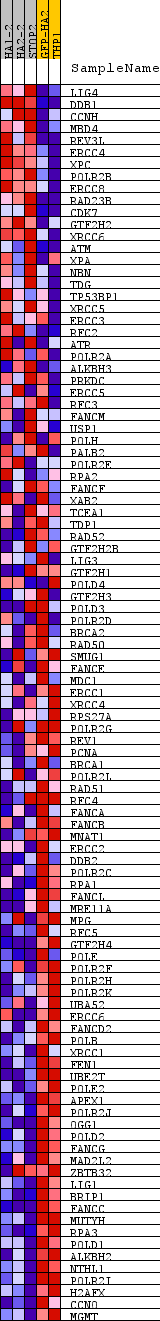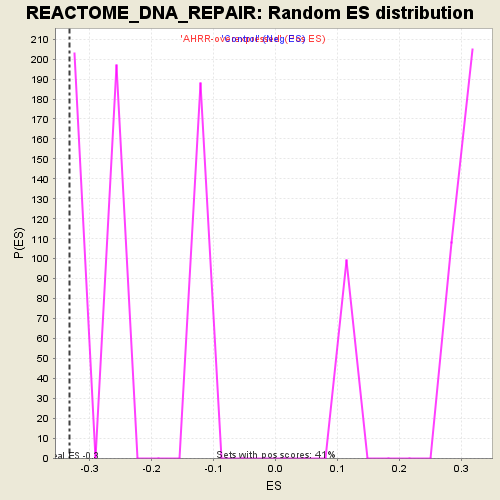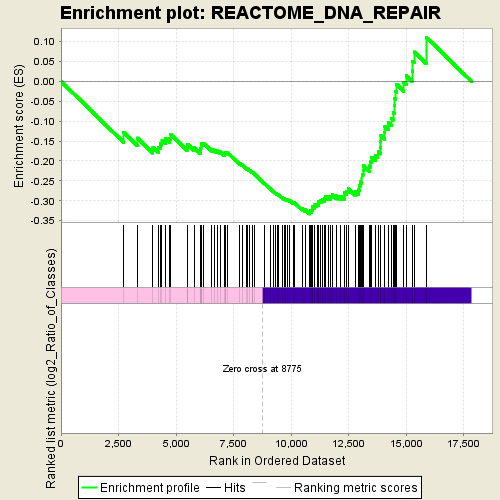
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA_2.class_2.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class_2.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | Control |
| GeneSet | REACTOME_DNA_REPAIR |
| Enrichment Score (ES) | -0.33330128 |
| Normalized Enrichment Score (NES) | -1.3619545 |
| Nominal p-value | 0.18197279 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | LIG4 | na | 2724 | 0.408 | -0.1283 | No | ||
| 2 | DDB1 | na | 3316 | 0.314 | -0.1422 | No | ||
| 3 | CCNH | na | 3991 | 0.243 | -0.1652 | No | ||
| 4 | MBD4 | na | 4244 | 0.221 | -0.1658 | No | ||
| 5 | REV3L | na | 4312 | 0.216 | -0.1562 | No | ||
| 6 | ERCC4 | na | 4386 | 0.210 | -0.1474 | No | ||
| 7 | XPC | na | 4538 | 0.198 | -0.1437 | No | ||
| 8 | POLR2B | na | 4728 | 0.185 | -0.1429 | No | ||
| 9 | ERCC8 | na | 4772 | 0.182 | -0.1341 | No | ||
| 10 | RAD23B | na | 5477 | 0.139 | -0.1652 | No | ||
| 11 | CDK7 | na | 5509 | 0.138 | -0.1584 | No | ||
| 12 | GTF2H2 | na | 5784 | 0.124 | -0.1662 | No | ||
| 13 | XRCC6 | na | 6046 | 0.110 | -0.1741 | No | ||
| 14 | ATM | na | 6065 | 0.109 | -0.1684 | No | ||
| 15 | XPA | na | 6085 | 0.108 | -0.1627 | No | ||
| 16 | NBN | na | 6087 | 0.108 | -0.1561 | No | ||
| 17 | TDG | na | 6198 | 0.103 | -0.1559 | No | ||
| 18 | TP53BP1 | na | 6554 | 0.088 | -0.1705 | No | ||
| 19 | XRCC5 | na | 6678 | 0.082 | -0.1724 | No | ||
| 20 | ERCC3 | na | 6797 | 0.076 | -0.1743 | No | ||
| 21 | RFC2 | na | 6913 | 0.071 | -0.1764 | No | ||
| 22 | ATR | na | 7102 | 0.064 | -0.1830 | No | ||
| 23 | POLR2A | na | 7130 | 0.063 | -0.1806 | No | ||
| 24 | ALKBH3 | na | 7163 | 0.062 | -0.1786 | No | ||
| 25 | PRKDC | na | 7220 | 0.059 | -0.1781 | No | ||
| 26 | ERCC5 | na | 7756 | 0.038 | -0.2059 | No | ||
| 27 | RFC3 | na | 7865 | 0.034 | -0.2099 | No | ||
| 28 | FANCM | na | 8057 | 0.027 | -0.2190 | No | ||
| 29 | USP1 | na | 8099 | 0.026 | -0.2197 | No | ||
