Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___PanKO_vs_ClusterKO.RPKM_matrix___PanKO_vs_ClusterKO.cls #PanKO_versus_ClusterKO |
| Phenotype | RPKM_matrix___PanKO_vs_ClusterKO.cls#PanKO_versus_ClusterKO |
| Upregulated in class | ClusterKO |
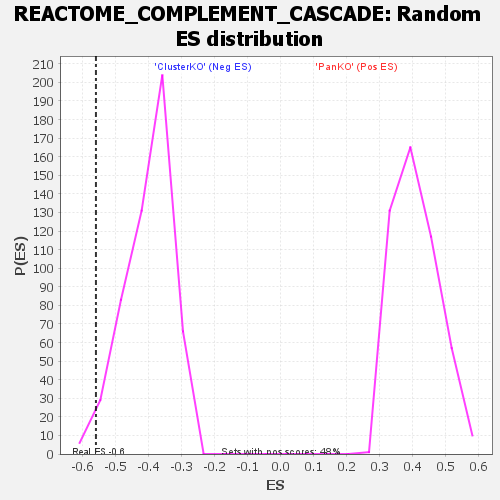
| GeneSet | REACTOME_COMPLEMENT_CASCADE |
| Enrichment Score (ES) | -0.5606923 |
| Normalized Enrichment Score (NES) | -1.3965703 |
| Nominal p-value | 0.028901733 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.763 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Ighv1-78 | na | 1237 | 0.439 | -0.0109 | No |
| 2 | Clu | na | 1709 | 0.390 | -0.0095 | No |
| 3 | C1s2 | na | 1838 | 0.379 | -0.0022 | No |
| 4 | C5ar1 | na | 1853 | 0.378 | 0.0072 | No |
| 5 | Ighv3-8 | na | 2054 | 0.362 | 0.0127 | No |
| 6 | Cfi | na | 2606 | 0.323 | 0.0111 | No |
| 7 | Ighv5-6 | na | 2732 | 0.316 | 0.0168 | No |
| 8 | Igkv14-100 | na | 2891 | 0.305 | 0.0217 | No |
| 9 | Igkv1-133 | na | 2921 | 0.303 | 0.0289 | No |
| 10 | Cfb | na | 2955 | 0.301 | 0.0359 | No |
| 11 | C8a | na | 3090 | 0.293 | 0.0409 | No |
| 12 | Ighv1-47 | na | 3355 | 0.279 | 0.0433 | No |
| 13 | Ighv1-81 | na | 3671 | 0.263 | 0.0443 | No |
| 14 | Cpn1 | na | 3778 | 0.258 | 0.0489 | No |
| 15 | Ighv4-1 | na | 4388 | 0.228 | 0.0438 | No |
| 16 | Ighv1-34 | na | 4498 | 0.223 | 0.0475 | No |
| 17 | Ighv1-49 | na | 4611 | 0.218 | 0.0510 | No |
| 18 | Igkv14-126 | na | 4847 | 0.208 | 0.0521 | No |
| 19 | Cpb2 | na | 4939 | 0.205 | 0.0557 | No |
| 20 | C8b | na | 5205 | 0.193 | 0.0558 | No |
| 21 | Vtn | na | 5349 | 0.189 | 0.0580 | No |
| 22 | Fcnb | na | 5881 | 0.169 | 0.0529 | No |
| 23 | Ighv1-4 | na | 6116 | 0.160 | 0.0527 | No |
| 24 | Mbl2 | na | 6294 | 0.154 | 0.0535 | No |
| 25 | Ighv1-12 | na | 6489 | 0.147 | 0.0537 | No |
| 26 | Ighv1-75 | na | 6656 | 0.140 | 0.0543 | No |
| 27 | Ighv5-9-1 | na | 6807 | 0.135 | 0.0551 | No |
| 28 | Ighv1-77 | na | 6818 | 0.134 | 0.0583 | No |
| 29 | Ighv8-12 | na | 7212 | 0.122 | 0.0544 | No |
| 30 | Ighv1-82 | na | 7337 | 0.118 | 0.0551 | No |
| 31 | Ighv1-66 | na | 7887 | 0.099 | 0.0479 | No |
| 32 | Elane | na | 7987 | 0.095 | 0.0485 | No |
| 33 | C6 | na | 8126 | 0.090 | 0.0483 | No |
| 34 | C9 | na | 8282 | 0.085 | 0.0477 | No |
| 35 | Ighv1-20 | na | 8436 | 0.080 | 0.0470 | No |
| 36 | C7 | na | 8585 | 0.078 | 0.0463 | No |
