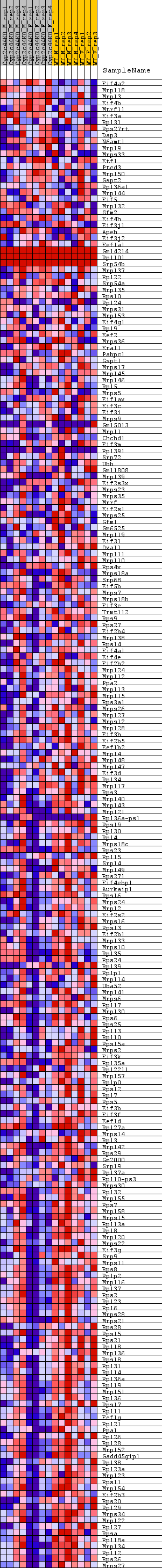
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___Cyp2c44KO_vs_WT.RPKM_matrix___Cyp2c44KO_vs_WT.cls #Cyp2c44KO_versus_WT |
| Phenotype | RPKM_matrix___Cyp2c44KO_vs_WT.cls#Cyp2c44KO_versus_WT |
| Upregulated in class | WT |
| GeneSet | REACTOME_TRANSLATION |
| Enrichment Score (ES) | -0.79955494 |
| Normalized Enrichment Score (NES) | -1.4507757 |
| Nominal p-value | 0.014344262 |
| FDR q-value | 0.16061492 |
| FWER p-Value | 0.637 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Eif4a2 | na | 1320 | 0.628 | -0.0122 | No |
| 2 | Mrpl18 | na | 3489 | 0.428 | -0.0431 | No |
| 3 | Mrpl3 | na | 4459 | 0.370 | -0.0537 | No |
| 4 | Eif4b | na | 5477 | 0.318 | -0.0661 | No |
| 5 | Mtrf1l | na | 6642 | 0.268 | -0.0821 | No |
| 6 | Eif3a | na | 7280 | 0.244 | -0.0890 | No |
| 7 | Rpl3l | na | 7738 | 0.226 | -0.0931 | No |
| 8 | Rps27rt | na | 11007 | 0.117 | -0.1492 | No |
| 9 | Dap3 | na | 11206 | 0.112 | -0.1507 | No |
| 10 | N6amt1 | na | 11589 | 0.100 | -0.1557 | No |
| 11 | Mrpl9 | na | 12512 | 0.075 | -0.1708 | No |
| 12 | Mrps33 | na | 12645 | 0.071 | -0.1718 | No |
| 13 | Etf1 | na | 13007 | 0.062 | -0.1772 | No |
| 14 | Ptcd3 | na | 13209 | 0.057 | -0.1797 | No |
| 15 | Mrpl50 | na | 13360 | 0.054 | -0.1814 | No |
| 16 | Gspt2 | na | 13628 | 0.048 | -0.1853 | No |
| 17 | Rpl36al | na | 13685 | 0.047 | -0.1855 | No |
| 18 | Mrpl44 | na | 13806 | 0.044 | -0.1868 | No |
| 19 | Eif5 | na | 13920 | 0.042 | -0.1881 | No |
| 20 | Mrpl32 | na | 13981 | 0.041 | -0.1884 | No |
| 21 | Gfm2 | na | 14682 | 0.030 | -0.2003 | No |
| 22 | Eif4h | na | 14709 | 0.029 | -0.2003 | No |
| 23 | Eif3j1 | na | 14871 | 0.027 | -0.2026 | No |
| 24 | Apeh | na | 14882 | 0.027 | -0.2023 | No |
| 25 | Eif3j2 | na | 16741 | 0.009 | -0.2353 | No |
| 26 | Eef1a1 | na | 18140 | 0.002 | -0.2601 | No |
| 27 | Gm14214 | na | 25859 | 0.000 | -0.3976 | No |
| 28 | Rpl10l | na | 26075 | 0.000 | -0.4014 | No |
| 29 | Srp54b | na | 44878 | 0.000 | -0.7363 | No |
| 30 | Mrpl37 | na | 47077 | -0.013 | -0.7752 | No |
| 31 | Rpl22 | na | 47702 | -0.028 | -0.7858 | No |
| 32 | Srp54a | na | 47705 | -0.028 | -0.7853 | No |
| 33 | Mrpl35 | na | 47800 | -0.031 | -0.7865 | No |
| 34 | Rps10 | na | 47950 | -0.036 | -0.7885 | No |
| 35 | Rpl24 | na | 48147 | -0.042 | -0.7912 | No |
| 36 | Mrps31 | na | 48239 | -0.045 | -0.7920 | No |
| 37 | Mrpl53 | na | 48398 | -0.049 | -0.7940 | No |
| 38 | Eif4g1 | na | 48470 | -0.051 | -0.7943 | No |
| 39 | Rpl9 | na | 48558 | -0.054 | -0.7949 | No |
| 40 | Eef2 | na | 48664 | -0.057 | -0.7957 | No |
