Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___Cyp2c44KO_vs_WT.RPKM_matrix___Cyp2c44KO_vs_WT.cls #Cyp2c44KO_versus_WT |
| Phenotype | RPKM_matrix___Cyp2c44KO_vs_WT.cls#Cyp2c44KO_versus_WT |
| Upregulated in class | WT |
| GeneSet | WP_METAPATHWAY_BIOTRANSFORMATION |
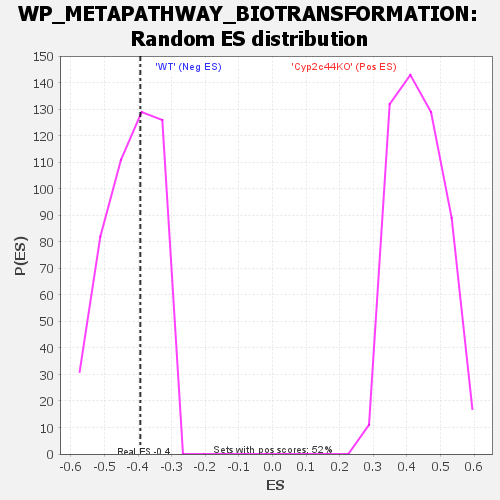
| Enrichment Score (ES) | -0.39343023 |
| Normalized Enrichment Score (NES) | -0.93407845 |
| Nominal p-value | 0.58246344 |
| FDR q-value | 0.9263713 |
| FWER p-Value | 0.994 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Ugt1a6a | na | 217 | 0.968 | 0.0340 | No |
| 2 | Chst3 | na | 304 | 0.910 | 0.0681 | No |
| 3 | Chst10 | na | 2052 | 0.542 | 0.0583 | No |
| 4 | Cyp4b1 | na | 2080 | 0.539 | 0.0789 | No |
| 5 | Nat14 | na | 2855 | 0.473 | 0.0837 | No |
| 6 | Naa80 | na | 3467 | 0.429 | 0.0896 | No |
| 7 | Naa40 | na | 3471 | 0.429 | 0.1064 | No |
| 8 | Cyp27b1 | na | 3496 | 0.428 | 0.1227 | No |
| 9 | Naa30 | na | 3886 | 0.403 | 0.1315 | No |
| 10 | Chst4 | na | 4065 | 0.391 | 0.1437 | No |
| 11 | Gsta1 | na | 4337 | 0.376 | 0.1536 | No |
| 12 | Akr1b10 | na | 4675 | 0.358 | 0.1616 | No |
| 13 | Nat10 | na | 4676 | 0.357 | 0.1756 | No |
| 14 | Ndst3 | na | 5646 | 0.310 | 0.1705 | No |
| 15 | Hs3st3a1 | na | 6109 | 0.290 | 0.1736 | No |
| 16 | Cyp4v3 | na | 6272 | 0.283 | 0.1818 | No |
| 17 | Sult1b1 | na | 7165 | 0.248 | 0.1757 | No |
| 18 | Cyp8b1 | na | 7175 | 0.247 | 0.1852 | No |
| 19 | Ndst2 | na | 7541 | 0.234 | 0.1879 | No |
| 20 | Nnmt | na | 7811 | 0.224 | 0.1918 | No |
| 21 | Gsta2 | na | 8234 | 0.209 | 0.1925 | No |
| 22 | Gsta4 | na | 8648 | 0.194 | 0.1928 | No |
| 23 | Chst11 | na | 8823 | 0.188 | 0.1971 | No |
| 24 | Cyp7a1 | na | 8961 | 0.183 | 0.2018 | No |
| 25 | Cyp2e1 | na | 9584 | 0.162 | 0.1971 | No |
| 26 | Ndst1 | na | 9740 | 0.157 | 0.2005 | No |
| 27 | Cyp46a1 | na | 9811 | 0.155 | 0.2053 | No |
| 28 | Ugt1a1 | na | 10213 | 0.141 | 0.2037 | No |
| 29 | Gstcd | na | 10421 | 0.135 | 0.2053 | No |
| 30 | Sult1c2 | na | 10520 | 0.132 | 0.2087 | No |
| 31 | Sult1e1 | na | 10831 | 0.123 | 0.2080 | No |
| 32 | Hs6st1 | na | 10896 | 0.120 | 0.2116 | No |
| 33 | Hs3st3b1 | na | 10925 | 0.120 | 0.2157 | No |
| 34 | Chst8 | na | 10968 | 0.118 | 0.2196 | No |
| 35 | Naa50 | na | 11270 | 0.110 | 0.2186 | No |
| 36 | Chst14 | na | 11278 | 0.110 | 0.2227 | No |
| 37 | Kcnab2 | na | 11474 | 0.103 | 0.2233 | No |
| 38 | Fmo2 | na | 11856 | 0.093 | 0.2202 | No |
| 39 | Cyp11a1 | na | 11952 | 0.090 | 0.2220 | No |
| 40 | Cyp21a1 | na | 12270 | 0.082 | 0.2196 | No |
| 41 | Ugt2a3 | na | 12569 | 0.073 | 0.2172 | No |
| 42 | Chst7 | na | 12761 | 0.068 | 0.2165 | No |
