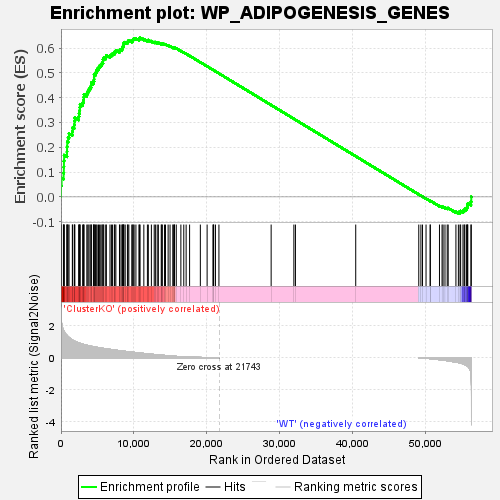
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___ClusterKO_vs_WT.RPKM_matrix___ClusterKO_vs_WT.cls #ClusterKO_versus_WT |
| Phenotype | RPKM_matrix___ClusterKO_vs_WT.cls#ClusterKO_versus_WT |
| Upregulated in class | ClusterKO |
| GeneSet | WP_ADIPOGENESIS_GENES |
| Enrichment Score (ES) | 0.64129007 |
| Normalized Enrichment Score (NES) | 1.4126287 |
| Nominal p-value | 0.015717093 |
| FDR q-value | 0.08413593 |
| FWER p-Value | 0.413 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Lpl | na | 1 | 3.125 | 0.0459 | Yes |
| 2 | Gdf10 | na | 83 | 2.171 | 0.0763 | Yes |
| 3 | Tgfb1 | na | 354 | 1.708 | 0.0966 | Yes |
| 4 | Smad3 | na | 389 | 1.673 | 0.1206 | Yes |
| 5 | Frzb | na | 402 | 1.665 | 0.1448 | Yes |
| 6 | Pck2 | na | 430 | 1.639 | 0.1684 | Yes |
| 7 | Mef2c | na | 822 | 1.392 | 0.1819 | Yes |
| 8 | Socs1 | na | 832 | 1.384 | 0.2020 | Yes |
| 9 | Egr2 | na | 858 | 1.372 | 0.2217 | Yes |
| 10 | Klf6 | na | 975 | 1.320 | 0.2391 | Yes |
| 11 | Wwtr1 | na | 1104 | 1.263 | 0.2553 | Yes |
| 12 | Gata2 | na | 1557 | 1.114 | 0.2637 | Yes |
| 13 | Hmga1 | na | 1590 | 1.101 | 0.2793 | Yes |
| 14 | Tnf | na | 1824 | 1.040 | 0.2904 | Yes |
| 15 | Lmna | na | 1861 | 1.033 | 0.3049 | Yes |
| 16 | Klf5 | na | 1899 | 1.027 | 0.3193 | Yes |
| 17 | Irs1 | na | 2432 | 0.928 | 0.3235 | Yes |
| 18 | Ctnnb1 | na | 2482 | 0.919 | 0.3361 | Yes |
| 19 | Rbl1 | na | 2558 | 0.906 | 0.3481 | Yes |
| 20 | Klf7 | na | 2581 | 0.902 | 0.3610 | Yes |
| 21 | Il6 | na | 2624 | 0.896 | 0.3734 | Yes |
| 22 | Ncoa2 | na | 2974 | 0.845 | 0.3796 | Yes |
| 23 | Nsg1 | na | 3081 | 0.833 | 0.3899 | Yes |
| 24 | Lpin3 | na | 3115 | 0.829 | 0.4015 | Yes |
| 25 | Mef2a | na | 3160 | 0.823 | 0.4128 | Yes |
| 26 | Stat1 | na | 3549 | 0.778 | 0.4173 | Yes |
| 27 | Celf1 | na | 3685 | 0.760 | 0.4261 | Yes |
| 28 | Fzd1 | na | 3829 | 0.745 | 0.4345 | Yes |
| 29 | Lif | na | 4001 | 0.727 | 0.4421 | Yes |
| 30 | Creb1 | na | 4142 | 0.715 | 0.4501 | Yes |
| 31 | Nrip1 | na | 4164 | 0.713 | 0.4602 | Yes |
| 32 | Serpine1 | na | 4454 | 0.688 | 0.4652 | Yes |
| 33 | Mbnl1 | na | 4536 | 0.680 | 0.4738 | Yes |
| 34 | Ebf1 | na | 4546 | 0.679 | 0.4836 | Yes |
| 35 | Nr3c1 | na | 4548 | 0.679 | 0.4935 | Yes |
| 36 | Hif1a | na | 4747 | 0.663 | 0.4997 | Yes |
| 37 | Slc2a4 | na | 4833 | 0.656 | 0.5079 | Yes |
| 38 | Mef2d | na | 4961 | 0.646 | 0.5151 | Yes |
