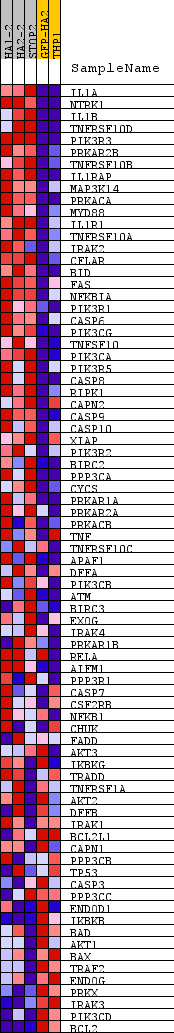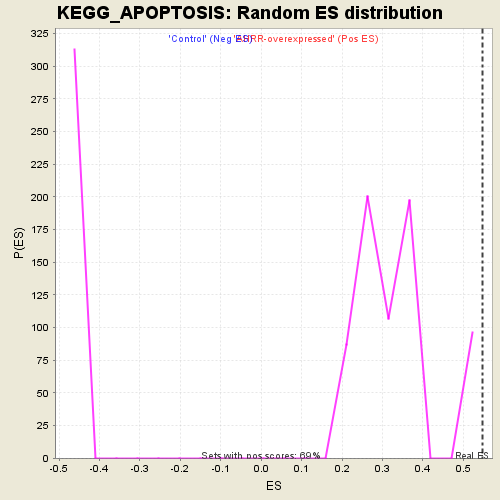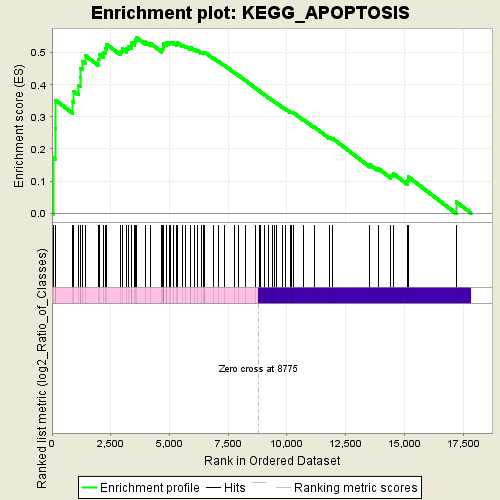
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA_2.class_2.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class_2.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | AHRR-overexpressed |
| GeneSet | KEGG_APOPTOSIS |
| Enrichment Score (ES) | 0.54737234 |
| Normalized Enrichment Score (NES) | 1.626635 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.18685265 |
| FWER p-Value | 0.491 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | IL1A | na | 49 | 5.421 | 0.1720 | Yes | ||
| 2 | NTRK1 | na | 131 | 2.985 | 0.2637 | Yes | ||
| 3 | IL1B | na | 148 | 2.766 | 0.3520 | Yes | ||
| 4 | TNFRSF10D | na | 884 | 1.123 | 0.3468 | Yes | ||
| 5 | PIK3R3 | na | 912 | 1.093 | 0.3805 | Yes | ||
| 6 | PRKAR2B | na | 1129 | 0.926 | 0.3982 | Yes | ||
| 7 | TNFRSF10B | na | 1198 | 0.885 | 0.4229 | Yes | ||
| 8 | IL1RAP | na | 1220 | 0.873 | 0.4499 | Yes | ||
| 9 | MAP3K14 | na | 1283 | 0.845 | 0.4737 | Yes | ||
| 10 | PRKACA | na | 1414 | 0.778 | 0.4914 | Yes | ||
| 11 | MYD88 | na | 1959 | 0.582 | 0.4796 | Yes | ||
| 12 | IL1R1 | na | 2000 | 0.567 | 0.4956 | Yes | ||
| 13 | TNFRSF10A | na | 2187 | 0.521 | 0.5019 | Yes | ||
| 14 | IRAK2 | na | 2278 | 0.495 | 0.5128 | Yes | ||
| 15 | CFLAR | na | 2308 | 0.489 | 0.5269 | Yes | ||
| 16 | BID | na | 2910 | 0.374 | 0.5052 | Yes | ||
| 17 | FAS | na | 2978 | 0.364 | 0.5131 | Yes | ||
| 18 | NFKBIA | na | 3178 | 0.332 | 0.5126 | Yes | ||
| 19 | PIK3R1 | na | 3251 | 0.323 | 0.5190 | Yes | ||
| 20 | CASP6 | na | 3377 | 0.306 | 0.5218 | Yes | ||
| 21 | PIK3CG | na | 3383 | 0.305 | 0.5314 | Yes | ||
| 22 | TNFSF10 | na | 3522 | 0.290 | 0.5329 | Yes | ||
| 23 | PIK3CA | na | 3554 | 0.286 | 0.5404 | Yes | ||
| 24 | PIK3R5 | na | 3593 | 0.282 | 0.5474 | Yes | ||
| 25 | CASP8 | na | 3981 | 0.244 | 0.5334 | No | ||
| 26 | RIPK1 | na | 4171 | 0.227 | 0.5301 | No | ||
| 27 | CAPN2 | na | 4678 | 0.188 | 0.5077 | No | ||
| 28 | CASP9 | na | 4717 | 0.185 | 0.5116 | No | ||
| 29 | CASP10 | na | 4725 | 0.185 | 0.5171 | No | ||
| 30 | XIAP | na | 4737 | 0.184 | 0.5224 | No | ||
| 31 | PIK3R2 | na | 4743 | 0.184 | 0.5281 | No | ||
| 32 | BIRC2 | na | 4866 | 0.175 | 0.5269 | No | ||
| 33 | PPP3CA | na | 4867 | 0.175 | 0.5325 | No | ||
| 34 | CYCS | na | 4997 | 0.166 | 0.5306 | No | ||