| 43 | Sult1a1 | na | 12815 | 0.067 | 0.2181 | No |
| 44 | Gstm7 | na | 13161 | 0.058 | 0.2143 | No |
| 45 | Gsto2 | na | 13345 | 0.054 | 0.2131 | No |
| 46 | Cyp4f18 | na | 14000 | 0.041 | 0.2031 | No |
| 47 | Hnmt | na | 14181 | 0.037 | 0.2014 | No |
| 48 | Gstt2 | na | 14228 | 0.037 | 0.2020 | No |
| 49 | Gstz1 | na | 14923 | 0.026 | 0.1907 | No |
| 50 | Nat8 | na | 15237 | 0.022 | 0.1860 | No |
| 51 | Hs6st3 | na | 16108 | 0.013 | 0.1710 | No |
| 52 | Chst1 | na | 16492 | 0.010 | 0.1646 | No |
| 53 | Fmo4 | na | 16584 | 0.010 | 0.1634 | No |
| 54 | Hs2st1 | na | 16872 | 0.008 | 0.1586 | No |
| 55 | Cyp2u1 | na | 16942 | 0.008 | 0.1577 | No |
| 56 | Gal3st3 | na | 17734 | 0.004 | 0.1437 | No |
| 57 | Cyp4x1 | na | 18155 | 0.002 | 0.1364 | No |
| 58 | Cyp11b1 | na | 18228 | 0.002 | 0.1352 | No |
| 59 | Hs3st2 | na | 18607 | 0.001 | 0.1285 | No |
| 60 | Hs3st4 | na | 20686 | 0.000 | 0.0915 | No |
| 61 | Hs3st5 | na | 22315 | 0.000 | 0.0626 | No |
| 62 | Cyp2w1 | na | 25088 | 0.000 | 0.0133 | No |
| 63 | Gal3st4 | na | 25654 | 0.000 | 0.0032 | No |
| 64 | Ndst4 | na | 32357 | 0.000 | -0.1159 | No |
| 65 | Cyp19a1 | na | 35337 | 0.000 | -0.1689 | No |
| 66 | Chst5 | na | 36796 | 0.000 | -0.1948 | No |
| 67 | Sult6b1 | na | 38283 | 0.000 | -0.2213 | No |
| 68 | Ugt2a2 | na | 45643 | 0.000 | -0.3521 | No |
| 69 | Ugt1a2 | na | 46477 | -0.000 | -0.3669 | No |
| 70 | Cyp24a1 | na | 46655 | -0.004 | -0.3699 | No |
| 71 | Cyp11b2 | na | 46861 | -0.008 | -0.3732 | No |
| 72 | Cyp27a1 | na | 47036 | -0.012 | -0.3759 | No |
| 73 | Gss | na | 47424 | -0.022 | -0.3819 | No |
| 74 | Gstp1 | na | 47515 | -0.024 | -0.3825 | No |
| 75 | Hs6st2 | na | 47602 | -0.026 | -0.3830 | No |
| 76 | Nat9 | na | 47736 | -0.029 | -0.3843 | No |
| 77 | Kcnab3 | na | 47802 | -0.031 | -0.3842 | No |
| 78 | Gstm5 | na | 48269 | -0.045 | -0.3907 | No |
| 79 | Mgst3 | na | 48423 | -0.050 | -0.3915 | Yes |
| 80 | Cyp20a1 | na | 48433 | -0.050 | -0.3897 | Yes |
| 81 | Cyp39a1 | na | 48443 | -0.050 | -0.3879 | Yes |
| 82 | Kcnab1 | na | 48597 | -0.055 | -0.3885 | Yes |
| 83 | Fmo1 | na | 48697 | -0.058 | -0.3880 | Yes |
| 84 | Gsta3 | na | 48738 | -0.060 | -0.3863 | Yes |
| 85 | Ugt1a9 | na | 48837 | -0.063 | -0.3856 | Yes |
| 86 | Akr1d1 | na | 49032 | -0.070 | -0.3863 | Yes |
| 87 | Chst12 | na | 49091 | -0.072 | -0.3845 | Yes |
| 88 | Gstk1 | na | 49297 | -0.079 | -0.3850 | Yes |
| 89 | Cyp26b1 | na | 49381 | -0.082 | -0.3833 | Yes |
| 90 | Naa20 | na | 49384 | -0.083 | -0.3801 | Yes |
| 91 | Cyp7b1 | na | 49468 | -0.085 | -0.3782 | Yes |
| 92 | Cyp2s1 | na | 49544 | -0.088 | -0.3761 | Yes |
