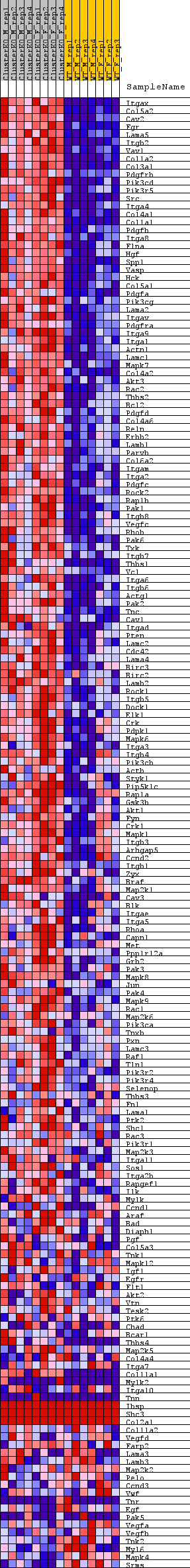
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___ClusterKO_vs_WT.RPKM_matrix___ClusterKO_vs_WT.cls #ClusterKO_versus_WT |
| Phenotype | RPKM_matrix___ClusterKO_vs_WT.cls#ClusterKO_versus_WT |
| Upregulated in class | ClusterKO |
| GeneSet | WP_FOCAL_ADHESION |
| Enrichment Score (ES) | 0.766532 |
| Normalized Enrichment Score (NES) | 1.4171457 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.08316219 |
| FWER p-Value | 0.392 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Itgax | na | 32 | 2.458 | 0.0162 | Yes |
| 2 | Col5a2 | na | 39 | 2.372 | 0.0323 | Yes |
| 3 | Cav2 | na | 99 | 2.114 | 0.0456 | Yes |
| 4 | Fgr | na | 105 | 2.097 | 0.0599 | Yes |
| 5 | Lama5 | na | 109 | 2.084 | 0.0740 | Yes |
| 6 | Itgb2 | na | 150 | 1.982 | 0.0868 | Yes |
| 7 | Vav1 | na | 153 | 1.976 | 0.1003 | Yes |
| 8 | Col1a2 | na | 157 | 1.971 | 0.1137 | Yes |
| 9 | Col3a1 | na | 165 | 1.951 | 0.1269 | Yes |
| 10 | Pdgfrb | na | 189 | 1.906 | 0.1395 | Yes |
| 11 | Pik3cd | na | 204 | 1.888 | 0.1521 | Yes |
| 12 | Pik3r5 | na | 246 | 1.833 | 0.1639 | Yes |
| 13 | Src | na | 302 | 1.762 | 0.1749 | Yes |
| 14 | Itga4 | na | 319 | 1.739 | 0.1865 | Yes |
| 15 | Col4a1 | na | 348 | 1.711 | 0.1977 | Yes |
| 16 | Col1a1 | na | 355 | 1.707 | 0.2092 | Yes |
| 17 | Pdgfb | na | 379 | 1.688 | 0.2203 | Yes |
| 18 | Itga8 | na | 383 | 1.682 | 0.2317 | Yes |
| 19 | Flna | na | 384 | 1.682 | 0.2432 | Yes |
| 20 | Hgf | na | 447 | 1.629 | 0.2532 | Yes |
| 21 | Spp1 | na | 459 | 1.624 | 0.2641 | Yes |
| 22 | Vasp | na | 462 | 1.622 | 0.2751 | Yes |
| 23 | Hck | na | 468 | 1.617 | 0.2861 | Yes |
| 24 | Col5a1 | na | 566 | 1.533 | 0.2948 | Yes |
| 25 | Pdgfa | na | 587 | 1.517 | 0.3048 | Yes |
| 26 | Pik3cg | na | 599 | 1.509 | 0.3149 | Yes |
| 27 | Lama2 | na | 614 | 1.501 | 0.3249 | Yes |
| 28 | Itgav | na | 635 | 1.489 | 0.3347 | Yes |
| 29 | Pdgfra | na | 661 | 1.468 | 0.3443 | Yes |
| 30 | Itga9 | na | 664 | 1.463 | 0.3542 | Yes |
| 31 | Itgal | na | 679 | 1.456 | 0.3639 | Yes |
| 32 | Actn1 | na | 710 | 1.443 | 0.3732 | Yes |
| 33 | Lamc1 | na | 742 | 1.428 | 0.3824 | Yes |
| 34 | Mapk7 | na | 743 | 1.428 | 0.3922 | Yes |
| 35 | Col4a2 | na | 788 | 1.409 | 0.4010 | Yes |
| 36 | Akt3 | na | 833 | 1.383 | 0.4096 | Yes |
| 37 | Rac2 | na | 840 | 1.378 | 0.4189 | Yes |
| 38 | Thbs2 | na | 965 | 1.323 | 0.4258 | Yes |
| 39 | Bcl2 | na | 993 | 1.312 | 0.4342 | Yes |