| 41 | Mrps36 | na | 48712 | -0.059 | -0.7955 | No |
| 42 | Eral1 | na | 48746 | -0.060 | -0.7950 | No |
| 43 | Pabpc1 | na | 48788 | -0.062 | -0.7946 | No |
| 44 | Gspt1 | na | 49060 | -0.071 | -0.7982 | Yes |
| 45 | Mrps17 | na | 49074 | -0.072 | -0.7971 | Yes |
| 46 | Mrpl45 | na | 49132 | -0.073 | -0.7968 | Yes |
| 47 | Mrpl46 | na | 49226 | -0.077 | -0.7971 | Yes |
| 48 | Rpl5 | na | 49251 | -0.078 | -0.7961 | Yes |
| 49 | Mrps5 | na | 49338 | -0.081 | -0.7962 | Yes |
| 50 | Eif1ax | na | 49349 | -0.082 | -0.7949 | Yes |
| 51 | Eif3c | na | 49383 | -0.083 | -0.7940 | Yes |
| 52 | Eif3i | na | 49604 | -0.090 | -0.7963 | Yes |
| 53 | Mrps9 | na | 49788 | -0.098 | -0.7978 | Yes |
| 54 | Gm15013 | na | 49822 | -0.098 | -0.7966 | Yes |
| 55 | Mrpl1 | na | 49835 | -0.099 | -0.7951 | Yes |
| 56 | Chchd1 | na | 49840 | -0.099 | -0.7933 | Yes |
| 57 | Eif3m | na | 50008 | -0.105 | -0.7944 | Yes |
| 58 | Rpl39l | na | 50124 | -0.108 | -0.7945 | Yes |
| 59 | Srp72 | na | 50332 | -0.116 | -0.7961 | Yes |
| 60 | Ubb | na | 50333 | -0.116 | -0.7940 | Yes |
| 61 | Gm11808 | na | 50371 | -0.118 | -0.7926 | Yes |
| 62 | Mrpl39 | na | 50447 | -0.121 | -0.7917 | Yes |
| 63 | Eif2s3x | na | 50469 | -0.122 | -0.7899 | Yes |
| 64 | Mrps23 | na | 50472 | -0.122 | -0.7878 | Yes |
| 65 | Mrps35 | na | 50524 | -0.125 | -0.7864 | Yes |
| 66 | Mrrf | na | 50604 | -0.128 | -0.7855 | Yes |
| 67 | Eif2s1 | na | 50794 | -0.135 | -0.7865 | Yes |
| 68 | Mrps25 | na | 50795 | -0.135 | -0.7840 | Yes |
| 69 | Gfm1 | na | 50847 | -0.138 | -0.7825 | Yes |
| 70 | Gm6525 | na | 50887 | -0.139 | -0.7807 | Yes |
| 71 | Mrpl19 | na | 50949 | -0.142 | -0.7792 | Yes |
| 72 | Eif3l | na | 50972 | -0.143 | -0.7770 | Yes |
| 73 | Oxa1l | na | 50988 | -0.143 | -0.7747 | Yes |
| 74 | Mrpl11 | na | 50993 | -0.143 | -0.7722 | Yes |
| 75 | Mrpl10 | na | 51134 | -0.150 | -0.7720 | Yes |
| 76 | Rps4x | na | 51161 | -0.150 | -0.7698 | Yes |
| 77 | Mrps18a | na | 51165 | -0.151 | -0.7671 | Yes |
| 78 | Srp68 | na | 51173 | -0.151 | -0.7645 | Yes |
| 79 | Eif5b | na | 51197 | -0.152 | -0.7622 | Yes |
| 80 | Mrps7 | na | 51240 | -0.154 | -0.7602 | Yes |
| 81 | Mrps18b | na | 51246 | -0.154 | -0.7575 | Yes |
| 82 | Eif3e | na | 51426 | -0.162 | -0.7578 | Yes |
| 83 | Trmt112 | na | 51513 | -0.165 | -0.7563 | Yes |
| 84 | Rps9 | na | 51569 | -0.168 | -0.7543 | Yes |
| 85 | Rps27 | na | 51600 | -0.169 | -0.7518 | Yes |
| 86 | Eif2b4 | na | 51622 | -0.170 | -0.7491 | Yes |
| 87 | Mrpl38 | na | 51717 | -0.173 | -0.7477 | Yes |
| 88 | Rps14 | na | 51824 | -0.177 | -0.7464 | Yes |
