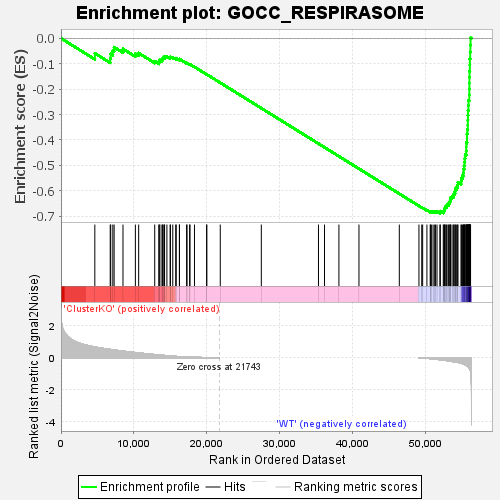
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___ClusterKO_vs_WT.RPKM_matrix___ClusterKO_vs_WT.cls #ClusterKO_versus_WT |
| Phenotype | RPKM_matrix___ClusterKO_vs_WT.cls#ClusterKO_versus_WT |
| Upregulated in class | WT |
| GeneSet | GOCC_RESPIRASOME |
| Enrichment Score (ES) | -0.6883782 |
| Normalized Enrichment Score (NES) | -1.3482177 |
| Nominal p-value | 0.07723577 |
| FDR q-value | 0.35392088 |
| FWER p-Value | 0.808 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | 1700066M21Rik | na | 4646 | 0.671 | -0.0588 | No |
| 2 | AA467197 | na | 6760 | 0.518 | -0.0780 | No |
| 3 | Cox6a2 | na | 6829 | 0.513 | -0.0610 | No |
| 4 | Cox6c2 | na | 7119 | 0.494 | -0.0487 | No |
| 5 | Ttc19 | na | 7296 | 0.483 | -0.0347 | No |
| 6 | Uqcc4 | na | 8513 | 0.414 | -0.0417 | No |
| 7 | Cox4i2 | na | 10214 | 0.323 | -0.0604 | No |
| 8 | Dmac1 | na | 10668 | 0.299 | -0.0579 | No |
| 9 | Uqcrfs1 | na | 12871 | 0.194 | -0.0902 | No |
| 10 | Ndufa4 | na | 13452 | 0.169 | -0.0945 | No |
| 11 | Ndufb6 | na | 13465 | 0.169 | -0.0887 | No |
| 12 | Ndufs1 | na | 13582 | 0.164 | -0.0850 | No |
| 13 | Rab5if | na | 13874 | 0.152 | -0.0848 | No |
| 14 | Coa6 | na | 13923 | 0.150 | -0.0803 | No |
| 15 | Sdhc | na | 14038 | 0.146 | -0.0772 | No |
| 16 | Sdhd | na | 14207 | 0.139 | -0.0752 | No |
| 17 | Cox5a | na | 14223 | 0.138 | -0.0706 | No |
| 18 | Cox6b2 | na | 14545 | 0.125 | -0.0719 | No |
| 19 | Ndufb4c | na | 14991 | 0.106 | -0.0760 | No |
| 20 | Cox7a2l | na | 15028 | 0.104 | -0.0730 | No |
| 21 | Ndufc2 | na | 15352 | 0.094 | -0.0754 | No |
| 22 | Ndufb8 | na | 15759 | 0.080 | -0.0798 | No |
| 23 | Ndufs4 | na | 15823 | 0.078 | -0.0781 | No |
| 24 | Ndufs2 | na | 16260 | 0.063 | -0.0836 | No |
| 25 | Ndufa11b | na | 16273 | 0.063 | -0.0816 | No |
| 26 | Sdha | na | 17252 | 0.038 | -0.0976 | No |
| 27 | mt-Nd6 | na | 17308 | 0.037 | -0.0973 | No |
| 28 | Ndufs5 | na | 17669 | 0.031 | -0.1026 | No |
| 29 | mt-Co2 | na | 17713 | 0.031 | -0.1023 | No |
| 30 | Cox7a1 | na | 18331 | 0.022 | -0.1125 | No |
| 31 | Ndufc1 | na | 20012 | 0.008 | -0.1420 | No |
| 32 | Ndufa2 | na | 20015 | 0.008 | -0.1418 | No |
| 33 | mt-Nd4l | na | 21874 | 0.000 | -0.1748 | No |
| 34 | Cox8c | na | 27512 | 0.000 | -0.2750 | No |
| 35 | Ndufb4b | na | 35363 | 0.000 | -0.4145 | No |
| 36 | Ndufb11b | na | 36192 | 0.000 | -0.4292 | No |
| 37 | Cyct | na | 38170 | 0.000 | -0.4643 | No |
| 38 | mt-Co3 | na | 40924 | 0.000 | -0.5132 | No |
| 39 | Cox7b2 | na | 46465 | 0.000 | -0.6117 | No |
| 40 | Ndufv1 | na | 49162 | -0.002 | -0.6595 | No |
| 41 | Cyc1 | na | 49545 | -0.013 | -0.6658 | No |
| 42 | Uqcrc2 | na | 49672 | -0.018 | -0.6675 | No |
| 43 | Uqcrc1 | na | 50257 | -0.040 | -0.6764 | No |
| 44 | Wdr93 | na | 50710 | -0.059 | -0.6824 | No |
| 45 | Ndufs3 | na | 50744 | -0.061 | -0.6808 | No |
| 46 | mt-Nd3 | na | 50904 | -0.068 | -0.6812 | No |
| 47 | Ndufa7 | na | 51108 | -0.079 | -0.6821 | No |
| 48 | Ndufa9 | na | 51325 | -0.088 | -0.6828 | No |