| 40 | Pdgfd | na | 1042 | 1.289 | 0.4422 | Yes |
| 41 | Col4a6 | na | 1043 | 1.288 | 0.4510 | Yes |
| 42 | Reln | na | 1055 | 1.284 | 0.4595 | Yes |
| 43 | Erbb2 | na | 1098 | 1.266 | 0.4674 | Yes |
| 44 | Lamb1 | na | 1101 | 1.265 | 0.4760 | Yes |
| 45 | Parvb | na | 1148 | 1.247 | 0.4837 | Yes |
| 46 | Col6a2 | na | 1152 | 1.245 | 0.4921 | Yes |
| 47 | Itgam | na | 1194 | 1.232 | 0.4998 | Yes |
| 48 | Itga2 | na | 1268 | 1.202 | 0.5067 | Yes |
| 49 | Pdgfc | na | 1375 | 1.170 | 0.5128 | Yes |
| 50 | Rock2 | na | 1380 | 1.169 | 0.5207 | Yes |
| 51 | Rap1b | na | 1480 | 1.138 | 0.5267 | Yes |
| 52 | Pak1 | na | 1521 | 1.124 | 0.5337 | Yes |
| 53 | Itgb8 | na | 1571 | 1.108 | 0.5404 | Yes |
| 54 | Vegfc | na | 1597 | 1.099 | 0.5474 | Yes |
| 55 | Rhob | na | 1636 | 1.088 | 0.5542 | Yes |
| 56 | Pak6 | na | 1700 | 1.069 | 0.5603 | Yes |
| 57 | Txk | na | 1717 | 1.065 | 0.5673 | Yes |
| 58 | Itgb7 | na | 1742 | 1.058 | 0.5741 | Yes |
| 59 | Thbs1 | na | 1825 | 1.040 | 0.5797 | Yes |
| 60 | Vcl | na | 1845 | 1.036 | 0.5865 | Yes |
| 61 | Itga6 | na | 1896 | 1.027 | 0.5926 | Yes |
| 62 | Itgb6 | na | 1906 | 1.026 | 0.5994 | Yes |
| 63 | Actg1 | na | 1912 | 1.024 | 0.6063 | Yes |
| 64 | Pak2 | na | 1980 | 1.010 | 0.6120 | Yes |
| 65 | Tnc | na | 2017 | 1.005 | 0.6182 | Yes |
| 66 | Cav1 | na | 2033 | 1.002 | 0.6248 | Yes |
| 67 | Itgad | na | 2307 | 0.950 | 0.6264 | Yes |
| 68 | Pten | na | 2392 | 0.935 | 0.6313 | Yes |
| 69 | Lamc2 | na | 2532 | 0.910 | 0.6351 | Yes |
| 70 | Cdc42 | na | 2563 | 0.906 | 0.6407 | Yes |
| 71 | Lama4 | na | 2636 | 0.895 | 0.6455 | Yes |
| 72 | Birc3 | na | 2798 | 0.869 | 0.6486 | Yes |
| 73 | Birc2 | na | 2809 | 0.868 | 0.6543 | Yes |
| 74 | Lamb2 | na | 2811 | 0.867 | 0.6602 | Yes |
| 75 | Rock1 | na | 2989 | 0.843 | 0.6628 | Yes |
| 76 | Itgb5 | na | 3008 | 0.841 | 0.6682 | Yes |
| 77 | Dock1 | na | 3138 | 0.825 | 0.6716 | Yes |
| 78 | Elk1 | na | 3181 | 0.820 | 0.6764 | Yes |
| 79 | Crk | na | 3328 | 0.804 | 0.6793 | Yes |
| 80 | Pdpk1 | na | 3543 | 0.778 | 0.6808 | Yes |
| 81 | Mapk6 | na | 3594 | 0.771 | 0.6852 | Yes |
| 82 | Itga3 | na | 3877 | 0.740 | 0.6852 | Yes |
| 83 | Itgb4 | na | 4008 | 0.726 | 0.6879 | Yes |
| 84 | Pik3cb | na | 4079 | 0.720 | 0.6915 | Yes |
| 85 | Actb | na | 4096 | 0.719 | 0.6961 | Yes |
| 86 | Styk1 | na | 4227 | 0.709 | 0.6987 | Yes |
| 87 | Pip5k1c | na | 4468 | 0.686 | 0.6991 | Yes |
| 88 | Rap1a | na | 4556 | 0.678 | 0.7022 | Yes |
