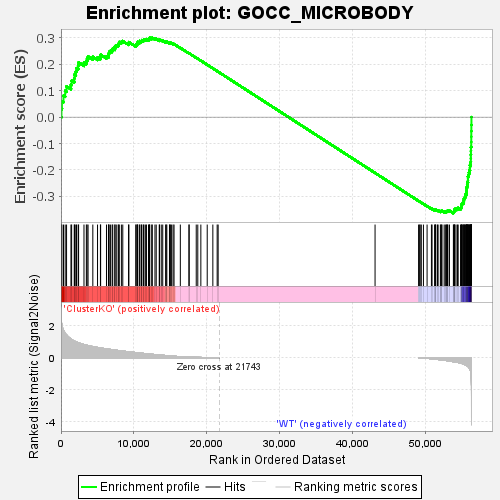
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___ClusterKO_vs_WT.RPKM_matrix___ClusterKO_vs_WT.cls #ClusterKO_versus_WT |
| Phenotype | RPKM_matrix___ClusterKO_vs_WT.cls#ClusterKO_versus_WT |
| Upregulated in class | WT |
| GeneSet | GOCC_MICROBODY |
| Enrichment Score (ES) | -0.3639196 |
| Normalized Enrichment Score (NES) | -0.8743506 |
| Nominal p-value | 0.68267226 |
| FDR q-value | 0.82594883 |
| FWER p-Value | 1.0 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Myo5a | na | 61 | 2.259 | 0.0318 | No |
| 2 | Sting1 | na | 124 | 2.032 | 0.0602 | No |
| 3 | Rab8b | na | 343 | 1.715 | 0.0812 | No |
| 4 | Far1 | na | 573 | 1.525 | 0.0993 | No |
| 5 | Vim | na | 763 | 1.418 | 0.1166 | No |
| 6 | Cd33 | na | 1368 | 1.171 | 0.1229 | No |
| 7 | Syt7 | na | 1460 | 1.146 | 0.1379 | No |
| 8 | Agps | na | 1823 | 1.040 | 0.1466 | No |
| 9 | Atm | na | 1849 | 1.035 | 0.1612 | No |
| 10 | Cav1 | na | 2033 | 1.002 | 0.1725 | No |
| 11 | Acsl4 | na | 2129 | 0.982 | 0.1851 | No |
| 12 | Trim37 | na | 2355 | 0.942 | 0.1948 | No |
| 13 | Kcnip4 | na | 2415 | 0.931 | 0.2073 | No |
| 14 | Xdh | na | 3156 | 0.824 | 0.2061 | No |
| 15 | Akap11 | na | 3481 | 0.787 | 0.2118 | No |
| 16 | Isoc1 | na | 3579 | 0.773 | 0.2213 | No |
| 17 | Nos2 | na | 3717 | 0.756 | 0.2299 | No |
| 18 | Lacc1 | na | 4371 | 0.695 | 0.2284 | No |
| 19 | Mff | na | 5017 | 0.643 | 0.2262 | No |
| 20 | Atad1 | na | 5415 | 0.611 | 0.2281 | No |
| 21 | Pik3c3 | na | 5443 | 0.608 | 0.2364 | No |
| 22 | Pex5l | na | 6247 | 0.550 | 0.2301 | No |
| 23 | Nudt17 | na | 6516 | 0.533 | 0.2331 | No |
| 24 | Zadh2 | na | 6543 | 0.531 | 0.2404 | No |
| 25 | Crot | na | 6610 | 0.526 | 0.2468 | No |
| 26 | Serhl | na | 6788 | 0.516 | 0.2512 | No |
| 27 | Pex26 | na | 7022 | 0.500 | 0.2543 | No |
| 28 | Mpv17 | na | 7104 | 0.495 | 0.2601 | No |
| 29 | Abcd1 | na | 7401 | 0.476 | 0.2617 | No |
| 30 | Nudt19 | na | 7406 | 0.476 | 0.2686 | No |
| 31 | Acbd5 | na | 7589 | 0.466 | 0.2721 | No |
| 32 | Pex7 | na | 7847 | 0.451 | 0.2741 | No |
| 33 | Marf1 | na | 7921 | 0.448 | 0.2793 | No |
| 34 | Acox3 | na | 8002 | 0.442 | 0.2843 | No |
| 35 | Pex2 | na | 8293 | 0.426 | 0.2854 | No |
| 36 | Uox | na | 8485 | 0.415 | 0.2880 | No |
| 37 | Fndc5 | na | 9278 | 0.371 | 0.2793 | No |
| 38 | Pnpla8 | na | 9323 | 0.369 | 0.2839 | No |
| 39 | Agxt | na | 10277 | 0.319 | 0.2716 | No |
| 40 | Far2 | na | 10315 | 0.317 | 0.2755 | No |
| 41 | Mul1 | na | 10463 | 0.309 | 0.2774 | No |
| 42 | Slc25a17 | na | 10502 | 0.307 | 0.2812 | No |
| 43 | Szt2 | na | 10549 | 0.305 | 0.2848 | No |
| 44 | Gnpat | na | 10803 | 0.291 | 0.2846 | No |
| 45 | Prdx1 | na | 10831 | 0.289 | 0.2883 | No |
| 46 | Tirap | na | 11056 | 0.279 | 0.2884 | No |
