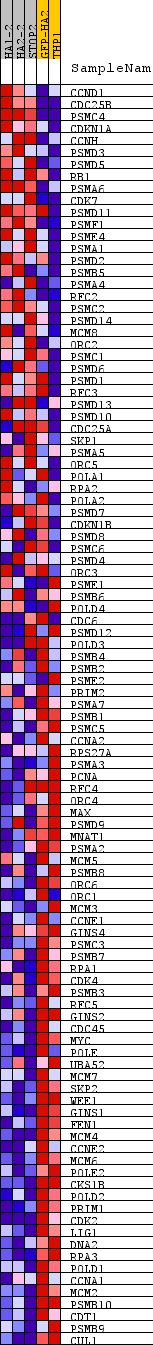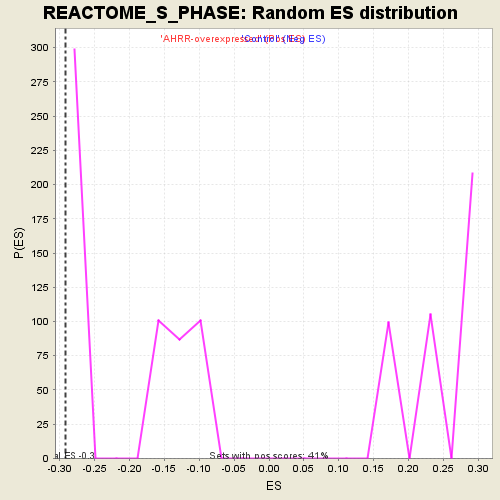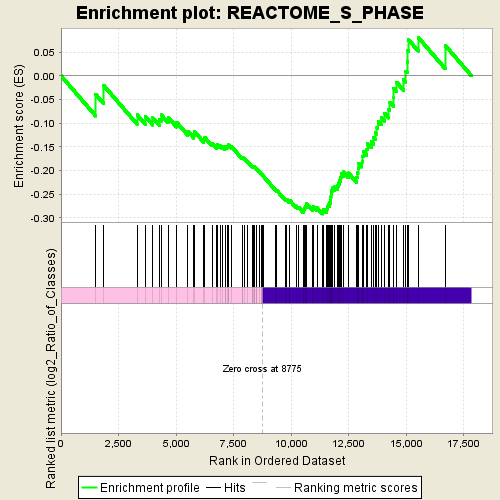
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA_2.class_2.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class_2.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | Control |
| GeneSet | REACTOME_S_PHASE |
| Enrichment Score (ES) | -0.2921359 |
| Normalized Enrichment Score (NES) | -1.3964175 |
| Nominal p-value | 0.1632653 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CCND1 | na | 1509 | 0.732 | -0.0395 | No | ||
| 2 | CDC25B | na | 1846 | 0.618 | -0.0200 | No | ||
| 3 | PSMC4 | na | 3303 | 0.317 | -0.0824 | No | ||
| 4 | CDKN1A | na | 3668 | 0.274 | -0.0859 | No | ||
| 5 | CCNH | na | 3991 | 0.243 | -0.0890 | No | ||
| 6 | PSMD3 | na | 4285 | 0.218 | -0.0919 | No | ||
| 7 | PSMD5 | na | 4354 | 0.213 | -0.0825 | No | ||
| 8 | RB1 | na | 4656 | 0.189 | -0.0877 | No | ||
| 9 | PSMA6 | na | 5023 | 0.164 | -0.0981 | No | ||
| 10 | CDK7 | na | 5509 | 0.138 | -0.1169 | No | ||
| 11 | PSMD11 | na | 5771 | 0.125 | -0.1239 | No | ||
| 12 | PSMF1 | na | 5807 | 0.123 | -0.1182 | No | ||
| 13 | PSME4 | na | 6203 | 0.103 | -0.1341 | No | ||
| 14 | PSMA1 | na | 6253 | 0.101 | -0.1306 | No | ||
| 15 | PSMD2 | na | 6562 | 0.087 | -0.1425 | No | ||
| 16 | PSMB5 | na | 6774 | 0.078 | -0.1496 | No | ||
| 17 | PSMA4 | na | 6778 | 0.078 | -0.1449 | No | ||
| 18 | RFC2 | na | 6913 | 0.071 | -0.1480 | No | ||
| 19 | PSMC2 | na | 7024 | 0.067 | -0.1501 | No | ||
| 20 | PSMD14 | na | 7127 | 0.063 | -0.1519 | No | ||
| 21 | MCM8 | na | 7144 | 0.062 | -0.1489 | No | ||
| 22 | ORC2 | na | 7216 | 0.059 | -0.1492 | No | ||
| 23 | PSMC1 | na | 7256 | 0.058 | -0.1478 | No | ||
| 24 | PSMD6 | na | 7286 | 0.057 | -0.1459 | No | ||
| 25 | PSMD1 | na | 7413 | 0.052 | -0.1498 | No | ||
| 26 | RFC3 | na | 7865 | 0.034 | -0.1731 | No | ||
| 27 | PSMD13 | na | 7900 | 0.033 | -0.1730 | No | ||
| 28 | PSMD10 | na | 7967 | 0.031 | -0.1748 | No | ||
| 29 | CDC25A | na | 8116 | 0.025 | -0.1816 | No | ||
| 30 | SKP1 | na | 8319 | 0.017 | -0.1919 | No | ||
