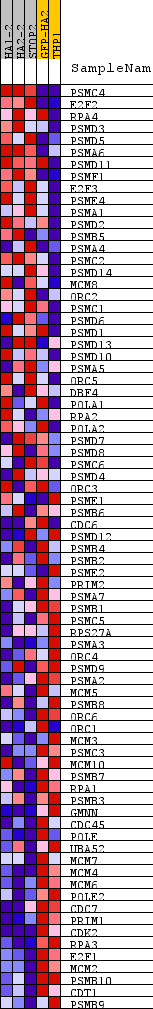
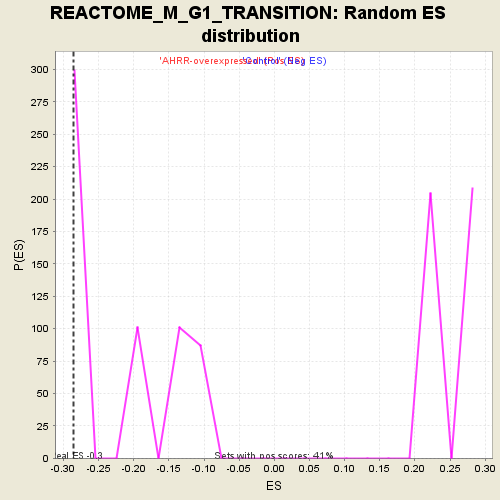
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA_2.class_2.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class_2.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | Control |
| GeneSet | REACTOME_M_G1_TRANSITION |
| Enrichment Score (ES) | -0.28616276 |
| Normalized Enrichment Score (NES) | -1.283857 |
| Nominal p-value | 0.3452381 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PSMC4 | na | 3303 | 0.317 | -0.1542 | No | ||
| 2 | E2F2 | na | 3773 | 0.263 | -0.1543 | No | ||
| 3 | RPA4 | na | 4101 | 0.233 | -0.1494 | No | ||
| 4 | PSMD3 | na | 4285 | 0.218 | -0.1378 | No | ||
| 5 | PSMD5 | na | 4354 | 0.213 | -0.1204 | No | ||
| 6 | PSMA6 | na | 5023 | 0.164 | -0.1415 | No | ||
| 7 | PSMD11 | na | 5771 | 0.125 | -0.1711 | No | ||
| 8 | PSMF1 | na | 5807 | 0.123 | -0.1608 | No | ||
| 9 | E2F3 | na | 6158 | 0.105 | -0.1700 | No | ||
| 10 | PSME4 | na | 6203 | 0.103 | -0.1621 | No | ||
| 11 | PSMA1 | na | 6253 | 0.101 | -0.1548 | No | ||
| 12 | PSMD2 | na | 6562 | 0.087 | -0.1634 | No | ||
| 13 | PSMB5 | na | 6774 | 0.078 | -0.1675 | No | ||
| 14 | PSMA4 | na | 6778 | 0.078 | -0.1599 | No | ||
| 15 | PSMC2 | na | 7024 | 0.067 | -0.1670 | No | ||
| 16 | PSMD14 | na | 7127 | 0.063 | -0.1664 | No | ||
| 17 | MCM8 | na | 7144 | 0.062 | -0.1610 | No | ||
| 18 | ORC2 | na | 7216 | 0.059 | -0.1591 | No | ||
| 19 | PSMC1 | na | 7256 | 0.058 | -0.1555 | No | ||
| 20 | PSMD6 | na | 7286 | 0.057 | -0.1514 | No | ||
| 21 | PSMD1 | na | 7413 | 0.052 | -0.1533 | No | ||
| 22 | PSMD13 | na | 7900 | 0.033 | -0.1773 | No | ||
| 23 | PSMD10 | na | 7967 | 0.031 | -0.1780 | No | ||
| 24 | PSMA5 | na | 8336 | 0.017 | -0.1971 | No | ||
| 25 | ORC5 | na | 8364 | 0.015 | -0.1970 | No | ||
| 26 | DBF4 | na | 8392 | 0.014 | -0.1971 | No | ||
| 27 | POLA1 | na | 8394 | 0.014 | -0.1958 | No | ||
| 28 | RPA2 | na | 8398 | 0.014 | -0.1945 | No | ||
| 29 | POLA2 | na | 8491 | 0.011 | -0.1987 | No | ||
| 30 | PSMD7 | na | 8515 | 0.009 | -0.1990 | No | ||
| 31 | PSMD8 | na | 8705 | 0.002 | -0.2094 | No | ||
| 32 | PSMC6 | na | 8715 | 0.002 | -0.2097 | No | ||
| 33 | PSMD4 | na | 8737 | 0.001 | -0.2108 | No | ||
| 34 | ORC3 | na | 8813 | -0.002 | -0.2148 | No | ||
| 35 | PSME1 | na | 9304 | -0.020 | -0.2404 | No | ||
