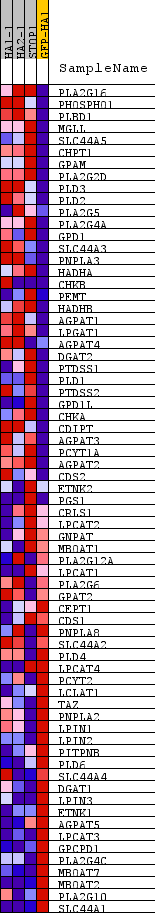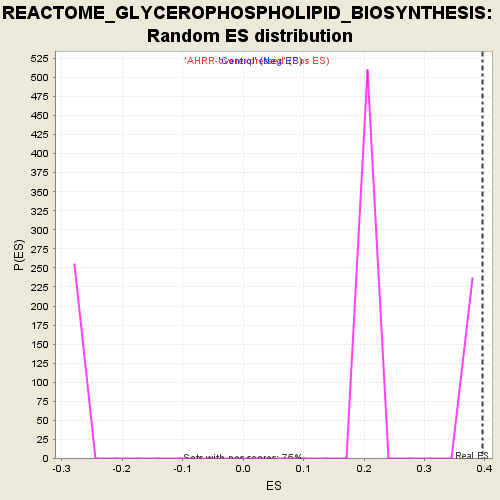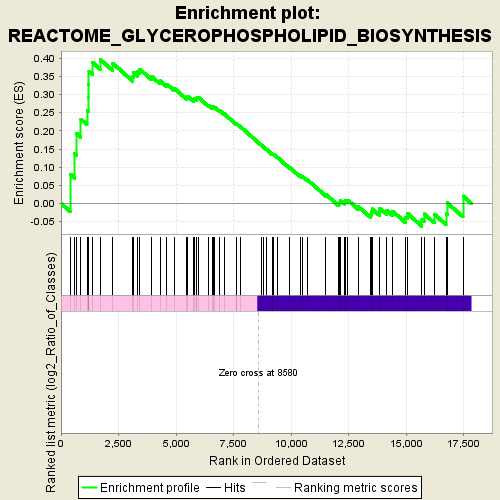
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA_1.class_1.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class_1.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | AHRR-overexpressed |
| GeneSet | REACTOME_GLYCEROPHOSPHOLIPID_BIOSYNTHESIS |
| Enrichment Score (ES) | 0.39623705 |
| Normalized Enrichment Score (NES) | 1.522868 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.5907338 |
| FWER p-Value | 0.752 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PLA2G16 | na | 415 | 2.975 | 0.0800 | Yes | ||
| 2 | PHOSPHO1 | na | 598 | 1.922 | 0.1366 | Yes | ||
| 3 | PLBD1 | na | 668 | 1.748 | 0.1935 | Yes | ||
| 4 | MGLL | na | 862 | 1.428 | 0.2322 | Yes | ||
| 5 | SLC44A5 | na | 1128 | 1.130 | 0.2566 | Yes | ||
| 6 | CHPT1 | na | 1184 | 1.086 | 0.2912 | Yes | ||
| 7 | GPAM | na | 1187 | 1.083 | 0.3288 | Yes | ||
| 8 | PLA2G2D | na | 1211 | 1.062 | 0.3644 | Yes | ||
| 9 | PLD3 | na | 1369 | 0.948 | 0.3885 | Yes | ||
| 10 | PLD2 | na | 1707 | 0.769 | 0.3962 | Yes | ||
| 11 | PLA2G5 | na | 2245 | 0.573 | 0.3859 | No | ||
| 12 | PLA2G4A | na | 3122 | 0.378 | 0.3498 | No | ||
| 13 | GPD1 | na | 3154 | 0.374 | 0.3611 | No | ||
| 14 | SLC44A3 | na | 3335 | 0.343 | 0.3629 | No | ||
| 15 | PNPLA3 | na | 3410 | 0.333 | 0.3702 | No | ||
| 16 | HADHA | na | 3935 | 0.273 | 0.3503 | No | ||
| 17 | CHKB | na | 4303 | 0.237 | 0.3379 | No | ||
| 18 | PEMT | na | 4598 | 0.212 | 0.3287 | No | ||
| 19 | HADHB | na | 4933 | 0.189 | 0.3165 | No | ||
| 20 | AGPAT1 | na | 5451 | 0.158 | 0.2929 | No | ||
| 21 | LPGAT1 | na | 5508 | 0.154 | 0.2951 | No | ||
| 22 | AGPAT4 | na | 5769 | 0.137 | 0.2852 | No | ||
| 23 | DGAT2 | na | 5794 | 0.135 | 0.2886 | No | ||
| 24 | PTDSS1 | na | 5880 | 0.130 | 0.2883 | No | ||
| 25 | PLD1 | na | 5908 | 0.128 | 0.2912 | No | ||
| 26 | PTDSS2 | na | 5952 | 0.126 | 0.2932 | No | ||
| 27 | GPD1L | na | 6416 | 0.099 | 0.2706 | No | ||
| 28 | CHKA | na | 6588 | 0.090 | 0.2641 | No | ||
| 29 | CDIPT | na | 6605 | 0.089 | 0.2663 | No | ||
| 30 | AGPAT3 | na | 6690 | 0.085 | 0.2645 | No | ||