| 31 | PSMA5 | na | 8336 | 0.017 | -0.1918 | No | ||
| 32 | ORC5 | na | 8364 | 0.015 | -0.1923 | No | ||
| 33 | POLA1 | na | 8394 | 0.014 | -0.1931 | No | ||
| 34 | RPA2 | na | 8398 | 0.014 | -0.1924 | No | ||
| 35 | POLA2 | na | 8491 | 0.011 | -0.1969 | No | ||
| 36 | PSMD7 | na | 8515 | 0.009 | -0.1976 | No | ||
| 37 | CDKN1B | na | 8605 | 0.006 | -0.2023 | No | ||
| 38 | PSMD8 | na | 8705 | 0.002 | -0.2077 | No | ||
| 39 | PSMC6 | na | 8715 | 0.002 | -0.2081 | No | ||
| 40 | PSMD4 | na | 8737 | 0.001 | -0.2092 | No | ||
| 41 | ORC3 | na | 8813 | -0.002 | -0.2133 | No | ||
| 42 | PSME1 | na | 9304 | -0.020 | -0.2397 | No | ||
| 43 | PSMB6 | na | 9385 | -0.024 | -0.2427 | No | ||
| 44 | POLD4 | na | 9774 | -0.039 | -0.2622 | No | ||
| 45 | CDC6 | na | 9800 | -0.040 | -0.2611 | No | ||
| 46 | PSMD12 | na | 9918 | -0.044 | -0.2649 | No | ||
| 47 | POLD3 | na | 9942 | -0.046 | -0.2634 | No | ||
| 48 | PSMB4 | na | 10214 | -0.056 | -0.2752 | No | ||
| 49 | PSMB2 | na | 10326 | -0.060 | -0.2777 | No | ||
| 50 | PSME2 | na | 10524 | -0.068 | -0.2845 | No | ||
| 51 | PRIM2 | na | 10559 | -0.070 | -0.2821 | No | ||
| 52 | PSMA7 | na | 10589 | -0.071 | -0.2793 | No | ||
| 53 | PSMB1 | na | 10605 | -0.071 | -0.2757 | No | ||
| 54 | PSMC5 | na | 10658 | -0.074 | -0.2741 | No | ||
| 55 | CCNA2 | na | 10675 | -0.075 | -0.2703 | No | ||
| 56 | RPS27A | na | 10937 | -0.086 | -0.2797 | No | ||
| 57 | PSMA3 | na | 10972 | -0.087 | -0.2762 | No | ||
| 58 | PCNA | na | 11127 | -0.093 | -0.2791 | No | ||
| 59 | RFC4 | na | 11359 | -0.104 | -0.2857 | Yes | ||
| 60 | ORC4 | na | 11403 | -0.106 | -0.2815 | Yes | ||
| 61 | MAX | na | 11546 | -0.112 | -0.2826 | Yes | ||
| 62 | PSMD9 | na | 11564 | -0.113 | -0.2765 | Yes | ||
| 63 | MNAT1 | na | 11604 | -0.115 | -0.2715 | Yes | ||
| 64 | PSMA2 | na | 11684 | -0.119 | -0.2686 | Yes | ||
| 65 | MCM5 | na | 11718 | -0.120 | -0.2630 | Yes | ||
| 66 | PSMB8 | na | 11730 | -0.121 | -0.2561 | Yes | ||
| 67 | ORC6 | na | 11735 | -0.121 | -0.2488 | Yes | ||
| 68 | ORC1 | na | 11761 | -0.122 | -0.2426 | Yes | ||