| 30 | POLH | na | 8184 | 0.023 | -0.2230 | No | ||
| 31 | PALB2 | na | 8303 | 0.018 | -0.2286 | No | ||
| 32 | POLR2E | na | 8326 | 0.017 | -0.2288 | No | ||
| 33 | RPA2 | na | 8398 | 0.014 | -0.2319 | No | ||
| 34 | FANCF | na | 8839 | -0.003 | -0.2565 | No | ||
| 35 | XAB2 | na | 9089 | -0.012 | -0.2698 | No | ||
| 36 | TCEA1 | na | 9227 | -0.017 | -0.2765 | No | ||
| 37 | TDP1 | na | 9333 | -0.022 | -0.2811 | No | ||
| 38 | RAD52 | na | 9400 | -0.025 | -0.2833 | No | ||
| 39 | GTF2H2B | na | 9452 | -0.027 | -0.2845 | No | ||
| 40 | LIG3 | na | 9635 | -0.034 | -0.2927 | No | ||
| 41 | GTF2H1 | na | 9692 | -0.036 | -0.2936 | No | ||
| 42 | POLD4 | na | 9774 | -0.039 | -0.2958 | No | ||
| 43 | GTF2H3 | na | 9849 | -0.042 | -0.2974 | No | ||
| 44 | POLD3 | na | 9942 | -0.046 | -0.2997 | No | ||
| 45 | POLR2D | na | 10089 | -0.052 | -0.3048 | No | ||
| 46 | BRCA2 | na | 10139 | -0.054 | -0.3042 | No | ||
| 47 | RAD50 | na | 10480 | -0.066 | -0.3193 | No | ||
| 48 | SMUG1 | na | 10607 | -0.072 | -0.3220 | No | ||
| 49 | FANCE | na | 10809 | -0.080 | -0.3284 | Yes | ||
| 50 | MDC1 | na | 10822 | -0.081 | -0.3240 | Yes | ||
| 51 | ERCC1 | na | 10904 | -0.084 | -0.3234 | Yes | ||
| 52 | XRCC4 | na | 10931 | -0.085 | -0.3196 | Yes | ||
| 53 | RPS27A | na | 10937 | -0.086 | -0.3146 | Yes | ||
| 54 | POLR2G | na | 11033 | -0.089 | -0.3145 | Yes | ||
| 55 | REV1 | na | 11037 | -0.089 | -0.3091 | Yes | ||
| 56 | PCNA | na | 11127 | -0.093 | -0.3084 | Yes | ||
| 57 | BRCA1 | na | 11195 | -0.096 | -0.3062 | Yes | ||
| 58 | POLR2L | na | 11209 | -0.097 | -0.3010 | Yes | ||
| 59 | RAD51 | na | 11291 | -0.100 | -0.2994 | Yes | ||
| 60 | RFC4 | na | 11359 | -0.104 | -0.2968 | Yes | ||
| 61 | FANCA | na | 11436 | -0.107 | -0.2944 | Yes | ||
| 62 | FANCB | na | 11477 | -0.109 | -0.2899 | Yes | ||
| 63 | MNAT1 | na | 11604 | -0.115 | -0.2899 | Yes | ||
| 64 | ERCC2 | na | 11729 | -0.121 | -0.2895 | Yes | ||
| 65 | DDB2 | na | 11783 | -0.123 | -0.2848 | Yes | ||
| 66 | POLR2C | na | 11975 | -0.132 | -0.2875 | Yes | ||