| 89 | Eif4a1 | na | 51836 | -0.178 | -0.7434 | Yes |
| 90 | Eif4e | na | 51838 | -0.178 | -0.7402 | Yes |
| 91 | Eif2b2 | na | 51867 | -0.179 | -0.7375 | Yes |
| 92 | Mrpl24 | na | 51923 | -0.182 | -0.7352 | Yes |
| 93 | Mrpl12 | na | 51944 | -0.184 | -0.7322 | Yes |
| 94 | Ppa2 | na | 51967 | -0.185 | -0.7293 | Yes |
| 95 | Mrpl13 | na | 51969 | -0.185 | -0.7260 | Yes |
| 96 | Mrpl15 | na | 52049 | -0.188 | -0.7240 | Yes |
| 97 | Rps3a1 | na | 52270 | -0.198 | -0.7244 | Yes |
| 98 | Mrps26 | na | 52276 | -0.199 | -0.7209 | Yes |
| 99 | Mrpl27 | na | 52330 | -0.200 | -0.7182 | Yes |
| 100 | Mrps12 | na | 52331 | -0.201 | -0.7146 | Yes |
| 101 | Mrpl28 | na | 52368 | -0.202 | -0.7116 | Yes |
| 102 | Eif3h | na | 52428 | -0.205 | -0.7090 | Yes |
| 103 | Eif2b5 | na | 52461 | -0.206 | -0.7059 | Yes |
| 104 | Eef1b2 | na | 52494 | -0.208 | -0.7027 | Yes |
| 105 | Mrpl4 | na | 52521 | -0.209 | -0.6994 | Yes |
| 106 | Mrpl48 | na | 52522 | -0.209 | -0.6956 | Yes |
| 107 | Mrpl47 | na | 52554 | -0.211 | -0.6924 | Yes |
| 108 | Eif3d | na | 52556 | -0.211 | -0.6886 | Yes |
| 109 | Rpl34 | na | 52574 | -0.212 | -0.6851 | Yes |
| 110 | Mrpl17 | na | 52611 | -0.214 | -0.6819 | Yes |
| 111 | Rps3 | na | 52657 | -0.216 | -0.6788 | Yes |
| 112 | Mrpl40 | na | 52662 | -0.216 | -0.6750 | Yes |
| 113 | Mrpl43 | na | 52676 | -0.217 | -0.6714 | Yes |
| 114 | Mrpl21 | na | 52691 | -0.217 | -0.6677 | Yes |
| 115 | Rpl36a-ps1 | na | 52813 | -0.223 | -0.6658 | Yes |
| 116 | Rps19 | na | 52927 | -0.228 | -0.6637 | Yes |
| 117 | Rpl30 | na | 52939 | -0.229 | -0.6598 | Yes |
| 118 | Rpl4 | na | 52991 | -0.232 | -0.6566 | Yes |
| 119 | Mrps18c | na | 53003 | -0.233 | -0.6526 | Yes |
| 120 | Rps23 | na | 53021 | -0.234 | -0.6487 | Yes |
| 121 | Rpl15 | na | 53050 | -0.235 | -0.6449 | Yes |
| 122 | Srp14 | na | 53060 | -0.236 | -0.6409 | Yes |
| 123 | Mrpl49 | na | 53125 | -0.239 | -0.6377 | Yes |
| 124 | Rps27l | na | 53149 | -0.241 | -0.6338 | Yes |
| 125 | Eif4ebp1 | na | 53223 | -0.245 | -0.6307 | Yes |
| 126 | Aurkaip1 | na | 53233 | -0.245 | -0.6264 | Yes |
| 127 | Rps16 | na | 53255 | -0.247 | -0.6224 | Yes |
| 128 | Mrps24 | na | 53262 | -0.247 | -0.6180 | Yes |
| 129 | Mrpl2 | na | 53317 | -0.250 | -0.6145 | Yes |
| 130 | Eif2s2 | na | 53321 | -0.250 | -0.6100 | Yes |
| 131 | Mrps16 | na | 53334 | -0.251 | -0.6057 | Yes |
| 132 | Rps13 | na | 53366 | -0.253 | -0.6018 | Yes |
| 133 | Eif2b1 | na | 53373 | -0.253 | -0.5973 | Yes |
| 134 | Mrpl33 | na | 53384 | -0.253 | -0.5929 | Yes |
| 135 | Mrps10 | na | 53389 | -0.254 | -0.5884 | Yes |
| 136 | Rpl35 | na | 53464 | -0.257 | -0.5851 | Yes |