| 37 | Serping1 | na | 8903 | 0.068 | 0.0424 | No |
| 38 | Ighv8-6 | na | 9099 | 0.061 | 0.0405 | No |
| 39 | Cd81 | na | 9134 | 0.060 | 0.0414 | No |
| 40 | Gzmm | na | 9342 | 0.054 | 0.0391 | No |
| 41 | Ighv3-4 | na | 9564 | 0.048 | 0.0364 | No |
| 42 | C1ra | na | 9599 | 0.048 | 0.0370 | No |
| 43 | Ighv1-84 | na | 9827 | 0.041 | 0.0340 | No |
| 44 | Ighv1-31 | na | 9830 | 0.041 | 0.0349 | No |
| 45 | C3ar1 | na | 10225 | 0.030 | 0.0287 | No |
| 46 | Ighv1-37 | na | 10431 | 0.025 | 0.0257 | No |
| 47 | Ighv3-6 | na | 10555 | 0.022 | 0.0241 | No |
| 48 | Igkv13-84 | na | 10733 | 0.018 | 0.0214 | No |
| 49 | F2 | na | 10855 | 0.016 | 0.0196 | No |
| 50 | Ighv12-3 | na | 11086 | 0.012 | 0.0158 | No |
| 51 | C3 | na | 11227 | 0.010 | 0.0136 | No |
| 52 | Ighv11-1 | na | 12444 | 0.000 | -0.0081 | No |
| 53 | Ighv8-2 | na | 13356 | 0.000 | -0.0243 | No |
| 54 | Igkv2-137 | na | 13595 | 0.000 | -0.0285 | No |
| 55 | Ighv5-12-4 | na | 14234 | 0.000 | -0.0398 | No |
| 56 | Ighv1-71 | na | 14940 | 0.000 | -0.0524 | No |
| 57 | Ighv1-69 | na | 15259 | 0.000 | -0.0580 | No |
| 58 | Ighv1-24 | na | 15495 | 0.000 | -0.0622 | No |
| 59 | Igkv1-35 | na | 16127 | 0.000 | -0.0734 | No |
| 60 | Igkv20-101-2 | na | 19795 | 0.000 | -0.1387 | No |
| 61 | Igkv1-132 | na | 22200 | 0.000 | -0.1814 | No |
| 62 | Ighv8-9 | na | 23076 | 0.000 | -0.1970 | No |
| 63 | Ighv5-2 | na | 25533 | 0.000 | -0.2407 | No |
| 64 | Ighv1-23 | na | 26362 | 0.000 | -0.2554 | No |
| 65 | Ighv3-3 | na | 26603 | 0.000 | -0.2597 | No |
| 66 | Ighv8-4 | na | 29013 | 0.000 | -0.3026 | No |
| 67 | Ighv1-16 | na | 30525 | 0.000 | -0.3294 | No |
| 68 | Igkv1-131 | na | 30783 | 0.000 | -0.3340 | No |
| 69 | Ighv8-11 | na | 30840 | 0.000 | -0.3350 | No |
| 70 | Ighv1-62-3 | na | 33034 | 0.000 | -0.3740 | No |
| 71 | Ighv1-43 | na | 35620 | 0.000 | -0.4200 | No |
| 72 | Ighv1-62-2 | na | 35630 | 0.000 | -0.4202 | No |
| 73 | Ighv15-2 | na | 36431 | 0.000 | -0.4344 | No |
| 74 | Igkv18-36 | na | 37299 | 0.000 | -0.4498 | No |
| 75 | Ighv8-13 | na | 40280 | -0.009 | -0.5026 | No |
| 76 | Igll1 | na | 40390 | -0.010 | -0.5043 | No |
| 77 | Crp | na | 40412 | -0.010 | -0.5044 | No |
| 78 | Ighv1-63 | na | 41544 | -0.024 | -0.5239 | No |
| 79 | Ighv1-22 | na | 42049 | -0.033 | -0.5320 | No |
| 80 | Igkv2-112 | na | 42099 | -0.034 | -0.5320 | No |
