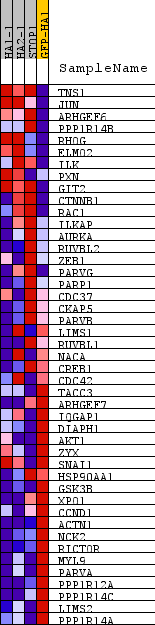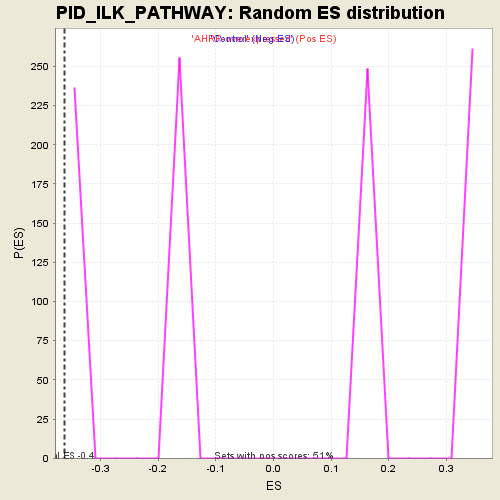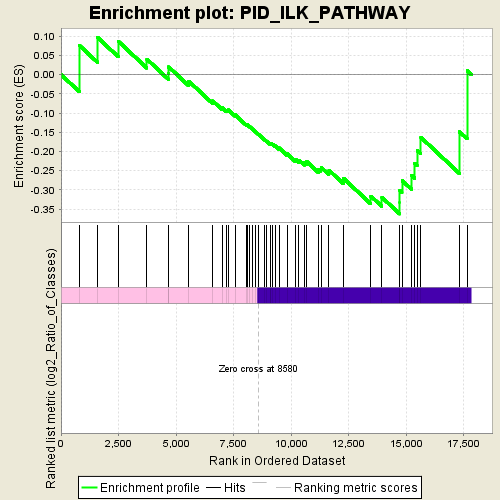
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA_1.class_1.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class_1.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | Control |
| GeneSet | PID_ILK_PATHWAY |
| Enrichment Score (ES) | -0.3630438 |
| Normalized Enrichment Score (NES) | -1.4060446 |
| Nominal p-value | 0.0 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TNS1 | na | 798 | 1.531 | 0.0760 | No | ||
| 2 | JUN | na | 1588 | 0.825 | 0.0968 | No | ||
| 3 | ARHGEF6 | na | 2487 | 0.513 | 0.0868 | No | ||
| 4 | PPP1R14B | na | 3730 | 0.295 | 0.0403 | No | ||
| 5 | RHOG | na | 4666 | 0.209 | 0.0042 | No | ||
| 6 | ELMO2 | na | 4676 | 0.208 | 0.0201 | No | ||
| 7 | ILK | na | 5550 | 0.151 | -0.0170 | No | ||
| 8 | PXN | na | 6592 | 0.090 | -0.0684 | No | ||
| 9 | GIT2 | na | 7001 | 0.070 | -0.0859 | No | ||
| 10 | CTNNB1 | na | 7200 | 0.060 | -0.0922 | No | ||
| 11 | RAC1 | na | 7273 | 0.058 | -0.0917 | No | ||
| 12 | ILKAP | na | 7565 | 0.044 | -0.1046 | No | ||
| 13 | AURKA | na | 8076 | 0.021 | -0.1316 | No | ||
| 14 | RUVBL2 | na | 8082 | 0.021 | -0.1302 | No | ||
| 15 | ZEB1 | na | 8197 | 0.015 | -0.1354 | No | ||
| 16 | PARVG | na | 8326 | 0.009 | -0.1419 | No | ||
| 17 | PARP1 | na | 8451 | 0.003 | -0.1486 | No | ||
| 18 | CDC37 | na | 8577 | -0.002 | -0.1554 | No | ||
| 19 | CKAP5 | na | 8845 | -0.014 | -0.1693 | No | ||
| 20 | PARVB | na | 8947 | -0.018 | -0.1736 | No | ||
| 21 | LIMS1 | na | 9105 | -0.024 | -0.1805 | No | ||
| 22 | RUVBL1 | na | 9109 | -0.024 | -0.1787 | No | ||
| 23 | NACA | na | 9188 | -0.028 | -0.1809 | No | ||
| 24 | CREB1 | na | 9319 | -0.034 | -0.1855 | No | ||
| 25 | CDC42 | na | 9474 | -0.041 | -0.1909 | No | ||
| 26 | TACC3 | na | 9822 | -0.058 | -0.2059 | No | ||
| 27 | ARHGEF7 | na | 10180 | -0.074 | -0.2201 | No | ||
| 28 | IQGAP1 | na | 10341 | -0.081 | -0.2226 | No | ||
| 29 | DIAPH1 | na | 10574 | -0.093 | -0.2283 | No | ||
| 30 | AKT1 | na | 10681 | -0.098 | -0.2265 | No | ||
| 31 | ZYX | na | 11201 | -0.125 | -0.2458 | No | ||
| 32 | SNAI1 | na | 11300 | -0.129 | -0.2412 | No | ||
| 33 | HSP90AA1 | na | 11645 | -0.146 | -0.2490 | No | ||
| 34 | GSK3B | na | 12277 | -0.183 | -0.2700 | No | ||
| 35 | XPO1 | na | 13467 | -0.258 | -0.3165 | No | ||
| 36 | CCND1 | na | 13947 | -0.303 | -0.3195 | No | ||
| 37 | ACTN1 | na | 14723 | -0.388 | -0.3324 | Yes | ||
| 38 | NCK2 | na | 14726 | -0.388 | -0.3019 | Yes | ||
| 39 | RICTOR | na | 14822 | -0.401 | -0.2756 | Yes | ||
| 40 | MYL9 | na | 15235 | -0.462 | -0.2622 | Yes | ||
| 41 | PARVA | na | 15376 | -0.488 | -0.2316 | Yes | ||
| 42 | PPP1R12A | na | 15479 | -0.508 | -0.1972 | Yes | ||
| 43 | PPP1R14C | na | 15642 | -0.544 | -0.1634 | Yes | ||
| 44 | LIMS2 | na | 17322 | -1.378 | -0.1489 | Yes | ||
| 45 | PPP1R14A | na | 17654 | -2.256 | 0.0105 | Yes |