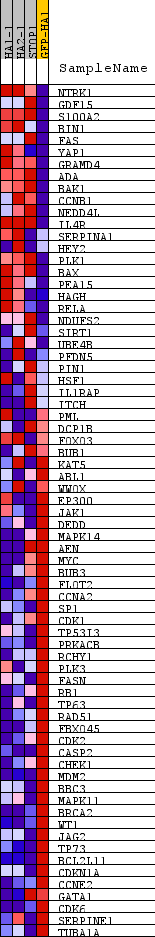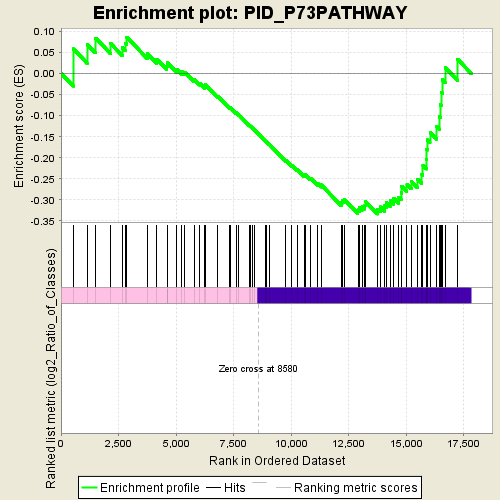
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA_1.class_1.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class_1.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | Control |
| GeneSet | PID_P73PATHWAY |
| Enrichment Score (ES) | -0.33376488 |
| Normalized Enrichment Score (NES) | -1.294659 |
| Nominal p-value | 0.0 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | NTRK1 | na | 526 | 2.210 | 0.0590 | No | ||
| 2 | GDF15 | na | 1134 | 1.124 | 0.0698 | No | ||
| 3 | S100A2 | na | 1492 | 0.871 | 0.0847 | No | ||
| 4 | BIN1 | na | 2138 | 0.604 | 0.0726 | No | ||
| 5 | FAS | na | 2657 | 0.471 | 0.0623 | No | ||
| 6 | YAP1 | na | 2786 | 0.444 | 0.0729 | No | ||
| 7 | GRAMD4 | na | 2859 | 0.430 | 0.0860 | No | ||
| 8 | ADA | na | 3748 | 0.293 | 0.0478 | No | ||
| 9 | BAK1 | na | 4161 | 0.250 | 0.0347 | No | ||
| 10 | CCNB1 | na | 4605 | 0.212 | 0.0182 | No | ||
| 11 | NEDD4L | na | 4624 | 0.211 | 0.0257 | No | ||
| 12 | IL4R | na | 5026 | 0.183 | 0.0104 | No | ||
| 13 | SERPINA1 | na | 5251 | 0.169 | 0.0046 | No | ||
| 14 | HEY2 | na | 5385 | 0.162 | 0.0036 | No | ||
| 15 | PLK1 | na | 5778 | 0.137 | -0.0130 | No | ||
| 16 | BAX | na | 6030 | 0.121 | -0.0223 | No | ||
| 17 | PEA15 | na | 6249 | 0.110 | -0.0301 | No | ||
| 18 | HAGH | na | 6260 | 0.109 | -0.0263 | No | ||
| 19 | RELA | na | 6799 | 0.080 | -0.0534 | No | ||
| 20 | NDUFS2 | na | 7320 | 0.055 | -0.0804 | No | ||
| 21 | SIRT1 | na | 7361 | 0.053 | -0.0806 | No | ||
| 22 | UBE4B | na | 7611 | 0.042 | -0.0929 | No | ||
| 23 | PFDN5 | na | 7731 | 0.036 | -0.0982 | No | ||
| 24 | PIN1 | na | 8188 | 0.016 | -0.1232 | No | ||
| 25 | HSF1 | na | 8220 | 0.015 | -0.1243 | No | ||
| 26 | IL1RAP | na | 8306 | 0.010 | -0.1287 | No | ||
| 27 | ITCH | na | 8416 | 0.005 | -0.1347 | No | ||
| 28 | PML | na | 8899 | -0.016 | -0.1611 | No | ||
| 29 | DCP1B | na | 8929 | -0.017 | -0.1621 | No | ||
| 30 | FOXO3 | na | 9055 | -0.022 | -0.1682 | No | ||
| 31 | BUB1 | na | 9775 | -0.055 | -0.2065 | No | ||
| 32 | KAT5 | na | 10023 | -0.067 | -0.2177 | No | ||
