Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___Cyp2c44KO_vs_WT.RPKM_matrix___Cyp2c44KO_vs_WT.cls #Cyp2c44KO_versus_WT |
| Phenotype | RPKM_matrix___Cyp2c44KO_vs_WT.cls#Cyp2c44KO_versus_WT |
| Upregulated in class | Cyp2c44KO |
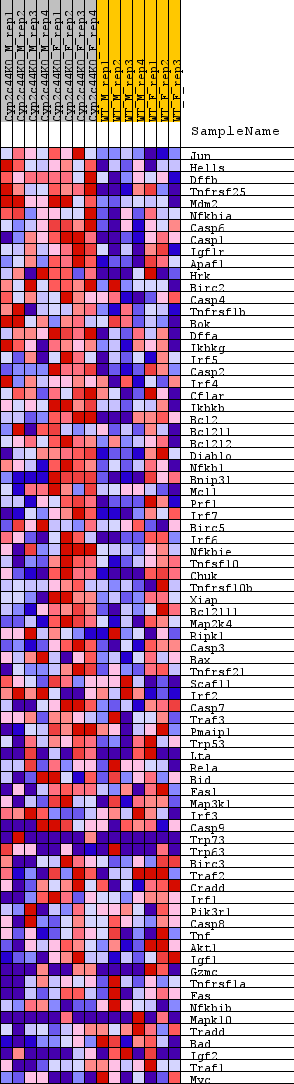
| GeneSet | WP_APOPTOSIS |
| Enrichment Score (ES) | 0.5869394 |
| Normalized Enrichment Score (NES) | 1.2514787 |
| Nominal p-value | 0.11044177 |
| FDR q-value | 0.62142843 |
| FWER p-Value | 0.835 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Jun | na | 157 | 1.020 | 0.0478 | Yes |
| 2 | Hells | na | 445 | 0.839 | 0.0843 | Yes |
| 3 | Dffb | na | 673 | 0.760 | 0.1180 | Yes |
| 4 | Tnfrsf25 | na | 747 | 0.741 | 0.1534 | Yes |
| 5 | Mdm2 | na | 1171 | 0.654 | 0.1783 | Yes |
| 6 | Nfkbia | na | 2181 | 0.530 | 0.1867 | Yes |
| 7 | Casp6 | na | 2417 | 0.511 | 0.2078 | Yes |
| 8 | Casp1 | na | 2580 | 0.496 | 0.2295 | Yes |
| 9 | Igf1r | na | 2850 | 0.473 | 0.2482 | Yes |
| 10 | Apaf1 | na | 2932 | 0.467 | 0.2699 | Yes |
| 11 | Hrk | na | 3217 | 0.447 | 0.2871 | Yes |
| 12 | Birc2 | na | 3307 | 0.442 | 0.3074 | Yes |
| 13 | Casp4 | na | 3439 | 0.431 | 0.3264 | Yes |
| 14 | Tnfrsf1b | na | 3838 | 0.406 | 0.3395 | Yes |
| 15 | Bok | na | 4122 | 0.387 | 0.3537 | Yes |
| 16 | Dffa | na | 4240 | 0.380 | 0.3705 | Yes |
| 17 | Ikbkg | na | 4333 | 0.376 | 0.3875 | Yes |
| 18 | Irf5 | na | 4677 | 0.357 | 0.3991 | Yes |
| 19 | Casp2 | na | 4678 | 0.357 | 0.4168 | Yes |
| 20 | Irf4 | na | 5126 | 0.334 | 0.4254 | Yes |
| 21 | Cflar | na | 5176 | 0.331 | 0.4410 | Yes |
| 22 | Ikbkb | na | 5353 | 0.323 | 0.4539 | Yes |
| 23 | Bcl2 | na | 5726 | 0.307 | 0.4625 | Yes |
| 24 | Bcl2l1 | na | 5830 | 0.302 | 0.4756 | Yes |
| 25 | Bcl2l2 | na | 6184 | 0.287 | 0.4836 | Yes |
| 26 | Diablo | na | 6529 | 0.272 | 0.4910 | Yes |
| 27 | Nfkb1 | na | 6802 | 0.262 | 0.4991 | Yes |
| 28 | Bnip3l | na | 6819 | 0.261 | 0.5118 | Yes |
| 29 | Mcl1 | na | 7125 | 0.249 | 0.5187 | Yes |
| 30 | Prf1 | na | 7324 | 0.242 | 0.5272 | Yes |
| 31 | Irf7 | na | 7744 | 0.226 | 0.5310 | Yes |
| 32 | Birc5 | na | 7790 | 0.225 | 0.5413 | Yes |
| 33 | Irf6 | na | 7861 | 0.222 | 0.5511 | Yes |
| 34 | Nfkbie | na | 8522 | 0.199 | 0.5493 | Yes |