| 137 | Rps24 | na | 53532 | -0.259 | -0.5817 | Yes |
| 138 | Rpl39 | na | 53569 | -0.261 | -0.5776 | Yes |
| 139 | Rplp1 | na | 53652 | -0.266 | -0.5743 | Yes |
| 140 | Mrpl14 | na | 53656 | -0.267 | -0.5696 | Yes |
| 141 | Uba52 | na | 53677 | -0.267 | -0.5651 | Yes |
| 142 | Mrpl41 | na | 53717 | -0.270 | -0.5610 | Yes |
| 143 | Mrps6 | na | 53742 | -0.271 | -0.5565 | Yes |
| 144 | Rpl17 | na | 53765 | -0.273 | -0.5520 | Yes |
| 145 | Mrpl30 | na | 53819 | -0.276 | -0.5480 | Yes |
| 146 | Rps6 | na | 53840 | -0.277 | -0.5433 | Yes |
| 147 | Rps25 | na | 53849 | -0.278 | -0.5385 | Yes |
| 148 | Rpl13 | na | 53916 | -0.283 | -0.5346 | Yes |
| 149 | Rpl10 | na | 53923 | -0.283 | -0.5296 | Yes |
| 150 | Rps15a | na | 53933 | -0.284 | -0.5246 | Yes |
| 151 | Mrps2 | na | 53994 | -0.287 | -0.5205 | Yes |
| 152 | Eif3k | na | 54052 | -0.291 | -0.5163 | Yes |
| 153 | Rpl35a | na | 54054 | -0.291 | -0.5111 | Yes |
| 154 | Rpl22l1 | na | 54058 | -0.291 | -0.5059 | Yes |
| 155 | Mrpl57 | na | 54067 | -0.292 | -0.5008 | Yes |
| 156 | Rplp0 | na | 54071 | -0.292 | -0.4956 | Yes |
| 157 | Rps12 | na | 54110 | -0.295 | -0.4910 | Yes |
| 158 | Rpl7 | na | 54115 | -0.295 | -0.4858 | Yes |
| 159 | Rps5 | na | 54145 | -0.297 | -0.4810 | Yes |
| 160 | Eif3b | na | 54155 | -0.297 | -0.4758 | Yes |
| 161 | Eif3f | na | 54157 | -0.297 | -0.4705 | Yes |
| 162 | Eef1d | na | 54178 | -0.299 | -0.4654 | Yes |
| 163 | Rpl27a | na | 54194 | -0.300 | -0.4603 | Yes |
| 164 | Mrps14 | na | 54287 | -0.305 | -0.4565 | Yes |
| 165 | Rpl3 | na | 54347 | -0.308 | -0.4520 | Yes |
| 166 | Mrpl42 | na | 54525 | -0.320 | -0.4494 | Yes |
| 167 | Rps29 | na | 54540 | -0.321 | -0.4439 | Yes |
| 168 | Gm2000 | na | 54545 | -0.321 | -0.4382 | Yes |
| 169 | Srp19 | na | 54554 | -0.322 | -0.4325 | Yes |
| 170 | Rpl37a | na | 54555 | -0.322 | -0.4267 | Yes |
| 171 | Rpl10-ps3 | na | 54562 | -0.323 | -0.4210 | Yes |
| 172 | Mrps30 | na | 54617 | -0.326 | -0.4161 | Yes |
| 173 | Rpl32 | na | 54674 | -0.330 | -0.4112 | Yes |
| 174 | Mrpl55 | na | 54751 | -0.335 | -0.4065 | Yes |
| 175 | Rps7 | na | 54753 | -0.336 | -0.4005 | Yes |
| 176 | Mrpl58 | na | 54760 | -0.336 | -0.3946 | Yes |
| 177 | Mrps15 | na | 54824 | -0.342 | -0.3896 | Yes |
| 178 | Rpl13a | na | 54829 | -0.342 | -0.3835 | Yes |
| 179 | Rpl8 | na | 54838 | -0.342 | -0.3775 | Yes |
| 180 | Mrpl20 | na | 54848 | -0.343 | -0.3715 | Yes |
| 181 | Mrps22 | na | 54970 | -0.354 | -0.3673 | Yes |
| 182 | Eif3g | na | 54989 | -0.356 | -0.3612 | Yes |
| 183 | Srp9 | na | 55052 | -0.362 | -0.3558 | Yes |