| 81 | Ighv1-19 | na | 42200 | -0.036 | -0.5329 | No |
| 82 | Ighv1-50 | na | 42626 | -0.045 | -0.5393 | No |
| 83 | Ighv14-4 | na | 42645 | -0.046 | -0.5385 | No |
| 84 | Ighv1-67 | na | 42863 | -0.051 | -0.5411 | No |
| 85 | Hc | na | 42954 | -0.053 | -0.5413 | No |
| 86 | Masp2 | na | 43019 | -0.054 | -0.5411 | No |
| 87 | Ighv5-15 | na | 43083 | -0.056 | -0.5408 | No |
| 88 | Ighv11-2 | na | 43366 | -0.063 | -0.5442 | No |
| 89 | Igkv13-85 | na | 43384 | -0.063 | -0.5429 | No |
| 90 | Igkv1-99 | na | 43414 | -0.064 | -0.5418 | No |
| 91 | Ighv1-36 | na | 43437 | -0.065 | -0.5406 | No |
| 92 | Igkv14-130 | na | 43438 | -0.065 | -0.5390 | No |
| 93 | Ighv1-80 | na | 43919 | -0.078 | -0.5455 | No |
| 94 | Ighv1-72 | na | 44135 | -0.084 | -0.5472 | No |
| 95 | Ighv1-61 | na | 44215 | -0.086 | -0.5465 | No |
| 96 | Ighv1-15 | na | 44602 | -0.099 | -0.5508 | No |
| 97 | Ighv1-52 | na | 44646 | -0.100 | -0.5491 | No |
| 98 | Ighv5-12 | na | 44824 | -0.106 | -0.5496 | No |
| 99 | Igkv12-98 | na | 44982 | -0.110 | -0.5496 | No |
| 100 | Ighv1-58 | na | 45156 | -0.115 | -0.5498 | No |
| 101 | Igkv1-122 | na | 45581 | -0.127 | -0.5541 | No |
| 102 | Cr1l | na | 45953 | -0.139 | -0.5572 | Yes |
| 103 | Masp1 | na | 46151 | -0.145 | -0.5570 | Yes |
| 104 | C1qb | na | 46226 | -0.148 | -0.5546 | Yes |
| 105 | Ighv8-5 | na | 46514 | -0.158 | -0.5557 | Yes |
| 106 | Ighv1-42 | na | 46671 | -0.163 | -0.5544 | Yes |
| 107 | C4b | na | 46911 | -0.170 | -0.5543 | Yes |
| 108 | Ighv1-5 | na | 47105 | -0.176 | -0.5533 | Yes |
| 109 | C1qa | na | 47149 | -0.177 | -0.5496 | Yes |
| 110 | Ighv14-1 | na | 47718 | -0.197 | -0.5547 | Yes |
| 111 | Ighv5-16 | na | 47792 | -0.200 | -0.5510 | Yes |
| 112 | Ighv3-1 | na | 48124 | -0.211 | -0.5515 | Yes |
| 113 | Ighv1-76 | na | 48309 | -0.218 | -0.5493 | Yes |
| 114 | Ighv1-64 | na | 48434 | -0.222 | -0.5459 | Yes |
| 115 | Igkv17-121 | na | 48602 | -0.227 | -0.5432 | Yes |
| 116 | Igkv1-110 | na | 48759 | -0.234 | -0.5400 | Yes |
| 117 | Igkv1-88 | na | 48780 | -0.234 | -0.5345 | Yes |
| 118 | C8g | na | 49246 | -0.253 | -0.5364 | Yes |
| 119 | C1qc | na | 49525 | -0.263 | -0.5347 | Yes |
| 120 | Ighg1 | na | 49529 | -0.263 | -0.5281 | Yes |
| 121 | Ighv1-59 | na | 49845 | -0.277 | -0.5267 | Yes |
| 122 | Cfhr4 | na | 49850 | -0.277 | -0.5198 | Yes |