| 35 | PRKAR1A | na | 5050 | 0.163 | 0.5330 | No | ||
| 36 | PRKAR2A | na | 5185 | 0.155 | 0.5304 | No | ||
| 37 | PRKACB | na | 5312 | 0.148 | 0.5281 | No | ||
| 38 | TNF | na | 5334 | 0.147 | 0.5316 | No | ||
| 39 | TNFRSF10C | na | 5571 | 0.135 | 0.5227 | No | ||
| 40 | APAF1 | na | 5681 | 0.129 | 0.5207 | No | ||
| 41 | DFFA | na | 5882 | 0.119 | 0.5133 | No | ||
| 42 | PIK3CB | na | 5899 | 0.118 | 0.5162 | No | ||
| 43 | ATM | na | 6065 | 0.109 | 0.5105 | No | ||
| 44 | BIRC3 | na | 6183 | 0.104 | 0.5072 | No | ||
| 45 | EXOG | na | 6379 | 0.095 | 0.4993 | No | ||
| 46 | IRAK4 | na | 6437 | 0.092 | 0.4991 | No | ||
| 47 | PRKAR1B | na | 6473 | 0.091 | 0.5000 | No | ||
| 48 | RELA | na | 6492 | 0.090 | 0.5019 | No | ||
| 49 | AIFM1 | na | 6864 | 0.074 | 0.4834 | No | ||
| 50 | PPP3R1 | na | 7088 | 0.064 | 0.4729 | No | ||
| 51 | CASP7 | na | 7325 | 0.056 | 0.4614 | No | ||
| 52 | CSF2RB | na | 7784 | 0.038 | 0.4369 | No | ||
| 53 | NFKB1 | na | 7942 | 0.032 | 0.4290 | No | ||
| 54 | CHUK | na | 8229 | 0.021 | 0.4136 | No | ||
| 55 | FADD | na | 8656 | 0.004 | 0.3898 | No | ||
| 56 | AKT3 | na | 8815 | -0.002 | 0.3809 | No | ||
| 57 | IKBKG | na | 8880 | -0.005 | 0.3775 | No | ||
| 58 | TRADD | na | 9032 | -0.010 | 0.3693 | No | ||
| 59 | TNFRSF1A | na | 9206 | -0.016 | 0.3601 | No | ||
| 60 | AKT2 | na | 9366 | -0.023 | 0.3519 | No | ||
| 61 | DFFB | na | 9470 | -0.028 | 0.3470 | No | ||
| 62 | IRAK1 | na | 9570 | -0.031 | 0.3424 | No | ||
| 63 | BCL2L1 | na | 9830 | -0.041 | 0.3292 | No | ||
| 64 | CAPN1 | na | 9927 | -0.045 | 0.3252 | No | ||
| 65 | PPP3CB | na | 10137 | -0.053 | 0.3152 | No | ||
| 66 | TP53 | na | 10180 | -0.055 | 0.3146 | No | ||
| 67 | CASP3 | na | 10208 | -0.056 | 0.3149 | No | ||
| 68 | PPP3CC | na | 10264 | -0.058 | 0.3137 | No | ||
| 69 | ENDOD1 | na | 10697 | -0.076 | 0.2918 | No | ||
| 70 | IKBKB | na | 11156 | -0.095 | 0.2691 | No | ||
| 71 | BAD | na | 11806 | -0.125 | 0.2365 | No | ||
| 72 | AKT1 | na | 11942 | -0.131 | 0.2331 | No | ||
| 73 | BAX | na | 13533 | -0.225 | 0.1509 | No | ||
| 74 | TRAF2 | na | 13880 | -0.254 | 0.1396 | No | ||
| 75 | ENDOG | na | 14431 | -0.307 | 0.1185 | No | ||
| 76 | PRKX | na | 14524 | -0.317 | 0.1236 | No | ||
| 77 | IRAK3 | na | 15147 | -0.399 | 0.1014 | No | ||
| 78 | PIK3CD | na | 15172 | -0.403 | 0.1131 | No | ||
| 79 | BCL2 | na | 17215 | -1.151 | 0.0352 | No |