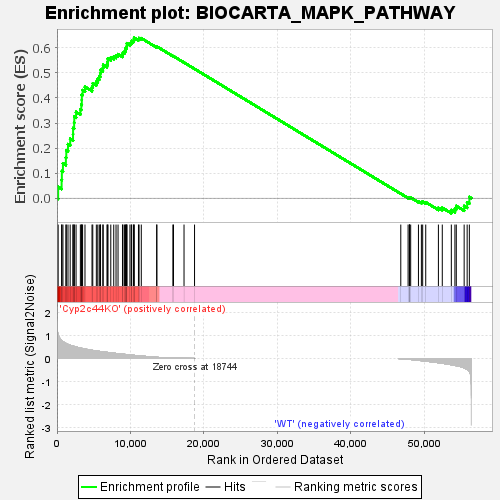
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___Cyp2c44KO_vs_WT.RPKM_matrix___Cyp2c44KO_vs_WT.cls #Cyp2c44KO_versus_WT |
| Phenotype | RPKM_matrix___Cyp2c44KO_vs_WT.cls#Cyp2c44KO_versus_WT |
| Upregulated in class | Cyp2c44KO |
| GeneSet | BIOCARTA_MAPK_PATHWAY |
| Enrichment Score (ES) | 0.6401837 |
| Normalized Enrichment Score (NES) | 1.2807772 |
| Nominal p-value | 0.070866145 |
| FDR q-value | 0.51590776 |
| FWER p-Value | 0.661 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Jun | na | 157 | 1.020 | 0.0450 | Yes |
| 2 | Mapk11 | na | 627 | 0.772 | 0.0729 | Yes |
| 3 | Map4k1 | na | 660 | 0.764 | 0.1081 | Yes |
| 4 | Map3k6 | na | 808 | 0.721 | 0.1393 | Yes |
| 5 | Map2k7 | na | 1210 | 0.647 | 0.1625 | Yes |
| 6 | Map3k5 | na | 1264 | 0.636 | 0.1914 | Yes |
| 7 | Map4k2 | na | 1495 | 0.604 | 0.2156 | Yes |
| 8 | Map3k2 | na | 1794 | 0.570 | 0.2370 | Yes |
| 9 | Nfkbia | na | 2181 | 0.530 | 0.2550 | Yes |
| 10 | Fos | na | 2187 | 0.529 | 0.2797 | Yes |
| 11 | Elk1 | na | 2333 | 0.517 | 0.3014 | Yes |
| 12 | Mapk12 | na | 2353 | 0.516 | 0.3252 | Yes |
| 13 | Map3k8 | na | 2594 | 0.494 | 0.3441 | Yes |
| 14 | Map3k12 | na | 3178 | 0.449 | 0.3548 | Yes |
| 15 | Map4k5 | na | 3306 | 0.442 | 0.3733 | Yes |
| 16 | Mapkapk5 | na | 3372 | 0.435 | 0.3925 | Yes |
| 17 | Mknk2 | na | 3376 | 0.435 | 0.4129 | Yes |
| 18 | Map3k3 | na | 3481 | 0.428 | 0.4311 | Yes |
| 19 | Mapk7 | na | 3808 | 0.408 | 0.4444 | Yes |
| 20 | Mapkapk2 | na | 4765 | 0.353 | 0.4440 | Yes |
| 21 | Tgfb1 | na | 4868 | 0.347 | 0.4584 | Yes |
| 22 | Ikbkb | na | 5353 | 0.323 | 0.4650 | Yes |
| 23 | Creb1 | na | 5549 | 0.315 | 0.4763 | Yes |
| 24 | Rps6ka3 | na | 5758 | 0.305 | 0.4869 | Yes |
| 25 | Atf2 | na | 5899 | 0.299 | 0.4984 | Yes |
| 26 | Rps6ka5 | na | 5925 | 0.298 | 0.5119 | Yes |
| 27 | Daxx | na | 6241 | 0.284 | 0.5197 | Yes |
| 28 | Map4k4 | na | 6282 | 0.282 | 0.5322 | Yes |
| 29 | Nfkb1 | na | 6802 | 0.262 | 0.5353 | Yes |
| 30 | Sp1 | na | 6904 | 0.258 | 0.5456 | Yes |
| 31 | Mapk8 | na | 6919 | 0.258 | 0.5574 | Yes |
| 32 | Max | na | 7319 | 0.243 | 0.5617 | Yes |
| 33 | Rapgef2 | na | 7721 | 0.227 | 0.5652 | Yes |
