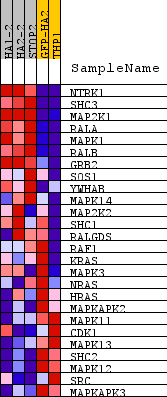
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA_2.class_2.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class_2.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | AHRR-overexpressed |
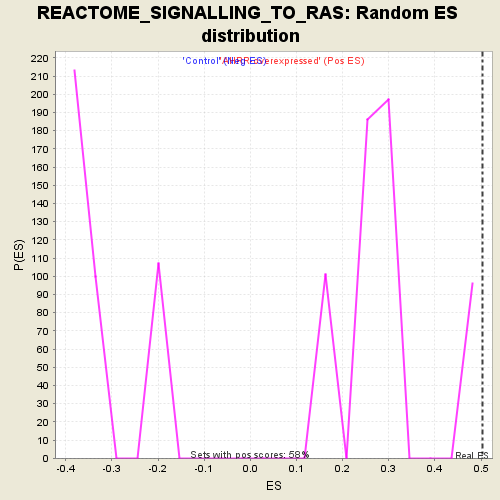
| GeneSet | REACTOME_SIGNALLING_TO_RAS |
| Enrichment Score (ES) | 0.503277 |
| Normalized Enrichment Score (NES) | 1.7332851 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.2771402 |
| FWER p-Value | 0.383 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | NTRK1 | na | 131 | 2.985 | 0.2898 | Yes | ||
| 2 | SHC3 | na | 246 | 2.209 | 0.5033 | Yes | ||
| 3 | MAP2K1 | na | 1559 | 0.713 | 0.5006 | No | ||
| 4 | RALA | na | 4176 | 0.227 | 0.3764 | No | ||
| 5 | MAPK1 | na | 4266 | 0.219 | 0.3932 | No | ||
| 6 | RALB | na | 4929 | 0.170 | 0.3729 | No | ||
| 7 | GRB2 | na | 4985 | 0.166 | 0.3864 | No | ||
| 8 | SOS1 | na | 5811 | 0.122 | 0.3523 | No | ||
| 9 | YWHAB | na | 6221 | 0.102 | 0.3395 | No | ||
| 10 | MAPK14 | na | 6344 | 0.097 | 0.3423 | No | ||
| 11 | MAP2K2 | na | 6878 | 0.073 | 0.3197 | No | ||
| 12 | SHC1 | na | 7644 | 0.042 | 0.2809 | No | ||
| 13 | RALGDS | na | 7841 | 0.035 | 0.2734 | No | ||
| 14 | RAF1 | na | 8761 | 0.000 | 0.2218 | No | ||
| 15 | KRAS | na | 9191 | -0.016 | 0.1993 | No | ||
| 16 | MAPK3 | na | 9820 | -0.041 | 0.1681 | No | ||
| 17 | NRAS | na | 9998 | -0.048 | 0.1630 | No | ||
| 18 | HRAS | na | 11793 | -0.124 | 0.0746 | No | ||
| 19 | MAPKAPK2 | na | 12125 | -0.140 | 0.0699 | No | ||
| 20 | MAPK11 | na | 12162 | -0.142 | 0.0820 | No | ||
| 21 | CDK1 | na | 13099 | -0.196 | 0.0490 | No | ||
| 22 | MAPK13 | na | 15066 | -0.387 | -0.0228 | No | ||
| 23 | SHC2 | na | 15132 | -0.397 | 0.0130 | No | ||
| 24 | MAPK12 | na | 15148 | -0.400 | 0.0520 | No | ||
| 25 | SRC | na | 15318 | -0.427 | 0.0850 | No | ||
| 26 | MAPKAPK3 | na | 16017 | -0.568 | 0.1024 | No |