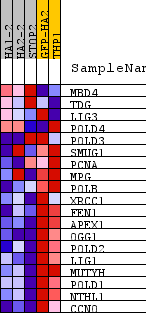
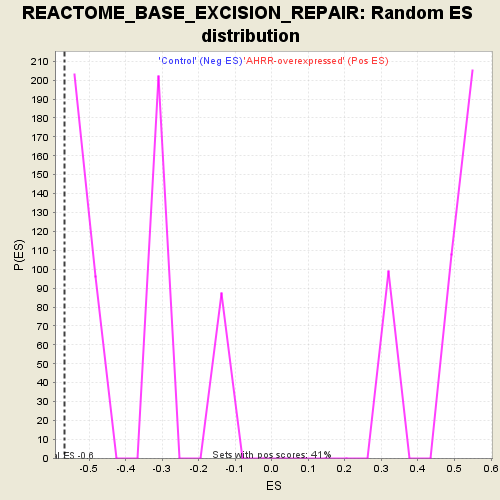
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA_2.class_2.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class_2.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | Control |
| GeneSet | REACTOME_BASE_EXCISION_REPAIR |
| Enrichment Score (ES) | -0.5670396 |
| Normalized Enrichment Score (NES) | -1.4134972 |
| Nominal p-value | 0.0 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MBD4 | na | 4244 | 0.221 | -0.1848 | No | ||
| 2 | TDG | na | 6198 | 0.103 | -0.2695 | No | ||
| 3 | LIG3 | na | 9635 | -0.034 | -0.4541 | No | ||
| 4 | POLD4 | na | 9774 | -0.039 | -0.4524 | No | ||
| 5 | POLD3 | na | 9942 | -0.046 | -0.4508 | No | ||
| 6 | SMUG1 | na | 10607 | -0.072 | -0.4708 | No | ||
| 7 | PCNA | na | 11127 | -0.093 | -0.4774 | No | ||
| 8 | MPG | na | 12421 | -0.155 | -0.5125 | No | ||
| 9 | POLB | na | 13394 | -0.215 | -0.5151 | Yes | ||
| 10 | XRCC1 | na | 13455 | -0.219 | -0.4656 | Yes | ||
| 11 | FEN1 | na | 13484 | -0.222 | -0.4137 | Yes | ||
| 12 | APEX1 | na | 13879 | -0.254 | -0.3744 | Yes | ||
| 13 | OGG1 | na | 13904 | -0.255 | -0.3142 | Yes | ||
| 14 | POLD2 | na | 14064 | -0.270 | -0.2578 | Yes | ||
| 15 | LIG1 | na | 14442 | -0.308 | -0.2044 | Yes | ||
| 16 | MUTYH | na | 14539 | -0.319 | -0.1328 | Yes | ||
| 17 | POLD1 | na | 14892 | -0.363 | -0.0649 | Yes | ||
| 18 | NTHL1 | na | 15259 | -0.417 | 0.0153 | Yes | ||
| 19 | CCNO | na | 15882 | -0.536 | 0.1099 | Yes |