| 36 | PSMB6 | na | 9385 | -0.024 | -0.2425 | No | ||
| 37 | CDC6 | na | 9800 | -0.040 | -0.2618 | No | ||
| 38 | PSMD12 | na | 9918 | -0.044 | -0.2639 | No | ||
| 39 | PSMB4 | na | 10214 | -0.056 | -0.2749 | No | ||
| 40 | PSMB2 | na | 10326 | -0.060 | -0.2751 | No | ||
| 41 | PSME2 | na | 10524 | -0.068 | -0.2793 | Yes | ||
| 42 | PRIM2 | na | 10559 | -0.070 | -0.2743 | Yes | ||
| 43 | PSMA7 | na | 10589 | -0.071 | -0.2688 | Yes | ||
| 44 | PSMB1 | na | 10605 | -0.071 | -0.2625 | Yes | ||
| 45 | PSMC5 | na | 10658 | -0.074 | -0.2580 | Yes | ||
| 46 | RPS27A | na | 10937 | -0.086 | -0.2651 | Yes | ||
| 47 | PSMA3 | na | 10972 | -0.087 | -0.2583 | Yes | ||
| 48 | ORC4 | na | 11403 | -0.106 | -0.2719 | Yes | ||
| 49 | PSMD9 | na | 11564 | -0.113 | -0.2696 | Yes | ||
| 50 | PSMA2 | na | 11684 | -0.119 | -0.2644 | Yes | ||
| 51 | MCM5 | na | 11718 | -0.120 | -0.2542 | Yes | ||
| 52 | PSMB8 | na | 11730 | -0.121 | -0.2427 | Yes | ||
| 53 | ORC6 | na | 11735 | -0.121 | -0.2308 | Yes | ||
| 54 | ORC1 | na | 11761 | -0.122 | -0.2200 | Yes | ||
| 55 | MCM3 | na | 11801 | -0.124 | -0.2098 | Yes | ||
| 56 | PSMC3 | na | 12055 | -0.136 | -0.2104 | Yes | ||
| 57 | MCM10 | na | 12056 | -0.136 | -0.1968 | Yes | ||
| 58 | PSMB7 | na | 12106 | -0.138 | -0.1857 | Yes | ||
| 59 | RPA1 | na | 12156 | -0.142 | -0.1743 | Yes | ||
| 60 | PSMB3 | na | 12273 | -0.148 | -0.1660 | Yes | ||
| 61 | GMNN | na | 12339 | -0.151 | -0.1546 | Yes | ||
| 62 | CDC45 | na | 12880 | -0.182 | -0.1668 | Yes | ||
| 63 | POLE | na | 12946 | -0.186 | -0.1518 | Yes | ||
| 64 | UBA52 | na | 13082 | -0.195 | -0.1398 | Yes | ||
| 65 | MCM7 | na | 13111 | -0.197 | -0.1217 | Yes | ||
| 66 | MCM4 | na | 13591 | -0.230 | -0.1256 | Yes | ||
| 67 | MCM6 | na | 13731 | -0.241 | -0.1093 | Yes | ||
| 68 | POLE2 | na | 13778 | -0.246 | -0.0873 | Yes | ||
| 69 | CDC7 | na | 14050 | -0.269 | -0.0756 | Yes | ||
| 70 | PRIM1 | na | 14247 | -0.287 | -0.0580 | Yes | ||
| 71 | CDK2 | na | 14296 | -0.292 | -0.0314 | Yes | ||
| 72 | RPA3 | na | 14591 | -0.326 | -0.0153 | Yes | ||
| 73 | E2F1 | na | 14772 | -0.348 | 0.0093 | Yes | ||
| 74 | MCM2 | na | 15059 | -0.385 | 0.0318 | Yes | ||
| 75 | PSMB10 | na | 15077 | -0.388 | 0.0698 | Yes | ||
| 76 | CDT1 | na | 15105 | -0.393 | 0.1076 | Yes | ||
| 77 | PSMB9 | na | 15528 | -0.462 | 0.1302 | Yes |