| 31 | PCYT1A | na | 6892 | 0.076 | 0.2558 | No | ||
| 32 | AGPAT2 | na | 7090 | 0.065 | 0.2470 | No | ||
| 33 | CDS2 | na | 7616 | 0.041 | 0.2189 | No | ||
| 34 | ETNK2 | na | 7633 | 0.041 | 0.2194 | No | ||
| 35 | PGS1 | na | 7794 | 0.034 | 0.2116 | No | ||
| 36 | CRLS1 | na | 8731 | -0.008 | 0.1592 | No | ||
| 37 | LPCAT2 | na | 8779 | -0.011 | 0.1569 | No | ||
| 38 | GNPAT | na | 8930 | -0.017 | 0.1491 | No | ||
| 39 | MBOAT1 | na | 9174 | -0.027 | 0.1363 | No | ||
| 40 | PLA2G12A | na | 9199 | -0.028 | 0.1360 | No | ||
| 41 | LPCAT1 | na | 9241 | -0.030 | 0.1347 | No | ||
| 42 | PLA2G6 | na | 9402 | -0.037 | 0.1270 | No | ||
| 43 | GPAT2 | na | 9946 | -0.063 | 0.0986 | No | ||
| 44 | CEPT1 | na | 10389 | -0.084 | 0.0767 | No | ||
| 45 | CDS1 | na | 10482 | -0.089 | 0.0746 | No | ||
| 46 | PNPLA8 | na | 10705 | -0.099 | 0.0655 | No | ||
| 47 | SLC44A2 | na | 11510 | -0.139 | 0.0251 | No | ||
| 48 | PLD4 | na | 12058 | -0.169 | 0.0002 | No | ||
| 49 | LPCAT4 | na | 12104 | -0.172 | 0.0037 | No | ||
| 50 | PCYT2 | na | 12139 | -0.174 | 0.0078 | No | ||
| 51 | LCLAT1 | na | 12307 | -0.185 | 0.0048 | No | ||
| 52 | TAZ | na | 12360 | -0.188 | 0.0084 | No | ||
| 53 | PNPLA2 | na | 12470 | -0.195 | 0.0091 | No | ||
| 54 | LPIN1 | na | 12916 | -0.224 | -0.0082 | No | ||
| 55 | LPIN2 | na | 13470 | -0.259 | -0.0303 | No | ||
| 56 | PITPNB | na | 13507 | -0.261 | -0.0233 | No | ||
| 57 | PLD6 | na | 13524 | -0.263 | -0.0150 | No | ||
| 58 | SLC44A4 | na | 13837 | -0.291 | -0.0225 | No | ||
| 59 | DGAT1 | na | 13852 | -0.292 | -0.0131 | No | ||
| 60 | LPIN3 | na | 14158 | -0.323 | -0.0190 | No | ||
| 61 | ETNK1 | na | 14411 | -0.352 | -0.0210 | No | ||
| 62 | AGPAT5 | na | 14974 | -0.424 | -0.0379 | No | ||
| 63 | LPCAT3 | na | 15053 | -0.435 | -0.0272 | No | ||
| 64 | GPCPD1 | na | 15688 | -0.554 | -0.0436 | No | ||
| 65 | PLA2G4C | na | 15785 | -0.579 | -0.0289 | No | ||
| 66 | MBOAT7 | na | 16219 | -0.690 | -0.0292 | No | ||
| 67 | MBOAT2 | na | 16736 | -0.889 | -0.0274 | No | ||
| 68 | PLA2G10 | na | 16787 | -0.915 | 0.0016 | No | ||
| 69 | SLC44A1 | na | 17475 | -1.659 | 0.0206 | No |