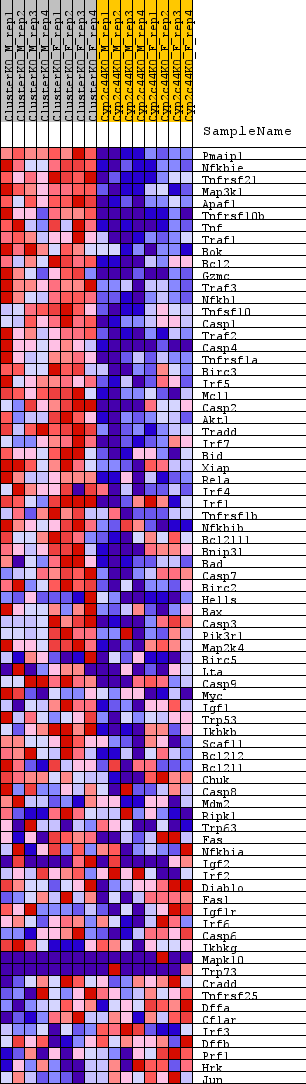
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___ClusterKO_vs_Cyp2c44KO.RPKM_matrix___ClusterKO_vs_Cyp2c44KO.cls #ClusterKO_versus_Cyp2c44KO |
| Phenotype | RPKM_matrix___ClusterKO_vs_Cyp2c44KO.cls#ClusterKO_versus_Cyp2c44KO |
| Upregulated in class | ClusterKO |
| GeneSet | WP_APOPTOSIS |
| Enrichment Score (ES) | 0.72375846 |
| Normalized Enrichment Score (NES) | 1.4233289 |
| Nominal p-value | 0.004016064 |
| FDR q-value | 0.067803554 |
| FWER p-Value | 0.325 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Pmaip1 | na | 75 | 2.076 | 0.0460 | Yes |
| 2 | Nfkbie | na | 139 | 1.867 | 0.0874 | Yes |
| 3 | Tnfrsf21 | na | 380 | 1.552 | 0.1184 | Yes |
| 4 | Map3k1 | na | 384 | 1.543 | 0.1535 | Yes |
| 5 | Apaf1 | na | 542 | 1.420 | 0.1831 | Yes |
| 6 | Tnfrsf10b | na | 549 | 1.414 | 0.2151 | Yes |
| 7 | Tnf | na | 682 | 1.332 | 0.2431 | Yes |
| 8 | Traf1 | na | 959 | 1.189 | 0.2653 | Yes |
| 9 | Bok | na | 1087 | 1.138 | 0.2890 | Yes |
| 10 | Bcl2 | na | 1285 | 1.071 | 0.3098 | Yes |
| 11 | Gzmc | na | 1334 | 1.052 | 0.3330 | Yes |
| 12 | Traf3 | na | 1374 | 1.040 | 0.3559 | Yes |
| 13 | Nfkb1 | na | 1528 | 0.998 | 0.3760 | Yes |
| 14 | Tnfsf10 | na | 1561 | 0.990 | 0.3979 | Yes |
| 15 | Casp1 | na | 1775 | 0.933 | 0.4154 | Yes |
| 16 | Traf2 | na | 1852 | 0.913 | 0.4348 | Yes |
| 17 | Casp4 | na | 2011 | 0.879 | 0.4520 | Yes |
| 18 | Tnfrsf1a | na | 2061 | 0.867 | 0.4709 | Yes |
| 19 | Birc3 | na | 2142 | 0.852 | 0.4889 | Yes |
| 20 | Irf5 | na | 2197 | 0.841 | 0.5071 | Yes |
| 21 | Mcl1 | na | 2380 | 0.809 | 0.5223 | Yes |
| 22 | Casp2 | na | 2428 | 0.801 | 0.5397 | Yes |
| 23 | Akt1 | na | 2566 | 0.775 | 0.5549 | Yes |
| 24 | Tradd | na | 2970 | 0.716 | 0.5641 | Yes |
| 25 | Irf7 | na | 3523 | 0.653 | 0.5691 | Yes |
| 26 | Bid | na | 3709 | 0.637 | 0.5803 | Yes |
| 27 | Xiap | na | 3887 | 0.622 | 0.5914 | Yes |
| 28 | Rela | na | 3986 | 0.612 | 0.6036 | Yes |
| 29 | Irf4 | na | 4015 | 0.609 | 0.6169 | Yes |
| 30 | Irf1 | na | 4085 | 0.603 | 0.6294 | Yes |
| 31 | Tnfrsf1b | na | 4185 | 0.594 | 0.6412 | Yes |
| 32 | Nfkbib | na | 4690 | 0.555 | 0.6449 | Yes |
| 33 | Bcl2l11 | na | 4885 | 0.541 | 0.6538 | Yes |
