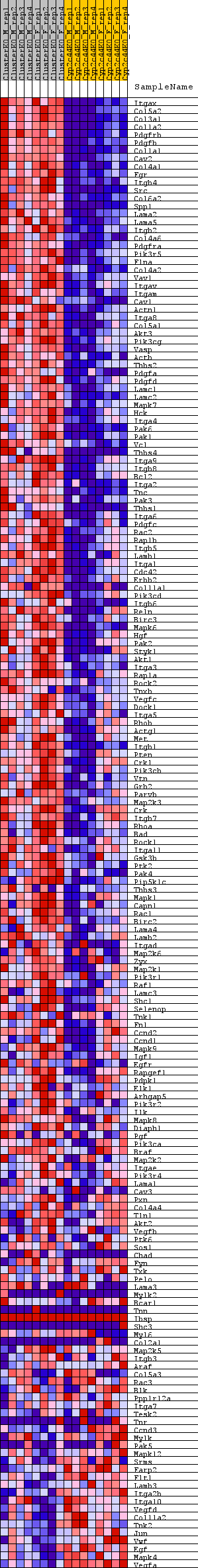
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___ClusterKO_vs_Cyp2c44KO.RPKM_matrix___ClusterKO_vs_Cyp2c44KO.cls #ClusterKO_versus_Cyp2c44KO |
| Phenotype | RPKM_matrix___ClusterKO_vs_Cyp2c44KO.cls#ClusterKO_versus_Cyp2c44KO |
| Upregulated in class | ClusterKO |
| GeneSet | WP_FOCAL_ADHESION |
| Enrichment Score (ES) | 0.7360021 |
| Normalized Enrichment Score (NES) | 1.4165789 |
| Nominal p-value | 0.001996008 |
| FDR q-value | 0.06926594 |
| FWER p-Value | 0.355 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Itgax | na | 27 | 2.329 | 0.0172 | Yes |
| 2 | Col5a2 | na | 51 | 2.188 | 0.0335 | Yes |
| 3 | Col3a1 | na | 89 | 2.027 | 0.0482 | Yes |
| 4 | Col1a2 | na | 94 | 2.011 | 0.0634 | Yes |
| 5 | Pdgfrb | na | 120 | 1.934 | 0.0777 | Yes |
| 6 | Pdgfb | na | 130 | 1.893 | 0.0919 | Yes |
| 7 | Col1a1 | na | 138 | 1.868 | 0.1060 | Yes |
| 8 | Cav2 | na | 155 | 1.831 | 0.1196 | Yes |
| 9 | Col4a1 | na | 215 | 1.736 | 0.1318 | Yes |
| 10 | Fgr | na | 219 | 1.726 | 0.1449 | Yes |
| 11 | Itgb4 | na | 236 | 1.711 | 0.1576 | Yes |
| 12 | Src | na | 286 | 1.640 | 0.1692 | Yes |
| 13 | Col6a2 | na | 305 | 1.618 | 0.1812 | Yes |
| 14 | Spp1 | na | 306 | 1.617 | 0.1935 | Yes |
| 15 | Lama2 | na | 320 | 1.602 | 0.2054 | Yes |
| 16 | Lama5 | na | 330 | 1.597 | 0.2174 | Yes |
| 17 | Itgb2 | na | 397 | 1.529 | 0.2278 | Yes |
| 18 | Col4a6 | na | 450 | 1.481 | 0.2382 | Yes |
| 19 | Pdgfra | na | 466 | 1.466 | 0.2491 | Yes |
| 20 | Pik3r5 | na | 485 | 1.456 | 0.2598 | Yes |
| 21 | Flna | na | 516 | 1.438 | 0.2702 | Yes |
| 22 | Col4a2 | na | 673 | 1.340 | 0.2776 | Yes |
| 23 | Vav1 | na | 679 | 1.334 | 0.2877 | Yes |
| 24 | Itgav | na | 717 | 1.312 | 0.2970 | Yes |
| 25 | Itgam | na | 718 | 1.311 | 0.3070 | Yes |
| 26 | Cav1 | na | 734 | 1.304 | 0.3166 | Yes |
| 27 | Actn1 | na | 744 | 1.298 | 0.3263 | Yes |
| 28 | Itga8 | na | 781 | 1.281 | 0.3354 | Yes |
| 29 | Col5a1 | na | 807 | 1.264 | 0.3446 | Yes |
| 30 | Akt3 | na | 817 | 1.258 | 0.3540 | Yes |
| 31 | Pik3cg | na | 855 | 1.239 | 0.3628 | Yes |
| 32 | Vasp | na | 893 | 1.222 | 0.3714 | Yes |
| 33 | Actb | na | 930 | 1.201 | 0.3799 | Yes |
| 34 | Thbs2 | na | 932 | 1.200 | 0.3890 | Yes |
| 35 | Pdgfa | na | 933 | 1.198 | 0.3981 | Yes |
| 36 | Pdgfd | na | 946 | 1.194 | 0.4070 | Yes |
| 37 | Lamc1 | na | 947 | 1.194 | 0.4161 | Yes |
| 38 | Lamc2 | na | 973 | 1.183 | 0.4246 | Yes |
| 39 | Mapk7 | na | 1010 | 1.164 | 0.4328 | Yes |
