0.4253 | Yes | ||
| 58 | CYCS | na | 5589 | 0.149 | 0.4271 | Yes | ||
| 59 | UBA1 | na | 5608 | 0.147 | 0.4321 | Yes | ||
| 60 | ATP5G1 | na | 5624 | 0.146 | 0.4371 | Yes | ||
| 61 | NDUFAB1 | na | 5711 | 0.141 | 0.4379 | Yes | ||
| 62 | NDUFB5 | na | 5749 | 0.139 | 0.4414 | Yes | ||
| 63 | NDUFA8 | na | 5829 | 0.133 | 0.4424 | Yes | ||
| 64 | NDUFV3 | na | 5895 | 0.129 | 0.4439 | Yes | ||
| 65 | UQCRH | na | 6086 | 0.118 | 0.4380 | Yes | ||
| 66 | UBE2L3 | na | 6182 | 0.113 | 0.4372 | Yes | ||
| 67 | VDAC1 | na | 6207 | 0.112 | 0.4403 | Yes | ||
| 68 | ATP5H | na | 6219 | 0.111 | 0.4442 | Yes | ||
| 69 | COX6C | na | 6247 | 0.110 | 0.4471 | Yes | ||
| 70 | NDUFA7 | na | 6265 | 0.109 | 0.4505 | Yes | ||
| 71 | ATP5O | na | 6464 | 0.097 | 0.4432 | No | ||
| 72 | NDUFA6 | na | 6524 | 0.093 | 0.4437 | No | ||
| 73 | SDHA | na | 6673 | 0.086 | 0.4388 | No | ||
| 74 | SLC18A2 | na | 6688 | 0.085 | 0.4414 | No | ||
| 75 | ATP5C1 | na | 6908 | 0.075 | 0.4321 | No | ||
| 76 | HTRA2 | na | 7220 | 0.060 | 0.4170 | No | ||
| 77 | NDUFC2 | na | 7301 | 0.056 | 0.4147 | No | ||
| 78 | NDUFS2 | na | 7320 | 0.055 | 0.4159 | No | ||
| 79 | NDUFB3 | na | 7362 | 0.053 | 0.4157 | No | ||
| 80 | VDAC2 | na | 7410 | 0.050 | 0.4151 | No | ||
| 81 | NDUFS1 | na | 7440 | 0.049 | 0.4155 | No | ||
| 82 | UBE2G1 | na | 7814 | 0.032 | 0.3957 | No | ||
| 83 | NDUFB9 | na | 7842 | 0.031 | 0.3955 | No | ||
| 84 | COX7A2 | na | 8032 | 0.023 | 0.3858 | No | ||
| 85 | SEPT5 | na | 8258 | 0.013 | 0.3736 | No | ||
| 86 | ATP5J | na | 8570 | -0.002 | 0.3561 | No | ||
| 87 | PPID | na | 8824 | -0.013 | 0.3424 | No | ||
| 88 | NDUFA4L2 | na | 8857 | -0.015 | 0.3412 | No | ||
| 89 | UBE2G2 | na | 8863 | -0.015 | 0.3415 | No | ||
| 90 | NDUFA5 | na | 8894 | -0.016 | 0.3404 | No | ||
| 91 | NDUFB4 | na | 9320 | -0.034 | 0.3178 | No | ||
| 92 | VDAC3 | na | 9660 | -0.050 | 0.3008 | No | ||
| 93 | NDUFV2 | na | 9908 | -0.061 | 0.2893 | No | ||
| 94 | NDUFB6 | na | 10031 | -0.067 | 0.2851 | No | ||
| 95 | APAF1 | na | 10357 | -0.082 | 0.2701 | No | ||
| 96 | CASP3 | na | 10710 | -0.099 | 0.2543 | No | ||
| 97 | NDUFS4 | na | 12094 | -0.171 | 0.1832 | No | ||
| 98 | PINK1 | na | 13071 | -0.234 | 0.1376 | No | ||
| 99 | UQCRHL | na | 13185 | -0.241 | 0.1410 | No | ||
| 100 | SNCAIP | na | 13379 | -0.253 | 0.1403 | No | ||
| 101 | UBA7 | na | 14879 | -0.410 | 0.0723 | No | ||
| 102 | UBE2J1 | na | 15629 | -0.542 | 0.0520 | No | ||
| 103 | COX7A1 | na | 16743 | -0.891 | 0.0252 | No | ||
| 104 | GPR37 | na | 16772 | -0.910 | 0.0603 | No |