| 69 | MCM3 | na | 11801 | -0.124 | -0.2371 | Yes | ||
| 70 | CCNE1 | na | 11903 | -0.129 | -0.2348 | Yes | ||
| 71 | GINS4 | na | 12010 | -0.134 | -0.2324 | Yes | ||
| 72 | PSMC3 | na | 12055 | -0.136 | -0.2265 | Yes | ||
| 73 | PSMB7 | na | 12106 | -0.138 | -0.2207 | Yes | ||
| 74 | RPA1 | na | 12156 | -0.142 | -0.2146 | Yes | ||
| 75 | CDK4 | na | 12180 | -0.143 | -0.2070 | Yes | ||
| 76 | PSMB3 | na | 12273 | -0.148 | -0.2030 | Yes | ||
| 77 | RFC5 | na | 12490 | -0.160 | -0.2053 | Yes | ||
| 78 | GINS2 | na | 12855 | -0.180 | -0.2146 | Yes | ||
| 79 | CDC45 | na | 12880 | -0.182 | -0.2046 | Yes | ||
| 80 | MYC | na | 12909 | -0.184 | -0.1948 | Yes | ||
| 81 | POLE | na | 12946 | -0.186 | -0.1852 | Yes | ||
| 82 | UBA52 | na | 13082 | -0.195 | -0.1807 | Yes | ||
| 83 | MCM7 | na | 13111 | -0.197 | -0.1700 | Yes | ||
| 84 | SKP2 | na | 13137 | -0.199 | -0.1591 | Yes | ||
| 85 | WEE1 | na | 13293 | -0.209 | -0.1548 | Yes | ||
| 86 | GINS1 | na | 13321 | -0.211 | -0.1432 | Yes | ||
| 87 | FEN1 | na | 13484 | -0.222 | -0.1386 | Yes | ||
| 88 | MCM4 | na | 13591 | -0.230 | -0.1303 | Yes | ||
| 89 | CCNE2 | na | 13664 | -0.236 | -0.1197 | Yes | ||
| 90 | MCM6 | na | 13731 | -0.241 | -0.1084 | Yes | ||
| 91 | POLE2 | na | 13778 | -0.246 | -0.0957 | Yes | ||
| 92 | CKS1B | na | 13917 | -0.256 | -0.0876 | Yes | ||
| 93 | POLD2 | na | 14064 | -0.270 | -0.0790 | Yes | ||
| 94 | PRIM1 | na | 14247 | -0.287 | -0.0714 | Yes | ||
| 95 | CDK2 | na | 14296 | -0.292 | -0.0560 | Yes | ||
| 96 | LIG1 | na | 14442 | -0.308 | -0.0449 | Yes | ||
| 97 | DNA2 | na | 14447 | -0.309 | -0.0259 | Yes | ||
| 98 | RPA3 | na | 14591 | -0.326 | -0.0138 | Yes | ||
| 99 | POLD1 | na | 14892 | -0.363 | -0.0081 | Yes | ||
| 100 | CCNA1 | na | 14986 | -0.376 | 0.0101 | Yes | ||
| 101 | MCM2 | na | 15059 | -0.385 | 0.0300 | Yes | ||
| 102 | PSMB10 | na | 15077 | -0.388 | 0.0532 | Yes | ||
| 103 | CDT1 | na | 15105 | -0.393 | 0.0761 | Yes | ||
| 104 | PSMB9 | na | 15528 | -0.462 | 0.0811 | Yes | ||
| 105 | CUL1 | na | 16697 | -0.792 | 0.0645 | Yes |