| 40 | Hck | na | 1130 | 1.123 | 0.4392 | Yes |
| 41 | Itga4 | na | 1166 | 1.105 | 0.4470 | Yes |
| 42 | Pak6 | na | 1168 | 1.105 | 0.4554 | Yes |
| 43 | Pak1 | na | 1172 | 1.102 | 0.4637 | Yes |
| 44 | Vcl | na | 1185 | 1.098 | 0.4719 | Yes |
| 45 | Thbs4 | na | 1228 | 1.088 | 0.4794 | Yes |
| 46 | Itga9 | na | 1242 | 1.083 | 0.4874 | Yes |
| 47 | Itgb8 | na | 1259 | 1.077 | 0.4953 | Yes |
| 48 | Bcl2 | na | 1285 | 1.071 | 0.5030 | Yes |
| 49 | Itga2 | na | 1286 | 1.071 | 0.5111 | Yes |
| 50 | Tnc | na | 1292 | 1.067 | 0.5191 | Yes |
| 51 | Pak3 | na | 1368 | 1.041 | 0.5257 | Yes |
| 52 | Thbs1 | na | 1376 | 1.040 | 0.5335 | Yes |
| 53 | Itga6 | na | 1415 | 1.028 | 0.5407 | Yes |
| 54 | Pdgfc | na | 1461 | 1.013 | 0.5476 | Yes |
| 55 | Rac2 | na | 1555 | 0.991 | 0.5534 | Yes |
| 56 | Rap1b | na | 1662 | 0.958 | 0.5588 | Yes |
| 57 | Itgb5 | na | 1724 | 0.944 | 0.5649 | Yes |
| 58 | Lamb1 | na | 1799 | 0.928 | 0.5707 | Yes |
| 59 | Itgal | na | 1838 | 0.918 | 0.5770 | Yes |
| 60 | Cdc42 | na | 1888 | 0.906 | 0.5830 | Yes |
| 61 | Erbb2 | na | 1909 | 0.904 | 0.5895 | Yes |
| 62 | Col11a1 | na | 1918 | 0.901 | 0.5962 | Yes |
| 63 | Pik3cd | na | 2000 | 0.883 | 0.6015 | Yes |
| 64 | Itgb6 | na | 2019 | 0.876 | 0.6078 | Yes |
| 65 | Reln | na | 2132 | 0.855 | 0.6123 | Yes |
| 66 | Birc3 | na | 2142 | 0.852 | 0.6187 | Yes |
| 67 | Mapk6 | na | 2183 | 0.844 | 0.6244 | Yes |
| 68 | Hgf | na | 2361 | 0.811 | 0.6274 | Yes |
| 69 | Pak2 | na | 2368 | 0.810 | 0.6334 | Yes |
| 70 | Styk1 | na | 2486 | 0.791 | 0.6374 | Yes |
| 71 | Akt1 | na | 2566 | 0.775 | 0.6419 | Yes |
| 72 | Itga3 | na | 2631 | 0.765 | 0.6465 | Yes |
| 73 | Rap1a | na | 2766 | 0.744 | 0.6498 | Yes |
| 74 | Rock2 | na | 2876 | 0.728 | 0.6534 | Yes |
| 75 | Tnxb | na | 2885 | 0.727 | 0.6588 | Yes |
| 76 | Vegfc | na | 3021 | 0.711 | 0.6618 | Yes |
| 77 | Dock1 | na | 3080 | 0.704 | 0.6661 | Yes |
| 78 | Itga5 | na | 3338 | 0.673 | 0.6666 | Yes |
| 79 | Rhob | na | 3508 | 0.654 | 0.6686 | Yes |
| 80 | Actg1 | na | 3914 | 0.619 | 0.6661 | Yes |
| 81 | Met | na | 3990 | 0.612 | 0.6694 | Yes |
| 82 | Itgb1 | na | 4038 | 0.607 | 0.6732 | Yes |
| 83 | Pten | na | 4212 | 0.590 | 0.6746 | Yes |
| 84 | Crkl | na | 4356 | 0.580 | 0.6765 | Yes |
| 85 | Pik3cb | na | 4728 | 0.552 | 0.6741 | Yes |
| 86 | Vtn | na | 4779 | 0.549 | 0.6774 | Yes |
| 87 | Grb2 | na | 4783 | 0.548 | 0.6815 | Yes |
| 88 | Parvb | na | 4837 | 0.544 | 0.6847 | Yes |