| 67 | RPA1 | na | 12156 | -0.142 | -0.2889 | Yes | ||
| 68 | FANCL | na | 12306 | -0.149 | -0.2881 | Yes | ||
| 69 | MRE11A | na | 12309 | -0.149 | -0.2789 | Yes | ||
| 70 | MPG | na | 12421 | -0.155 | -0.2756 | Yes | ||
| 71 | RFC5 | na | 12490 | -0.160 | -0.2696 | Yes | ||
| 72 | GTF2H4 | na | 12801 | -0.177 | -0.2761 | Yes | ||
| 73 | POLE | na | 12946 | -0.186 | -0.2727 | Yes | ||
| 74 | POLR2F | na | 12959 | -0.187 | -0.2619 | Yes | ||
| 75 | POLR2H | na | 13002 | -0.190 | -0.2525 | Yes | ||
| 76 | POLR2K | na | 13081 | -0.195 | -0.2448 | Yes | ||
| 77 | UBA52 | na | 13082 | -0.195 | -0.2328 | Yes | ||
| 78 | ERCC6 | na | 13129 | -0.198 | -0.2231 | Yes | ||
| 79 | FANCD2 | na | 13141 | -0.199 | -0.2115 | Yes | ||
| 80 | POLB | na | 13394 | -0.215 | -0.2124 | Yes | ||
| 81 | XRCC1 | na | 13455 | -0.219 | -0.2022 | Yes | ||
| 82 | FEN1 | na | 13484 | -0.222 | -0.1901 | Yes | ||
| 83 | UBE2T | na | 13660 | -0.236 | -0.1854 | Yes | ||
| 84 | POLE2 | na | 13778 | -0.246 | -0.1768 | Yes | ||
| 85 | APEX1 | na | 13879 | -0.254 | -0.1668 | Yes | ||
| 86 | POLR2J | na | 13891 | -0.254 | -0.1517 | Yes | ||
| 87 | OGG1 | na | 13904 | -0.255 | -0.1366 | Yes | ||
| 88 | POLD2 | na | 14064 | -0.270 | -0.1289 | Yes | ||
| 89 | FANCG | na | 14077 | -0.271 | -0.1128 | Yes | ||
| 90 | MAD2L2 | na | 14235 | -0.284 | -0.1041 | Yes | ||
| 91 | ZBTB32 | na | 14362 | -0.300 | -0.0926 | Yes | ||
| 92 | LIG1 | na | 14442 | -0.308 | -0.0780 | Yes | ||
| 93 | BRIP1 | na | 14483 | -0.312 | -0.0610 | Yes | ||
| 94 | FANCC | na | 14491 | -0.313 | -0.0421 | Yes | ||
| 95 | MUTYH | na | 14539 | -0.319 | -0.0250 | Yes | ||
| 96 | RPA3 | na | 14591 | -0.326 | -0.0078 | Yes | ||
| 97 | POLD1 | na | 14892 | -0.363 | -0.0023 | Yes | ||
| 98 | ALKBH2 | na | 15006 | -0.378 | 0.0147 | Yes | ||
| 99 | NTHL1 | na | 15259 | -0.417 | 0.0263 | Yes | ||
| 100 | POLR2I | na | 15287 | -0.421 | 0.0508 | Yes | ||
| 101 | H2AFX | na | 15352 | -0.433 | 0.0739 | Yes | ||
| 102 | CCNO | na | 15882 | -0.536 | 0.0772 | Yes | ||
| 103 | MGMT | na | 15883 | -0.536 | 0.1104 | Yes |