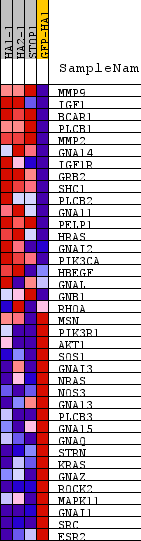
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA_1.class_1.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class_1.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | AHRR-overexpressed |
| GeneSet | PID_ER_NONGENOMIC_PATHWAY |
| Enrichment Score (ES) | 0.47279847 |
| Normalized Enrichment Score (NES) | 1.5948998 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.5370638 |
| FWER p-Value | 0.752 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MMP9 | na | 353 | 6.185 | 0.2760 | Yes | ||
| 2 | IGF1 | na | 639 | 1.802 | 0.3462 | Yes | ||
| 3 | BCAR1 | na | 993 | 1.258 | 0.3866 | Yes | ||
| 4 | PLCB1 | na | 1109 | 1.149 | 0.4351 | Yes | ||
| 5 | MMP2 | na | 1295 | 1.005 | 0.4728 | Yes | ||
| 6 | GNA14 | na | 3103 | 0.382 | 0.3896 | No | ||
| 7 | IGF1R | na | 4432 | 0.227 | 0.3258 | No | ||
| 8 | GRB2 | na | 4599 | 0.212 | 0.3267 | No | ||
| 9 | SHC1 | na | 4623 | 0.211 | 0.3354 | No | ||
| 10 | PLCB2 | na | 4943 | 0.189 | 0.3266 | No | ||
| 11 | GNA11 | na | 5261 | 0.168 | 0.3168 | No | ||
| 12 | PELP1 | na | 5341 | 0.164 | 0.3202 | No | ||
| 13 | HRAS | na | 6149 | 0.115 | 0.2804 | No | ||
| 14 | GNAI2 | na | 6766 | 0.082 | 0.2497 | No | ||
| 15 | PIK3CA | na | 7226 | 0.059 | 0.2268 | No | ||
| 16 | HBEGF | na | 7271 | 0.058 | 0.2270 | No | ||
| 17 | GNAL | na | 7337 | 0.054 | 0.2260 | No | ||
| 18 | GNB1 | na | 7569 | 0.044 | 0.2151 | No | ||
| 19 | RHOA | na | 8702 | -0.007 | 0.1519 | No | ||
| 20 | MSN | na | 9880 | -0.060 | 0.0886 | No | ||
| 21 | PIK3R1 | na | 10367 | -0.083 | 0.0653 | No | ||
| 22 | AKT1 | na | 10681 | -0.098 | 0.0524 | No | ||
| 23 | SOS1 | na | 11594 | -0.144 | 0.0081 | No | ||
| 24 | GNAI3 | na | 11705 | -0.150 | 0.0090 | No | ||
| 25 | NRAS | na | 11941 | -0.163 | 0.0036 | No | ||
| 26 | NOS3 | na | 12263 | -0.181 | -0.0057 | No | ||
| 27 | GNA13 | na | 12586 | -0.202 | -0.0141 | No | ||
| 28 | PLCB3 | na | 13597 | -0.269 | -0.0580 | No | ||
| 29 | GNA15 | na | 14622 | -0.375 | -0.0976 | No | ||
| 30 | GNAQ | na | 14664 | -0.379 | -0.0817 | No | ||
| 31 | STRN | na | 14692 | -0.382 | -0.0650 | No | ||
| 32 | KRAS | na | 14978 | -0.424 | -0.0607 | No | ||
| 33 | GNAZ | na | 15401 | -0.493 | -0.0608 | No | ||
| 34 | ROCK2 | na | 15664 | -0.549 | -0.0493 | No | ||
| 35 | MAPK11 | na | 15731 | -0.564 | -0.0260 | No | ||
| 36 | GNAI1 | na | 15908 | -0.601 | -0.0072 | No | ||
| 37 | SRC | na | 16527 | -0.788 | -0.0042 | No | ||
| 38 | ESR2 | na | 17468 | -1.630 | 0.0210 | No |