| 39 | Ptgis | na | 5150 | 0.634 | 0.5211 | Yes |
| 40 | Sp1 | na | 5273 | 0.623 | 0.5280 | Yes |
| 41 | Bmp2 | na | 5462 | 0.606 | 0.5336 | Yes |
| 42 | Osm | na | 5622 | 0.593 | 0.5395 | Yes |
| 43 | Stat6 | na | 5765 | 0.583 | 0.5455 | Yes |
| 44 | Zmpste24 | na | 5766 | 0.583 | 0.5541 | Yes |
| 45 | Stat3 | na | 5875 | 0.576 | 0.5606 | Yes |
| 46 | Il6st | na | 6147 | 0.556 | 0.5640 | Yes |
| 47 | Ahr | na | 6228 | 0.551 | 0.5706 | Yes |
| 48 | Ddit3 | na | 6728 | 0.520 | 0.5694 | Yes |
| 49 | Ncoa1 | na | 6924 | 0.507 | 0.5734 | Yes |
| 50 | Rb1 | na | 7082 | 0.497 | 0.5779 | Yes |
| 51 | Gadd45b | na | 7332 | 0.480 | 0.5805 | Yes |
| 52 | Klf15 | na | 7389 | 0.476 | 0.5865 | Yes |
| 53 | Foxo1 | na | 7535 | 0.468 | 0.5908 | Yes |
| 54 | Srebf1 | na | 8031 | 0.441 | 0.5885 | Yes |
| 55 | Pck1 | na | 8104 | 0.437 | 0.5936 | Yes |
| 56 | Cyp26a1 | na | 8367 | 0.422 | 0.5951 | Yes |
| 57 | Nampt | na | 8417 | 0.419 | 0.6004 | Yes |
| 58 | Rora | na | 8466 | 0.416 | 0.6057 | Yes |
| 59 | Rxra | na | 8587 | 0.411 | 0.6096 | Yes |
| 60 | Gata3 | na | 8607 | 0.409 | 0.6153 | Yes |
| 61 | Nr2f1 | na | 8625 | 0.408 | 0.6209 | Yes |
| 62 | Ncor2 | na | 8815 | 0.397 | 0.6234 | Yes |
| 63 | Bmp4 | na | 9123 | 0.379 | 0.6235 | Yes |
| 64 | E2f1 | na | 9190 | 0.376 | 0.6279 | Yes |
| 65 | Rbl2 | na | 9318 | 0.369 | 0.6310 | Yes |
| 66 | E2f4 | na | 9723 | 0.348 | 0.6290 | Yes |
| 67 | Fabp4 | na | 9848 | 0.341 | 0.6318 | Yes |
| 68 | Tle3 | na | 9918 | 0.338 | 0.6355 | Yes |
| 69 | Cebpd | na | 10040 | 0.332 | 0.6382 | Yes |
| 70 | Trib3 | na | 10290 | 0.319 | 0.6385 | Yes |
| 71 | Stat5b | na | 10711 | 0.296 | 0.6354 | Yes |
| 72 | Dlk1 | na | 10768 | 0.293 | 0.6387 | Yes |
| 73 | Gata4 | na | 10860 | 0.287 | 0.6413 | Yes |
| 74 | Foxc2 | na | 11377 | 0.264 | 0.6360 | No |
| 75 | Cyp26b1 | na | 11867 | 0.239 | 0.6308 | No |
| 76 | Stat5a | na | 11993 | 0.233 | 0.6320 | No |
| 77 | Ppargc1a | na | 12413 | 0.214 | 0.6277 | No |
| 78 | Igf1 | na | 12802 | 0.197 | 0.6237 | No |
| 79 | Cebpa | na | 12961 | 0.190 | 0.6237 | No |
| 80 | Id3 | na | 13211 | 0.180 | 0.6219 | No |
| 81 | Socs3 | na | 13373 | 0.172 | 0.6216 | No |
| 82 | Retn | na | 13766 | 0.156 | 0.6169 | No |
| 83 | Nr1h3 | na | 13838 | 0.153 | 0.6179 | No |
| 84 | Sfrp4 | na | 13914 | 0.151 | 0.6188 | No |
| 85 | Dvl1 | na | 14211 | 0.139 | 0.6155 | No |
| 86 | Ncor1 | na | 14350 | 0.133 | 0.6150 | No |
