Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___ClusterKO_vs_WT.RPKM_matrix___ClusterKO_vs_WT.cls #ClusterKO_versus_WT |
| Phenotype | RPKM_matrix___ClusterKO_vs_WT.cls#ClusterKO_versus_WT |
| Upregulated in class | WT |
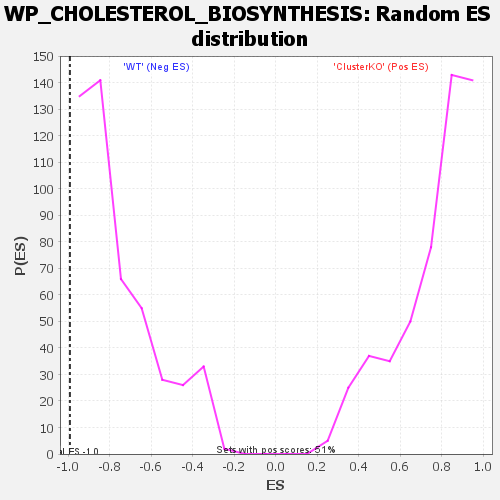
| GeneSet | WP_CHOLESTEROL_BIOSYNTHESIS |
| Enrichment Score (ES) | -0.99169666 |
| Normalized Enrichment Score (NES) | -1.2974671 |
| Nominal p-value | 0.002057613 |
| FDR q-value | 0.3022095 |
| FWER p-Value | 0.775 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Hmgcr | na | 55895 | -0.547 | -0.9630 | Yes |
| 2 | Sc5d | na | 56117 | -0.682 | -0.9312 | Yes |
| 3 | Idi1 | na | 56164 | -0.723 | -0.8941 | Yes |
| 4 | Mvk | na | 56287 | -0.947 | -0.8467 | Yes |
| 5 | Sqle | na | 56298 | -0.989 | -0.7951 | Yes |
| 6 | Hmgcs1 | na | 56306 | -1.018 | -0.7419 | Yes |
| 7 | Cyp51 | na | 56334 | -1.221 | -0.6784 | Yes |
| 8 | Msmo1 | na | 56341 | -1.284 | -0.6112 | Yes |
| 9 | Dhcr7 | na | 56348 | -1.365 | -0.5398 | Yes |
| 10 | Mvd | na | 56353 | -1.438 | -0.4646 | Yes |
| 11 | Fdft1 | na | 56358 | -1.524 | -0.3848 | Yes |
| 12 | Pmvk | na | 56362 | -1.649 | -0.2985 | Yes |
| 13 | Lss | na | 56367 | -1.723 | -0.2083 | Yes |
| 14 | Nsdhl | na | 56370 | -1.910 | -0.1083 | Yes |
| 15 | Fdps | na | 56372 | -2.069 | 0.0001 | Yes |