| 93 | Gstm4 | na | 49661 | -0.093 | -0.3745 | Yes |
| 94 | Gpx5 | na | 49692 | -0.094 | -0.3714 | Yes |
| 95 | Fmo3 | na | 49820 | -0.098 | -0.3698 | Yes |
| 96 | Gstm1 | na | 49912 | -0.101 | -0.3674 | Yes |
| 97 | Ugt1a5 | na | 49913 | -0.101 | -0.3635 | Yes |
| 98 | Mgst1 | na | 50002 | -0.105 | -0.3609 | Yes |
| 99 | Fmo5 | na | 50030 | -0.106 | -0.3573 | Yes |
| 100 | Cyp26c1 | na | 50286 | -0.115 | -0.3573 | Yes |
| 101 | Gsr | na | 50527 | -0.125 | -0.3567 | Yes |
| 102 | Hs3st1 | na | 50622 | -0.128 | -0.3533 | Yes |
| 103 | Akr1a1 | na | 50651 | -0.129 | -0.3488 | Yes |
| 104 | Nat2 | na | 51198 | -0.152 | -0.3525 | Yes |
| 105 | Glyat | na | 51223 | -0.153 | -0.3470 | Yes |
| 106 | Inmt | na | 51410 | -0.161 | -0.3440 | Yes |
| 107 | Gstt1 | na | 51416 | -0.162 | -0.3377 | Yes |
| 108 | Gsto1 | na | 51456 | -0.164 | -0.3320 | Yes |
| 109 | Baat | na | 51472 | -0.164 | -0.3259 | Yes |
| 110 | Akr7a5 | na | 51636 | -0.171 | -0.3221 | Yes |
| 111 | Gal3st1 | na | 51856 | -0.179 | -0.3190 | Yes |
| 112 | Cyp4f39 | na | 52475 | -0.207 | -0.3219 | Yes |
| 113 | Chst13 | na | 52524 | -0.209 | -0.3145 | Yes |
| 114 | Cyp1a2 | na | 52577 | -0.212 | -0.3071 | Yes |
| 115 | Cyp1b1 | na | 52778 | -0.221 | -0.3020 | Yes |
| 116 | Cyp1a1 | na | 52912 | -0.227 | -0.2955 | Yes |
| 117 | Ephx2 | na | 52914 | -0.227 | -0.2866 | Yes |
| 118 | Chst2 | na | 53062 | -0.236 | -0.2800 | Yes |
| 119 | Hs3st6 | na | 53101 | -0.238 | -0.2713 | Yes |
| 120 | Cyp2r1 | na | 53186 | -0.243 | -0.2633 | Yes |
| 121 | Nat8l | na | 53195 | -0.244 | -0.2539 | Yes |
| 122 | Gpx4 | na | 53403 | -0.254 | -0.2476 | Yes |
| 123 | Cyp2f2 | na | 53563 | -0.260 | -0.2403 | Yes |
| 124 | Chst9 | na | 53595 | -0.262 | -0.2305 | Yes |
| 125 | Gpx1 | na | 53648 | -0.266 | -0.2211 | Yes |
| 126 | Cyp17a1 | na | 53714 | -0.270 | -0.2117 | Yes |
| 127 | Mgst2 | na | 53832 | -0.277 | -0.2029 | Yes |
| 128 | Sult2b1 | na | 53906 | -0.282 | -0.1932 | Yes |
| 129 | Sult4a1 | na | 54136 | -0.296 | -0.1856 | Yes |
| 130 | Comt | na | 54431 | -0.313 | -0.1786 | Yes |
| 131 | Ephx1 | na | 54478 | -0.316 | -0.1671 | Yes |
| 132 | Gpx2 | na | 54524 | -0.320 | -0.1553 | Yes |
| 133 | Tpmt | na | 54925 | -0.350 | -0.1488 | Yes |
| 134 | Ugt1a10 | na | 55022 | -0.359 | -0.1364 | Yes |
| 135 | Gal3st2 | na | 55087 | -0.366 | -0.1232 | Yes |
| 136 | Sult2a1 | na | 55384 | -0.398 | -0.1129 | Yes |
| 137 | Akr1b3 | na | 56054 | -0.539 | -0.1037 | Yes |
| 138 | Gpx3 | na | 56305 | -0.730 | -0.0796 | Yes |
| 139 | Cyp51 | na | 56359 | -0.971 | -0.0425 | Yes |
| 140 | Cyp26a1 | na | 56371 | -1.094 | 0.0001 | Yes |