| 87 | Ndn | na | 14713 | 0.118 | 0.6103 | No |
| 88 | Plin1 | na | 14978 | 0.106 | 0.6072 | No |
| 89 | Cfd | na | 15364 | 0.094 | 0.6017 | No |
| 90 | Adipoq | na | 15425 | 0.092 | 0.6020 | No |
| 91 | Ppara | na | 15574 | 0.087 | 0.6006 | No |
| 92 | Cisd1 | na | 15588 | 0.087 | 0.6017 | No |
| 93 | Rxrg | na | 15831 | 0.078 | 0.5985 | No |
| 94 | Stat2 | na | 16464 | 0.057 | 0.5881 | No |
| 95 | Irs3 | na | 16862 | 0.046 | 0.5818 | No |
| 96 | Scd1 | na | 17187 | 0.039 | 0.5766 | No |
| 97 | Cebpb | na | 17662 | 0.031 | 0.5686 | No |
| 98 | Lep | na | 19138 | 0.014 | 0.5426 | No |
| 99 | Mef2b | na | 20068 | 0.007 | 0.5262 | No |
| 100 | Wnt1 | na | 20868 | 0.004 | 0.5120 | No |
| 101 | Gh | na | 20954 | 0.003 | 0.5106 | No |
| 102 | Ins2 | na | 20966 | 0.003 | 0.5104 | No |
| 103 | Wnt10b | na | 21213 | 0.002 | 0.5061 | No |
| 104 | Spock1 | na | 21690 | 0.001 | 0.4976 | No |
| 105 | Ins1 | na | 28862 | 0.000 | 0.3701 | No |
| 106 | Irs4 | na | 31984 | 0.000 | 0.3146 | No |
| 107 | Mixl1 | na | 32199 | 0.000 | 0.3108 | No |
| 108 | Agrp | na | 40479 | 0.000 | 0.1636 | No |
| 109 | Irs2 | na | 49138 | -0.001 | 0.0097 | No |
| 110 | Rara | na | 49432 | -0.010 | 0.0047 | No |
| 111 | Ucp1 | na | 49653 | -0.017 | 0.0010 | No |
| 112 | Prlr | na | 50136 | -0.035 | -0.0071 | No |
| 113 | Lifr | na | 50705 | -0.059 | -0.0163 | No |
| 114 | Fas | na | 50720 | -0.059 | -0.0157 | No |
| 115 | Lpin1 | na | 51981 | -0.123 | -0.0363 | No |
| 116 | Epas1 | na | 52335 | -0.144 | -0.0404 | No |
| 117 | Gadd45a | na | 52474 | -0.152 | -0.0407 | No |
| 118 | Lpin2 | na | 52737 | -0.168 | -0.0428 | No |
| 119 | Agt | na | 53043 | -0.186 | -0.0455 | No |
| 120 | Bscl2 | na | 53178 | -0.194 | -0.0451 | No |
| 121 | Bmp1 | na | 54240 | -0.279 | -0.0598 | No |
| 122 | Ppard | na | 54578 | -0.307 | -0.0613 | No |
| 123 | Agpat2 | na | 54800 | -0.330 | -0.0604 | No |
| 124 | Hnf1a | na | 54889 | -0.341 | -0.0570 | No |
| 125 | Bmp3 | na | 55209 | -0.387 | -0.0570 | No |
| 126 | Wnt5b | na | 55377 | -0.414 | -0.0539 | No |
| 127 | Plin2 | na | 55454 | -0.427 | -0.0489 | No |
| 128 | Mif | na | 55679 | -0.480 | -0.0459 | No |
| 129 | Pnpla3 | na | 55775 | -0.507 | -0.0401 | No |
| 130 | Lipe | na | 55823 | -0.520 | -0.0333 | No |
| 131 | Pparg | na | 55873 | -0.536 | -0.0263 | No |
| 132 | Cntfr | na | 56302 | -1.004 | -0.0192 | No |
| 133 | Twist1 | na | 56349 | -1.395 | 0.0005 | No |