| 123 | Ighv1-85 | na | 50119 | -0.288 | -0.5173 | Yes |
| 124 | Ighv1-9 | na | 50597 | -0.307 | -0.5180 | Yes |
| 125 | Ighv5-9 | na | 50739 | -0.313 | -0.5126 | Yes |
| 126 | Cpn2 | na | 50842 | -0.318 | -0.5064 | Yes |
| 127 | Ighv14-3 | na | 50911 | -0.321 | -0.4995 | Yes |
| 128 | Ighv1-56 | na | 51342 | -0.341 | -0.4985 | Yes |
| 129 | Ighv1-11 | na | 52141 | -0.378 | -0.5032 | Yes |
| 130 | Ighv1-26 | na | 52434 | -0.394 | -0.4984 | Yes |
| 131 | Ighv1-53 | na | 52683 | -0.410 | -0.4925 | Yes |
| 132 | Cfp | na | 53106 | -0.436 | -0.4890 | Yes |
| 133 | Igkv8-21 | na | 53261 | -0.445 | -0.4805 | Yes |
| 134 | Cfd | na | 53313 | -0.448 | -0.4701 | Yes |
| 135 | C2 | na | 53500 | -0.461 | -0.4618 | Yes |
| 136 | Ighg3 | na | 53659 | -0.471 | -0.4527 | Yes |
| 137 | Igkv14-111 | na | 53864 | -0.486 | -0.4441 | Yes |
| 138 | Ighv1-74 | na | 54031 | -0.498 | -0.4345 | Yes |
| 139 | Cd59b | na | 54122 | -0.506 | -0.4233 | Yes |
| 140 | Cfhr1 | na | 54179 | -0.511 | -0.4114 | Yes |
| 141 | Igkv1-117 | na | 54198 | -0.512 | -0.3988 | Yes |
| 142 | Ighv1-39 | na | 54524 | -0.542 | -0.3909 | Yes |
| 143 | Igkv1-135 | na | 54590 | -0.549 | -0.3782 | Yes |
| 144 | C5ar2 | na | 54623 | -0.553 | -0.3648 | Yes |
| 145 | Igkv15-103 | na | 54677 | -0.557 | -0.3517 | Yes |
| 146 | Fcna | na | 54695 | -0.560 | -0.3378 | Yes |
| 147 | Igkv17-127 | na | 54746 | -0.566 | -0.3245 | Yes |
| 148 | Cd55 | na | 54801 | -0.572 | -0.3110 | Yes |
| 149 | Ighg2c | na | 55013 | -0.598 | -0.2996 | Yes |
| 150 | Ighv1-55 | na | 55149 | -0.616 | -0.2865 | Yes |
| 151 | Ighv14-2 | na | 55219 | -0.625 | -0.2719 | Yes |
| 152 | Ighv1-7 | na | 55505 | -0.667 | -0.2602 | Yes |
| 153 | Ighv8-8 | na | 55616 | -0.689 | -0.2447 | Yes |
| 154 | Ighv1-54 | na | 55706 | -0.709 | -0.2284 | Yes |
| 155 | Iglc2 | na | 55832 | -0.744 | -0.2118 | Yes |
| 156 | Cr2 | na | 55949 | -0.775 | -0.1943 | Yes |
| 157 | Ighv5-4 | na | 56089 | -0.829 | -0.1759 | Yes |
| 158 | Ighv1-18 | na | 56150 | -0.862 | -0.1552 | Yes |
| 159 | Colec11 | na | 56191 | -0.893 | -0.1334 | Yes |
| 160 | Cd46 | na | 56241 | -0.953 | -0.1102 | Yes |
| 161 | Iglc1 | na | 56251 | -0.970 | -0.0859 | Yes |
| 162 | Cd19 | na | 56327 | -1.112 | -0.0591 | Yes |
| 163 | Colec10 | na | 56342 | -1.167 | -0.0299 | Yes |
| 164 | Ighv5-17 | na | 56355 | -1.208 | 0.0004 | Yes |