| 89 | Gsk3b | na | 4576 | 0.677 | 0.7064 | Yes |
| 90 | Akt1 | na | 4700 | 0.667 | 0.7088 | Yes |
| 91 | Fyn | na | 4734 | 0.665 | 0.7127 | Yes |
| 92 | Crkl | na | 4761 | 0.662 | 0.7168 | Yes |
| 93 | Mapk1 | na | 4864 | 0.654 | 0.7194 | Yes |
| 94 | Itgb3 | na | 5022 | 0.642 | 0.7210 | Yes |
| 95 | Arhgap5 | na | 5129 | 0.635 | 0.7235 | Yes |
| 96 | Ccnd2 | na | 5351 | 0.616 | 0.7238 | Yes |
| 97 | Itgb1 | na | 5629 | 0.593 | 0.7229 | Yes |
| 98 | Zyx | na | 5666 | 0.589 | 0.7262 | Yes |
| 99 | Braf | na | 5688 | 0.588 | 0.7299 | Yes |
| 100 | Map2k1 | na | 5879 | 0.576 | 0.7304 | Yes |
| 101 | Cav3 | na | 5922 | 0.573 | 0.7336 | Yes |
| 102 | Blk | na | 6100 | 0.560 | 0.7343 | Yes |
| 103 | Itgae | na | 6250 | 0.549 | 0.7354 | Yes |
| 104 | Itga5 | na | 6368 | 0.543 | 0.7370 | Yes |
| 105 | Rhoa | na | 6434 | 0.538 | 0.7395 | Yes |
| 106 | Capn1 | na | 6449 | 0.537 | 0.7429 | Yes |
| 107 | Met | na | 6487 | 0.535 | 0.7459 | Yes |
| 108 | Ppp1r12a | na | 6603 | 0.527 | 0.7474 | Yes |
| 109 | Grb2 | na | 6753 | 0.518 | 0.7483 | Yes |
| 110 | Pak3 | na | 6770 | 0.517 | 0.7516 | Yes |
| 111 | Mapk8 | na | 6857 | 0.511 | 0.7535 | Yes |
| 112 | Jun | na | 7021 | 0.500 | 0.7540 | Yes |
| 113 | Pak4 | na | 7080 | 0.497 | 0.7564 | Yes |
| 114 | Mapk9 | na | 7275 | 0.484 | 0.7563 | Yes |
| 115 | Rac1 | na | 7279 | 0.484 | 0.7595 | Yes |
| 116 | Map2k6 | na | 7489 | 0.471 | 0.7590 | Yes |
| 117 | Pik3ca | na | 7719 | 0.459 | 0.7580 | Yes |
| 118 | Tnxb | na | 7807 | 0.453 | 0.7596 | Yes |
| 119 | Pxn | na | 7836 | 0.452 | 0.7622 | Yes |
| 120 | Lamc3 | na | 7884 | 0.450 | 0.7644 | Yes |
| 121 | Raf1 | na | 7937 | 0.447 | 0.7665 | Yes |
| 122 | Tln1 | na | 8443 | 0.417 | 0.7604 | No |
| 123 | Pik3r2 | na | 8508 | 0.414 | 0.7621 | No |
| 124 | Pik3r4 | na | 9006 | 0.386 | 0.7559 | No |
| 125 | Selenop | na | 9044 | 0.384 | 0.7578 | No |
| 126 | Thbs3 | na | 9255 | 0.372 | 0.7566 | No |
| 127 | Fn1 | na | 9433 | 0.364 | 0.7560 | No |
| 128 | Lama1 | na | 9493 | 0.361 | 0.7574 | No |
| 129 | Ptk2 | na | 9615 | 0.354 | 0.7576 | No |
| 130 | Shc1 | na | 9636 | 0.353 | 0.7597 | No |
| 131 | Rac3 | na | 9686 | 0.350 | 0.7612 | No |
| 132 | Pik3r1 | na | 9928 | 0.337 | 0.7592 | No |
| 133 | Map2k3 | na | 9973 | 0.335 | 0.7607 | No |
| 134 | Itga11 | na | 10240 | 0.322 | 0.7582 | No |
| 135 | Sos1 | na | 10413 | 0.312 | 0.7573 | No |
| 136 | Itga2b | na | 10516 | 0.306 | 0.7575 | No |
| 137 | Rapgef1 | na | 11232 | 0.271 | 0.7467 | No |