| 35 | Tnfsf10 | na | 8728 | 0.192 | 0.5551 | Yes |
| 36 | Chuk | na | 8968 | 0.183 | 0.5600 | Yes |
| 37 | Tnfrsf10b | na | 9517 | 0.164 | 0.5584 | Yes |
| 38 | Xiap | na | 9546 | 0.163 | 0.5660 | Yes |
| 39 | Bcl2l11 | na | 9702 | 0.158 | 0.5711 | Yes |
| 40 | Map2k4 | na | 9927 | 0.151 | 0.5746 | Yes |
| 41 | Ripk1 | na | 10109 | 0.144 | 0.5785 | Yes |
| 42 | Casp3 | na | 10300 | 0.138 | 0.5820 | Yes |
| 43 | Bax | na | 10594 | 0.130 | 0.5832 | Yes |
| 44 | Tnfrsf21 | na | 10767 | 0.125 | 0.5863 | Yes |
| 45 | Scaf11 | na | 11057 | 0.116 | 0.5869 | Yes |
| 46 | Irf2 | na | 11570 | 0.101 | 0.5828 | No |
| 47 | Casp7 | na | 11829 | 0.094 | 0.5829 | No |
| 48 | Traf3 | na | 11893 | 0.092 | 0.5864 | No |
| 49 | Pmaip1 | na | 12161 | 0.084 | 0.5858 | No |
| 50 | Trp53 | na | 12674 | 0.071 | 0.5802 | No |
| 51 | Lta | na | 12876 | 0.065 | 0.5799 | No |
| 52 | Rela | na | 13587 | 0.049 | 0.5697 | No |
| 53 | Bid | na | 15056 | 0.025 | 0.5448 | No |
| 54 | Fasl | na | 15579 | 0.018 | 0.5365 | No |
| 55 | Map3k1 | na | 15852 | 0.016 | 0.5324 | No |
| 56 | Irf3 | na | 16894 | 0.008 | 0.5143 | No |
| 57 | Casp9 | na | 17273 | 0.006 | 0.5079 | No |
| 58 | Trp73 | na | 18233 | 0.002 | 0.4909 | No |
| 59 | Trp63 | na | 46972 | -0.010 | -0.0190 | No |
| 60 | Birc3 | na | 47264 | -0.018 | -0.0233 | No |
| 61 | Traf2 | na | 47959 | -0.036 | -0.0338 | No |
| 62 | Cradd | na | 49318 | -0.080 | -0.0539 | No |
| 63 | Irf1 | na | 49487 | -0.086 | -0.0527 | No |
| 64 | Pik3r1 | na | 49539 | -0.088 | -0.0492 | No |
| 65 | Casp8 | na | 49952 | -0.103 | -0.0514 | No |
| 66 | Tnf | na | 50208 | -0.112 | -0.0504 | No |
| 67 | Akt1 | na | 50442 | -0.121 | -0.0485 | No |
| 68 | Igf1 | na | 50907 | -0.141 | -0.0498 | No |
| 69 | Gzmc | na | 51306 | -0.157 | -0.0491 | No |
| 70 | Tnfrsf1a | na | 51804 | -0.177 | -0.0492 | No |
| 71 | Fas | na | 52026 | -0.187 | -0.0438 | No |
| 72 | Nfkbib | na | 52654 | -0.216 | -0.0443 | No |
| 73 | Mapk10 | na | 53661 | -0.267 | -0.0489 | No |
| 74 | Tradd | na | 54336 | -0.307 | -0.0456 | No |
| 75 | Bad | na | 54437 | -0.313 | -0.0319 | No |
| 76 | Igf2 | na | 55581 | -0.429 | -0.0309 | No |
| 77 | Traf1 | na | 55622 | -0.436 | -0.0100 | No |
| 78 | Myc | na | 55814 | -0.472 | 0.0100 | No |