| 47 | Slc22a21 | na | 11109 | 0.277 | 0.2915 | No |
| 48 | Nbr1 | na | 11336 | 0.265 | 0.2913 | No |
| 49 | Pex12 | na | 11394 | 0.263 | 0.2941 | No |
| 50 | Acot6 | na | 11625 | 0.252 | 0.2937 | No |
| 51 | Pjvk | na | 11735 | 0.245 | 0.2953 | No |
| 52 | Dnm1l | na | 12017 | 0.232 | 0.2937 | No |
| 53 | Pex10 | na | 12098 | 0.229 | 0.2956 | No |
| 54 | Arf1 | na | 12164 | 0.225 | 0.2977 | No |
| 55 | Pex11b | na | 12184 | 0.224 | 0.3006 | No |
| 56 | Pex13 | na | 12462 | 0.212 | 0.2988 | No |
| 57 | Pex5 | na | 12535 | 0.209 | 0.3005 | No |
| 58 | Tmem35a | na | 12906 | 0.193 | 0.2968 | No |
| 59 | Prdx5 | na | 13078 | 0.185 | 0.2964 | No |
| 60 | Ide | na | 13494 | 0.168 | 0.2915 | No |
| 61 | Amacr | na | 13528 | 0.166 | 0.2933 | No |
| 62 | Acsbg1 | na | 13768 | 0.156 | 0.2913 | No |
| 63 | Idh2 | na | 13972 | 0.148 | 0.2898 | No |
| 64 | Mgst1 | na | 14374 | 0.132 | 0.2846 | No |
| 65 | Acnat2 | na | 14442 | 0.130 | 0.2853 | No |
| 66 | Nudt12 | na | 14534 | 0.126 | 0.2855 | No |
| 67 | Pex3 | na | 14899 | 0.109 | 0.2807 | No |
| 68 | Dao | na | 14951 | 0.107 | 0.2813 | No |
| 69 | Acnat1 | na | 15019 | 0.105 | 0.2816 | No |
| 70 | Pex6 | na | 15162 | 0.100 | 0.2806 | No |
| 71 | Fis1 | na | 15386 | 0.093 | 0.2780 | No |
| 72 | Pex11g | na | 15525 | 0.089 | 0.2768 | No |
| 73 | Scp2 | na | 16389 | 0.060 | 0.2623 | No |
| 74 | Tmem135 | na | 17578 | 0.033 | 0.2417 | No |
| 75 | Baat | na | 17596 | 0.032 | 0.2418 | No |
| 76 | Gstk1 | na | 18574 | 0.019 | 0.2247 | No |
| 77 | Dhrs7b | na | 18787 | 0.017 | 0.2212 | No |
| 78 | Plaat1 | na | 19207 | 0.013 | 0.2139 | No |
| 79 | Rida | na | 20082 | 0.007 | 0.1985 | No |
| 80 | Idi2 | na | 20855 | 0.004 | 0.1848 | No |
| 81 | Ccdc33 | na | 21448 | 0.001 | 0.1743 | No |
| 82 | Acot5 | na | 21583 | 0.001 | 0.1720 | No |
| 83 | Idi2l | na | 43134 | 0.000 | -0.2113 | No |
| 84 | Pecr | na | 49112 | -0.000 | -0.3176 | No |
| 85 | Depp1 | na | 49261 | -0.005 | -0.3202 | No |
| 86 | Pex19 | na | 49323 | -0.007 | -0.3212 | No |
| 87 | Pxmp2 | na | 49485 | -0.011 | -0.3239 | No |
| 88 | Acad11 | na | 49778 | -0.021 | -0.3288 | No |
| 89 | Cat | na | 50280 | -0.041 | -0.3371 | No |
| 90 | Mtarc2 | na | 50876 | -0.067 | -0.3467 | No |
| 91 | Mavs | na | 50962 | -0.071 | -0.3472 | No |
| 92 | Pipox | na | 51318 | -0.088 | -0.3522 | No |
| 93 | Acoxl | na | 51369 | -0.091 | -0.3518 | No |
| 94 | Acot4 | na | 51448 | -0.094 | -0.3518 | No |
| 95 | Hao1 | na | 51466 | -0.095 | -0.3507 | No |
| 96 | Hspd1 | na | 51705 | -0.108 | -0.3534 | No |
| 97 | Hmgcl | na | 51792 | -0.112 | -0.3533 | No |
| 98 | Gbf1 | na | 52104 | -0.130 | -0.3569 | No |
| 99 | Plaat3 | na | 52172 | -0.134 | -0.3562 | No |
| 100 | Nudt7 | na | 52284 | -0.140 | -0.3561 | No |
| 101 | Phyh | na | 52315 | -0.142 | -0.3546 | No |