| 34 | Bnip3l | na | 4937 | 0.537 | 0.6651 | Yes |
| 35 | Bad | na | 4961 | 0.535 | 0.6769 | Yes |
| 36 | Casp7 | na | 5835 | 0.475 | 0.6722 | Yes |
| 37 | Birc2 | na | 6158 | 0.456 | 0.6768 | Yes |
| 38 | Hells | na | 6737 | 0.425 | 0.6763 | Yes |
| 39 | Bax | na | 6902 | 0.416 | 0.6828 | Yes |
| 40 | Casp3 | na | 6964 | 0.413 | 0.6911 | Yes |
| 41 | Pik3r1 | na | 7254 | 0.399 | 0.6951 | Yes |
| 42 | Map2k4 | na | 7410 | 0.392 | 0.7013 | Yes |
| 43 | Birc5 | na | 7612 | 0.383 | 0.7064 | Yes |
| 44 | Lta | na | 7932 | 0.367 | 0.7091 | Yes |
| 45 | Casp9 | na | 8122 | 0.359 | 0.7139 | Yes |
| 46 | Myc | na | 8354 | 0.347 | 0.7177 | Yes |
| 47 | Igf1 | na | 8454 | 0.342 | 0.7238 | Yes |
| 48 | Trp53 | na | 8901 | 0.321 | 0.7232 | No |
| 49 | Ikbkb | na | 9635 | 0.289 | 0.7167 | No |
| 50 | Scaf11 | na | 10372 | 0.257 | 0.7095 | No |
| 51 | Bcl2l2 | na | 10672 | 0.244 | 0.7098 | No |
| 52 | Bcl2l1 | na | 10756 | 0.241 | 0.7138 | No |
| 53 | Chuk | na | 11423 | 0.215 | 0.7068 | No |
| 54 | Casp8 | na | 11424 | 0.215 | 0.7117 | No |
| 55 | Mdm2 | na | 11704 | 0.205 | 0.7115 | No |
| 56 | Ripk1 | na | 11998 | 0.194 | 0.7107 | No |
| 57 | Trp63 | na | 12444 | 0.178 | 0.7068 | No |
| 58 | Fas | na | 13090 | 0.151 | 0.6988 | No |
| 59 | Nfkbia | na | 13470 | 0.138 | 0.6952 | No |
| 60 | Igf2 | na | 13646 | 0.131 | 0.6951 | No |
| 61 | Irf2 | na | 14619 | 0.095 | 0.6800 | No |
| 62 | Diablo | na | 14790 | 0.089 | 0.6790 | No |
| 63 | Fasl | na | 15266 | 0.071 | 0.6721 | No |
| 64 | Igf1r | na | 15601 | 0.060 | 0.6676 | No |
| 65 | Irf6 | na | 17237 | 0.018 | 0.6389 | No |
| 66 | Casp6 | na | 17501 | 0.015 | 0.6346 | No |
| 67 | Ikbkg | na | 17611 | 0.013 | 0.6330 | No |
| 68 | Mapk10 | na | 45373 | -0.000 | 0.1399 | No |
| 69 | Trp73 | na | 45644 | -0.002 | 0.1351 | No |
| 70 | Cradd | na | 47292 | -0.024 | 0.1064 | No |
| 71 | Tnfrsf25 | na | 49961 | -0.123 | 0.0618 | No |
| 72 | Dffa | na | 50157 | -0.132 | 0.0614 | No |
| 73 | Cflar | na | 50489 | -0.146 | 0.0588 | No |
| 74 | Irf3 | na | 51805 | -0.217 | 0.0404 | No |
| 75 | Dffb | na | 52711 | -0.271 | 0.0305 | No |
| 76 | Prf1 | na | 54301 | -0.400 | 0.0114 | No |
| 77 | Hrk | na | 55266 | -0.534 | 0.0064 | No |
| 78 | Jun | na | 55488 | -0.584 | 0.0158 | No |