| 49 | Ndufb2 | na | 51447 | -0.094 | -0.6816 | No |
| 50 | Ndufv2 | na | 51643 | -0.104 | -0.6814 | No |
| 51 | Cox7c | na | 52038 | -0.126 | -0.6839 | Yes |
| 52 | Cox4i1 | na | 52100 | -0.130 | -0.6804 | Yes |
| 53 | Sdhb | na | 52503 | -0.154 | -0.6821 | Yes |
| 54 | Ndufab1-ps | na | 52592 | -0.158 | -0.6780 | Yes |
| 55 | Ndufb4 | na | 52680 | -0.164 | -0.6738 | Yes |
| 56 | Cox7a2 | na | 52713 | -0.166 | -0.6685 | Yes |
| 57 | Foxred1 | na | 52767 | -0.170 | -0.6634 | Yes |
| 58 | Ndufb5 | na | 52967 | -0.182 | -0.6606 | Yes |
| 59 | Cox8a | na | 53018 | -0.185 | -0.6549 | Yes |
| 60 | Ndufb1 | na | 53239 | -0.198 | -0.6518 | Yes |
| 61 | Ndufs6b | na | 53303 | -0.202 | -0.6458 | Yes |
| 62 | Uqcrq | na | 53419 | -0.211 | -0.6403 | Yes |
| 63 | Cox8b | na | 53461 | -0.214 | -0.6335 | Yes |
| 64 | Uqcr10 | na | 53509 | -0.218 | -0.6266 | Yes |
| 65 | Uqcrh | na | 53712 | -0.236 | -0.6218 | Yes |
| 66 | Stmp1 | na | 53927 | -0.254 | -0.6166 | Yes |
| 67 | Ndufs7 | na | 53990 | -0.259 | -0.6086 | Yes |
| 68 | Uqcrh-ps1 | na | 54138 | -0.270 | -0.6016 | Yes |
| 69 | Ndufa4l2 | na | 54151 | -0.272 | -0.5922 | Yes |
| 70 | Cox7b | na | 54342 | -0.286 | -0.5855 | Yes |
| 71 | Ndufab1 | na | 54487 | -0.300 | -0.5774 | Yes |
| 72 | Ndufa11 | na | 54507 | -0.301 | -0.5671 | Yes |
| 73 | Higd1a | na | 54941 | -0.346 | -0.5625 | Yes |
| 74 | Ndufa6 | na | 54990 | -0.354 | -0.5508 | Yes |
| 75 | Ndufb11 | na | 55137 | -0.376 | -0.5401 | Yes |
| 76 | Ndufa10 | na | 55280 | -0.398 | -0.5286 | Yes |
| 77 | Ndufa3 | na | 55293 | -0.400 | -0.5146 | Yes |
| 78 | mt-Co1 | na | 55363 | -0.412 | -0.5013 | Yes |
| 79 | mt-Nd5 | na | 55416 | -0.420 | -0.4873 | Yes |
| 80 | Ndufs8 | na | 55463 | -0.429 | -0.4729 | Yes |
| 81 | Cox6c | na | 55511 | -0.442 | -0.4581 | Yes |
| 82 | Ndufa8 | na | 55652 | -0.473 | -0.4439 | Yes |
| 83 | Ndufb7 | na | 55663 | -0.475 | -0.4272 | Yes |
| 84 | Cox6a1 | na | 55665 | -0.476 | -0.4104 | Yes |
| 85 | Ndufa1 | na | 55761 | -0.502 | -0.3943 | Yes |
| 86 | Dmac2 | na | 55763 | -0.503 | -0.3765 | Yes |
| 87 | Uqcc3 | na | 55809 | -0.515 | -0.3591 | Yes |
| 88 | Ndufa12 | na | 55871 | -0.536 | -0.3412 | Yes |
| 89 | Ndufb9 | na | 55874 | -0.537 | -0.3222 | Yes |
| 90 | mt-Nd2 | na | 55886 | -0.543 | -0.3032 | Yes |
| 91 | Cox6b1 | na | 55906 | -0.553 | -0.2839 | Yes |
| 92 | Ndufv3 | na | 55958 | -0.578 | -0.2644 | Yes |
| 93 | Ndufs6 | na | 55970 | -0.584 | -0.2439 | Yes |
| 94 | Cox5b | na | 56075 | -0.648 | -0.2228 | Yes |
| 95 | Ndufa13 | na | 56082 | -0.650 | -0.1999 | Yes |
| 96 | Uqcr11 | na | 56089 | -0.659 | -0.1767 | Yes |
| 97 | mt-Nd4 | na | 56093 | -0.666 | -0.1532 | Yes |
| 98 | Uqcrb | na | 56105 | -0.674 | -0.1295 | Yes |
| 99 | mt-Nd1 | na | 56149 | -0.704 | -0.1053 | Yes |
| 100 | Ndufb10 | na | 56154 | -0.708 | -0.0803 | Yes |
| 101 | Ndufb3 | na | 56200 | -0.754 | -0.0545 | Yes |
| 102 | mt-Cytb | na | 56229 | -0.796 | -0.0268 | Yes |
| 103 | Ndufa5 | na | 56251 | -0.830 | 0.0022 | Yes |