| 33 | ABL1 | na | 10269 | -0.079 | -0.2283 | No | ||
| 34 | WWOX | na | 10572 | -0.093 | -0.2416 | No | ||
| 35 | EP300 | na | 10620 | -0.095 | -0.2404 | No | ||
| 36 | JAK1 | na | 10839 | -0.106 | -0.2484 | No | ||
| 37 | DEDD | na | 11136 | -0.121 | -0.2602 | No | ||
| 38 | MAPK14 | na | 11305 | -0.129 | -0.2645 | No | ||
| 39 | AEN | na | 12185 | -0.176 | -0.3069 | No | ||
| 40 | MYC | na | 12214 | -0.178 | -0.3013 | No | ||
| 41 | BUB3 | na | 12316 | -0.185 | -0.2996 | No | ||
| 42 | FLOT2 | na | 12908 | -0.223 | -0.3239 | Yes | ||
| 43 | CCNA2 | na | 12975 | -0.227 | -0.3185 | Yes | ||
| 44 | SP1 | na | 13084 | -0.234 | -0.3152 | Yes | ||
| 45 | CDK1 | na | 13206 | -0.242 | -0.3123 | Yes | ||
| 46 | TP53I3 | na | 13247 | -0.246 | -0.3047 | Yes | ||
| 47 | PRKACB | na | 13764 | -0.285 | -0.3224 | Yes | ||
| 48 | RCHY1 | na | 13878 | -0.295 | -0.3169 | Yes | ||
| 49 | PLK3 | na | 14058 | -0.314 | -0.3144 | Yes | ||
| 50 | FASN | na | 14147 | -0.322 | -0.3064 | Yes | ||
| 51 | RB1 | na | 14304 | -0.340 | -0.3016 | Yes | ||
| 52 | TP63 | na | 14458 | -0.356 | -0.2959 | Yes | ||
| 53 | RAD51 | na | 14684 | -0.381 | -0.2933 | Yes | ||
| 54 | FBXO45 | na | 14784 | -0.397 | -0.2830 | Yes | ||
| 55 | CDK2 | na | 14803 | -0.399 | -0.2680 | Yes | ||
| 56 | CASP2 | na | 15037 | -0.433 | -0.2638 | Yes | ||
| 57 | CHEK1 | na | 15221 | -0.459 | -0.2557 | Yes | ||
| 58 | MDM2 | na | 15504 | -0.514 | -0.2510 | Yes | ||
| 59 | BBC3 | na | 15684 | -0.553 | -0.2389 | Yes | ||
| 60 | MAPK11 | na | 15731 | -0.564 | -0.2189 | Yes | ||
| 61 | BRCA2 | na | 15872 | -0.592 | -0.2030 | Yes | ||
| 62 | WT1 | na | 15889 | -0.595 | -0.1801 | Yes | ||
| 63 | JAG2 | na | 15917 | -0.603 | -0.1575 | Yes | ||
| 64 | TP73 | na | 16039 | -0.632 | -0.1390 | Yes | ||
| 65 | BCL2L11 | na | 16336 | -0.730 | -0.1264 | Yes | ||
| 66 | CDKN1A | na | 16439 | -0.757 | -0.1018 | Yes | ||
| 67 | CCNE2 | na | 16495 | -0.777 | -0.0738 | Yes | ||
| 68 | GATA1 | na | 16523 | -0.787 | -0.0437 | Yes | ||
| 69 | CDK6 | na | 16583 | -0.813 | -0.0145 | Yes | ||
| 70 | SERPINE1 | na | 16696 | -0.867 | 0.0140 | Yes | ||
| 71 | TUBA1A | na | 17236 | -1.257 | 0.0340 | Yes |