| 89 | Map2k3 | na | 4915 | 0.539 | 0.6874 | Yes |
| 90 | Crk | na | 4924 | 0.538 | 0.6914 | Yes |
| 91 | Itgb7 | na | 4932 | 0.537 | 0.6953 | Yes |
| 92 | Rhoa | na | 4959 | 0.535 | 0.6989 | Yes |
| 93 | Bad | na | 4961 | 0.535 | 0.7030 | Yes |
| 94 | Rock1 | na | 5099 | 0.525 | 0.7045 | Yes |
| 95 | Itga11 | na | 5109 | 0.524 | 0.7084 | Yes |
| 96 | Gsk3b | na | 5233 | 0.515 | 0.7101 | Yes |
| 97 | Ptk2 | na | 5236 | 0.515 | 0.7140 | Yes |
| 98 | Pak4 | na | 5244 | 0.514 | 0.7177 | Yes |
| 99 | Pip5k1c | na | 5579 | 0.490 | 0.7155 | Yes |
| 100 | Thbs3 | na | 5601 | 0.488 | 0.7189 | Yes |
| 101 | Mapk1 | na | 5650 | 0.485 | 0.7217 | Yes |
| 102 | Capn1 | na | 6019 | 0.463 | 0.7187 | Yes |
| 103 | Rac1 | na | 6117 | 0.458 | 0.7204 | Yes |
| 104 | Birc2 | na | 6158 | 0.456 | 0.7232 | Yes |
| 105 | Lama4 | na | 6308 | 0.447 | 0.7239 | Yes |
| 106 | Lamb2 | na | 6344 | 0.445 | 0.7267 | Yes |
| 107 | Itgad | na | 6602 | 0.431 | 0.7254 | Yes |
| 108 | Map2k6 | na | 6948 | 0.413 | 0.7224 | Yes |
| 109 | Zyx | na | 7023 | 0.409 | 0.7242 | Yes |
| 110 | Map2k1 | na | 7116 | 0.405 | 0.7256 | Yes |
| 111 | Pik3r1 | na | 7254 | 0.399 | 0.7262 | Yes |
| 112 | Raf1 | na | 7290 | 0.398 | 0.7286 | Yes |
| 113 | Lamc3 | na | 7357 | 0.395 | 0.7305 | Yes |
| 114 | Shc1 | na | 7378 | 0.394 | 0.7331 | Yes |
| 115 | Selenop | na | 7384 | 0.394 | 0.7360 | Yes |
| 116 | Tnk1 | na | 7589 | 0.384 | 0.7353 | No |
| 117 | Fn1 | na | 7934 | 0.367 | 0.7320 | No |
| 118 | Ccnd2 | na | 8235 | 0.353 | 0.7293 | No |
| 119 | Ccnd1 | na | 8289 | 0.351 | 0.7310 | No |
| 120 | Mapk9 | na | 8443 | 0.343 | 0.7309 | No |
| 121 | Igf1 | na | 8454 | 0.342 | 0.7333 | No |
| 122 | Egfr | na | 8660 | 0.333 | 0.7322 | No |
| 123 | Rapgef1 | na | 8867 | 0.323 | 0.7310 | No |
| 124 | Pdpk1 | na | 9161 | 0.309 | 0.7282 | No |
| 125 | Elk1 | na | 9185 | 0.308 | 0.7301 | No |
| 126 | Arhgap5 | na | 9380 | 0.301 | 0.7289 | No |
| 127 | Pik3r2 | na | 9603 | 0.291 | 0.7272 | No |
| 128 | Ilk | na | 10134 | 0.268 | 0.7198 | No |
| 129 | Mapk8 | na | 10431 | 0.255 | 0.7165 | No |
| 130 | Diaph1 | na | 10746 | 0.242 | 0.7127 | No |
| 131 | Pgf | na | 10915 | 0.234 | 0.7115 | No |
| 132 | Pik3ca | na | 11645 | 0.207 | 0.7001 | No |
| 133 | Braf | na | 11910 | 0.196 | 0.6969 | No |
| 134 | Map2k2 | na | 12044 | 0.192 | 0.6960 | No |
| 135 | Itgae | na | 12120 | 0.190 | 0.6961 | No |
| 136 | Pik3r4 | na | 12131 | 0.189 | 0.6974 | No |
| 137 | Lama1 | na | 12309 | 0.183 | 0.6956 | No |