| 34 | Mknk1 | na | 8033 | 0.216 | 0.5698 | Yes |
| 35 | Pak2 | na | 8306 | 0.207 | 0.5746 | Yes |
| 36 | Rps6ka2 | na | 8911 | 0.185 | 0.5726 | Yes |
| 37 | Chuk | na | 8968 | 0.183 | 0.5802 | Yes |
| 38 | Map3k7 | na | 9173 | 0.176 | 0.5848 | Yes |
| 39 | Map2k1 | na | 9289 | 0.172 | 0.5908 | Yes |
| 40 | Mapk1 | na | 9328 | 0.171 | 0.5982 | Yes |
| 41 | Map4k3 | na | 9456 | 0.166 | 0.6037 | Yes |
| 42 | Rps6kb1 | na | 9503 | 0.165 | 0.6106 | Yes |
| 43 | Mapk3 | na | 9505 | 0.165 | 0.6183 | Yes |
| 44 | Map2k4 | na | 9927 | 0.151 | 0.6179 | Yes |
| 45 | Ripk1 | na | 10109 | 0.144 | 0.6215 | Yes |
| 46 | Mapk9 | na | 10166 | 0.142 | 0.6271 | Yes |
| 47 | Tgfbr1 | na | 10413 | 0.135 | 0.6291 | Yes |
| 48 | Stat1 | na | 10488 | 0.133 | 0.6340 | Yes |
| 49 | Rps6kb2 | na | 10491 | 0.133 | 0.6402 | Yes |
| 50 | Mapk14 | na | 11085 | 0.115 | 0.6350 | No |
| 51 | Tgfb3 | na | 11167 | 0.113 | 0.6389 | No |
| 52 | Map2k5 | na | 11470 | 0.103 | 0.6384 | No |
| 53 | Raf1 | na | 13568 | 0.049 | 0.6034 | No |
| 54 | Rela | na | 13587 | 0.049 | 0.6054 | No |
| 55 | Map3k11 | na | 15770 | 0.016 | 0.5674 | No |
| 56 | Map3k1 | na | 15852 | 0.016 | 0.5667 | No |
| 57 | Map3k4 | na | 17286 | 0.006 | 0.5415 | No |
| 58 | Map3k14 | na | 18717 | 0.000 | 0.5161 | No |
| 59 | Map2k6 | na | 46783 | -0.007 | 0.0180 | No |
| 60 | Grb2 | na | 47778 | -0.030 | 0.0018 | No |
| 61 | Traf2 | na | 47959 | -0.036 | 0.0002 | No |
| 62 | Pak1 | na | 47966 | -0.036 | 0.0018 | No |
| 63 | Rps6ka4 | na | 47968 | -0.036 | 0.0035 | No |
| 64 | Shc1 | na | 48130 | -0.041 | 0.0026 | No |
| 65 | Mapk6 | na | 49186 | -0.075 | -0.0126 | No |
| 66 | Rps6ka1 | na | 49586 | -0.090 | -0.0155 | No |
| 67 | Cebpa | na | 49762 | -0.096 | -0.0141 | No |
| 68 | Map3k10 | na | 50178 | -0.111 | -0.0163 | No |
| 69 | Tgfb2 | na | 51902 | -0.181 | -0.0384 | No |
| 70 | Map2k3 | na | 52430 | -0.205 | -0.0381 | No |
| 71 | Mapk10 | na | 53661 | -0.267 | -0.0475 | No |
| 72 | Map2k2 | na | 54150 | -0.297 | -0.0422 | No |
| 73 | Tradd | na | 54336 | -0.307 | -0.0311 | No |
| 74 | Mapk13 | na | 55406 | -0.401 | -0.0313 | No |
| 75 | Myc | na | 55814 | -0.472 | -0.0164 | No |
| 76 | Hras | na | 56113 | -0.564 | 0.0047 | No |