| 102 | Acsl6 | na | 52625 | -0.160 | -0.3578 | No |
| 103 | Tkt | na | 52787 | -0.170 | -0.3581 | No |
| 104 | Abcd4 | na | 52929 | -0.179 | -0.3580 | No |
| 105 | Pomc | na | 52975 | -0.182 | -0.3562 | No |
| 106 | Lonp2 | na | 53085 | -0.188 | -0.3554 | No |
| 107 | Mlycd | na | 53134 | -0.192 | -0.3535 | No |
| 108 | Pex1 | na | 53352 | -0.206 | -0.3543 | No |
| 109 | Decr2 | na | 53892 | -0.250 | -0.3603 | Yes |
| 110 | Idh1 | na | 53939 | -0.255 | -0.3574 | Yes |
| 111 | Pex14 | na | 54019 | -0.261 | -0.3550 | Yes |
| 112 | Tysnd1 | na | 54053 | -0.263 | -0.3518 | Yes |
| 113 | Acaa1a | na | 54070 | -0.265 | -0.3482 | Yes |
| 114 | Ddo | na | 54253 | -0.280 | -0.3474 | Yes |
| 115 | Abcd2 | na | 54451 | -0.297 | -0.3466 | Yes |
| 116 | Acsl3 | na | 54548 | -0.304 | -0.3439 | Yes |
| 117 | Eci3 | na | 54877 | -0.339 | -0.3448 | Yes |
| 118 | Pxt1 | na | 54969 | -0.350 | -0.3413 | Yes |
| 119 | Idi1-ps1 | na | 55024 | -0.358 | -0.3371 | Yes |
| 120 | Vwa8 | na | 55029 | -0.358 | -0.3319 | Yes |
| 121 | Ephx2 | na | 55076 | -0.364 | -0.3274 | Yes |
| 122 | Slc27a2 | na | 55177 | -0.383 | -0.3237 | Yes |
| 123 | Acot3 | na | 55319 | -0.404 | -0.3203 | Yes |
| 124 | Hacl1 | na | 55327 | -0.405 | -0.3145 | Yes |
| 125 | Acot8 | na | 55344 | -0.408 | -0.3089 | Yes |
| 126 | Acox2 | na | 55433 | -0.423 | -0.3043 | Yes |
| 127 | Urah | na | 55522 | -0.445 | -0.2994 | Yes |
| 128 | Acox1 | na | 55537 | -0.448 | -0.2931 | Yes |
| 129 | Hsd17b4 | na | 55677 | -0.480 | -0.2886 | Yes |
| 130 | Ehhadh | na | 55684 | -0.481 | -0.2817 | Yes |
| 131 | Dhrs4 | na | 55711 | -0.488 | -0.2751 | Yes |
| 132 | Hao2 | na | 55716 | -0.491 | -0.2680 | Yes |
| 133 | Pex16 | na | 55767 | -0.505 | -0.2616 | Yes |
| 134 | Hsdl2 | na | 55796 | -0.512 | -0.2546 | Yes |
| 135 | Abcd3 | na | 55814 | -0.517 | -0.2474 | Yes |
| 136 | Pxmp4 | na | 55891 | -0.546 | -0.2408 | Yes |
| 137 | Hmgcr | na | 55895 | -0.547 | -0.2329 | Yes |
| 138 | Urad | na | 55897 | -0.548 | -0.2250 | Yes |
| 139 | Mpv17l | na | 55980 | -0.587 | -0.2179 | Yes |
| 140 | Paox | na | 56036 | -0.626 | -0.2098 | Yes |
| 141 | Aldh3a2 | na | 56103 | -0.671 | -0.2012 | Yes |
| 142 | Idi1 | na | 56164 | -0.723 | -0.1917 | Yes |
| 143 | Eci2 | na | 56169 | -0.730 | -0.1812 | Yes |
| 144 | Acsl1 | na | 56272 | -0.893 | -0.1700 | Yes |
| 145 | Mvk | na | 56287 | -0.947 | -0.1565 | Yes |
| 146 | Acaa1b | na | 56290 | -0.974 | -0.1424 | Yes |
| 147 | Sod1 | na | 56300 | -0.995 | -0.1281 | Yes |
| 148 | Ech1 | na | 56305 | -1.013 | -0.1134 | Yes |
| 149 | Pex11a | na | 56346 | -1.351 | -0.0945 | Yes |
| 150 | Fabp1 | na | 56350 | -1.400 | -0.0742 | Yes |
| 151 | Mvd | na | 56353 | -1.438 | -0.0533 | Yes |
| 152 | Crat | na | 56361 | -1.626 | -0.0298 | Yes |
| 153 | Fdps | na | 56372 | -2.069 | 0.0001 | Yes |