| 138 | Ilk | na | 11245 | 0.270 | 0.7483 | No |
| 139 | Mylk | na | 11354 | 0.264 | 0.7482 | No |
| 140 | Ccnd1 | na | 11359 | 0.264 | 0.7499 | No |
| 141 | Araf | na | 11645 | 0.251 | 0.7465 | No |
| 142 | Bad | na | 12212 | 0.223 | 0.7380 | No |
| 143 | Diaph1 | na | 12322 | 0.219 | 0.7375 | No |
| 144 | Pgf | na | 12345 | 0.218 | 0.7386 | No |
| 145 | Col5a3 | na | 12373 | 0.217 | 0.7396 | No |
| 146 | Tnk1 | na | 12386 | 0.216 | 0.7409 | No |
| 147 | Mapk12 | na | 12763 | 0.199 | 0.7356 | No |
| 148 | Igf1 | na | 12802 | 0.197 | 0.7362 | No |
| 149 | Egfr | na | 13342 | 0.174 | 0.7278 | No |
| 150 | Flt1 | na | 13399 | 0.171 | 0.7280 | No |
| 151 | Akt2 | na | 13947 | 0.149 | 0.7193 | No |
| 152 | Vtn | na | 14190 | 0.140 | 0.7159 | No |
| 153 | Tesk2 | na | 14209 | 0.139 | 0.7166 | No |
| 154 | Ptk6 | na | 14458 | 0.129 | 0.7130 | No |
| 155 | Chad | na | 14822 | 0.113 | 0.7073 | No |
| 156 | Bcar1 | na | 15163 | 0.100 | 0.7020 | No |
| 157 | Thbs4 | na | 16051 | 0.071 | 0.6867 | No |
| 158 | Map2k5 | na | 16266 | 0.063 | 0.6833 | No |
| 159 | Col4a4 | na | 16618 | 0.052 | 0.6774 | No |
| 160 | Itga7 | na | 17401 | 0.035 | 0.6637 | No |
| 161 | Col11a1 | na | 19523 | 0.011 | 0.6261 | No |
| 162 | Mylk2 | na | 19870 | 0.009 | 0.6200 | No |
| 163 | Itga10 | na | 19999 | 0.008 | 0.6177 | No |
| 164 | Tnn | na | 21568 | 0.001 | 0.5898 | No |
| 165 | Ibsp | na | 34077 | 0.000 | 0.3672 | No |
| 166 | Shc3 | na | 36405 | 0.000 | 0.3258 | No |
| 167 | Col2a1 | na | 37874 | 0.000 | 0.2997 | No |
| 168 | Col11a2 | na | 49106 | -0.000 | 0.0998 | No |
| 169 | Vegfd | na | 49767 | -0.020 | 0.0882 | No |
| 170 | Farp2 | na | 49883 | -0.025 | 0.0864 | No |
| 171 | Lama3 | na | 50656 | -0.057 | 0.0730 | No |
| 172 | Lamb3 | na | 50887 | -0.067 | 0.0694 | No |
| 173 | Map2k2 | na | 51670 | -0.105 | 0.0562 | No |
| 174 | Pelo | na | 52073 | -0.128 | 0.0499 | No |
| 175 | Ccnd3 | na | 52353 | -0.146 | 0.0459 | No |
| 176 | Vwf | na | 53218 | -0.197 | 0.0319 | No |
| 177 | Tnr | na | 53279 | -0.200 | 0.0322 | No |
| 178 | Egf | na | 53335 | -0.204 | 0.0326 | No |
| 179 | Pak5 | na | 53452 | -0.213 | 0.0320 | No |
| 180 | Vegfa | na | 54197 | -0.276 | 0.0206 | No |
| 181 | Vegfb | na | 54402 | -0.293 | 0.0190 | No |
| 182 | Tnk2 | na | 54625 | -0.313 | 0.0172 | No |
| 183 | Myl6 | na | 54873 | -0.339 | 0.0151 | No |
| 184 | Mapk4 | na | 56230 | -0.796 | -0.0036 | No |
| 185 | Srms | na | 56275 | -0.907 | 0.0018 | No |