| 138 | Cav3 | na | 12316 | 0.183 | 0.6969 | No |
| 139 | Pxn | na | 12509 | 0.175 | 0.6948 | No |
| 140 | Col4a4 | na | 13142 | 0.149 | 0.6847 | No |
| 141 | Tln1 | na | 13576 | 0.132 | 0.6780 | No |
| 142 | Akt2 | na | 13614 | 0.132 | 0.6783 | No |
| 143 | Vegfb | na | 14559 | 0.096 | 0.6623 | No |
| 144 | Ptk6 | na | 14588 | 0.096 | 0.6625 | No |
| 145 | Sos1 | na | 15662 | 0.058 | 0.6438 | No |
| 146 | Chad | na | 16098 | 0.044 | 0.6364 | No |
| 147 | Fyn | na | 16201 | 0.041 | 0.6349 | No |
| 148 | Txk | na | 16845 | 0.025 | 0.6237 | No |
| 149 | Pelo | na | 17281 | 0.017 | 0.6161 | No |
| 150 | Lama3 | na | 17972 | 0.009 | 0.6039 | No |
| 151 | Mylk2 | na | 18034 | 0.009 | 0.6028 | No |
| 152 | Bcar1 | na | 18139 | 0.008 | 0.6010 | No |
| 153 | Tnn | na | 19138 | 0.001 | 0.5833 | No |
| 154 | Ibsp | na | 23488 | 0.000 | 0.5059 | No |
| 155 | Shc3 | na | 45400 | -0.001 | 0.1160 | No |
| 156 | Myl6 | na | 45497 | -0.001 | 0.1143 | No |
| 157 | Col2a1 | na | 45817 | -0.003 | 0.1086 | No |
| 158 | Map2k5 | na | 47914 | -0.040 | 0.0716 | No |
| 159 | Itgb3 | na | 48205 | -0.050 | 0.0668 | No |
| 160 | Araf | na | 49401 | -0.098 | 0.0463 | No |
| 161 | Col5a3 | na | 49509 | -0.103 | 0.0452 | No |
| 162 | Rac3 | na | 50165 | -0.132 | 0.0346 | No |
| 163 | Blk | na | 50294 | -0.137 | 0.0333 | No |
| 164 | Ppp1r12a | na | 50327 | -0.138 | 0.0338 | No |
| 165 | Itga7 | na | 50435 | -0.143 | 0.0330 | No |
| 166 | Tesk2 | na | 50561 | -0.149 | 0.0319 | No |
| 167 | Tnr | na | 51689 | -0.212 | 0.0134 | No |
| 168 | Ccnd3 | na | 51911 | -0.223 | 0.0112 | No |
| 169 | Mylk | na | 52272 | -0.242 | 0.0066 | No |
| 170 | Pak5 | na | 52630 | -0.265 | 0.0023 | No |
| 171 | Mapk12 | na | 53628 | -0.339 | -0.0129 | No |
| 172 | Srms | na | 54005 | -0.372 | -0.0167 | No |
| 173 | Farp2 | na | 54567 | -0.428 | -0.0235 | No |
| 174 | Flt1 | na | 54568 | -0.428 | -0.0202 | No |
| 175 | Lamb3 | na | 54941 | -0.477 | -0.0232 | No |
| 176 | Itga2b | na | 55091 | -0.502 | -0.0220 | No |
| 177 | Itga10 | na | 55312 | -0.544 | -0.0218 | No |
| 178 | Vegfd | na | 55444 | -0.576 | -0.0198 | No |
| 179 | Col11a2 | na | 55459 | -0.579 | -0.0156 | No |
| 180 | Tnk2 | na | 55486 | -0.583 | -0.0116 | No |
| 181 | Jun | na | 55488 | -0.584 | -0.0072 | No |
| 182 | Vwf | na | 55671 | -0.633 | -0.0056 | No |
| 183 | Egf | na | 55719 | -0.651 | -0.0015 | No |
| 184 | Mapk4 | na | 56045 | -0.819 | -0.0011 | No |
| 185 | Vegfa | na | 56181 | -0.917 | 0.0035 | No |