| 184 | Mrps11 | na | 55054 | -0.362 | -0.3493 | Yes |
| 185 | Rps8 | na | 55056 | -0.362 | -0.3428 | Yes |
| 186 | Rplp2 | na | 55110 | -0.369 | -0.3371 | Yes |
| 187 | Mrpl16 | na | 55127 | -0.370 | -0.3307 | Yes |
| 188 | Rpl37 | na | 55144 | -0.371 | -0.3243 | Yes |
| 189 | Rps2 | na | 55187 | -0.376 | -0.3183 | Yes |
| 190 | Rpl23 | na | 55195 | -0.376 | -0.3117 | Yes |
| 191 | Rpl6 | na | 55208 | -0.378 | -0.3051 | Yes |
| 192 | Mrps28 | na | 55235 | -0.381 | -0.2987 | Yes |
| 193 | Mrps21 | na | 55247 | -0.382 | -0.2921 | Yes |
| 194 | Rps28 | na | 55282 | -0.386 | -0.2857 | Yes |
| 195 | Rps15 | na | 55348 | -0.394 | -0.2798 | Yes |
| 196 | Rps21 | na | 55349 | -0.394 | -0.2727 | Yes |
| 197 | Rpl18 | na | 55360 | -0.395 | -0.2658 | Yes |
| 198 | Mrpl36 | na | 55377 | -0.397 | -0.2590 | Yes |
| 199 | Rps18 | na | 55404 | -0.401 | -0.2522 | Yes |
| 200 | Rpl31 | na | 55460 | -0.408 | -0.2459 | Yes |
| 201 | Rpl14 | na | 55467 | -0.409 | -0.2386 | Yes |
| 202 | Rpl36a | na | 55484 | -0.412 | -0.2315 | Yes |
| 203 | Rpl19 | na | 55492 | -0.413 | -0.2242 | Yes |
| 204 | Mrpl51 | na | 55504 | -0.415 | -0.2169 | Yes |
| 205 | Rpl36 | na | 55514 | -0.417 | -0.2096 | Yes |
| 206 | Rps17 | na | 55528 | -0.420 | -0.2022 | Yes |
| 207 | Rpl11 | na | 55553 | -0.425 | -0.1950 | Yes |
| 208 | Eef1g | na | 55556 | -0.425 | -0.1874 | Yes |
| 209 | Rpl21 | na | 55562 | -0.426 | -0.1798 | Yes |
| 210 | Ppa1 | na | 55567 | -0.427 | -0.1722 | Yes |
| 211 | Rpl26 | na | 55596 | -0.431 | -0.1650 | Yes |
| 212 | Rpl28 | na | 55621 | -0.436 | -0.1576 | Yes |
| 213 | Mrpl52 | na | 55625 | -0.437 | -0.1498 | Yes |
| 214 | Gadd45gip1 | na | 55639 | -0.440 | -0.1421 | Yes |
| 215 | Rpl38 | na | 55666 | -0.444 | -0.1346 | Yes |
| 216 | Rpl23a | na | 55684 | -0.448 | -0.1268 | Yes |
| 217 | Mrpl23 | na | 55745 | -0.459 | -0.1197 | Yes |
| 218 | Rps11 | na | 55767 | -0.465 | -0.1117 | Yes |
| 219 | Mrpl54 | na | 55791 | -0.469 | -0.1037 | Yes |
| 220 | Eif2b3 | na | 55834 | -0.476 | -0.0958 | Yes |
| 221 | Rps20 | na | 55837 | -0.476 | -0.0873 | Yes |
| 222 | Rpl29 | na | 55864 | -0.484 | -0.0791 | Yes |
| 223 | Mrps34 | na | 55895 | -0.492 | -0.0708 | Yes |
| 224 | Mrpl22 | na | 55902 | -0.494 | -0.0620 | Yes |
| 225 | Rpl27 | na | 55909 | -0.495 | -0.0532 | Yes |
| 226 | Rpsa | na | 55937 | -0.504 | -0.0446 | Yes |
| 227 | Rpl18a | na | 55998 | -0.519 | -0.0364 | Yes |
| 228 | Mrpl34 | na | 56146 | -0.579 | -0.0286 | Yes |
| 229 | Rpl12 | na | 56147 | -0.579 | -0.0182 | Yes |
| 230 | Rps26 | na | 56163 | -0.588 | -0.0079 | Yes |
| 231 | Mrps27 | na | 56247 | -0.649 | 0.0023 | Yes |