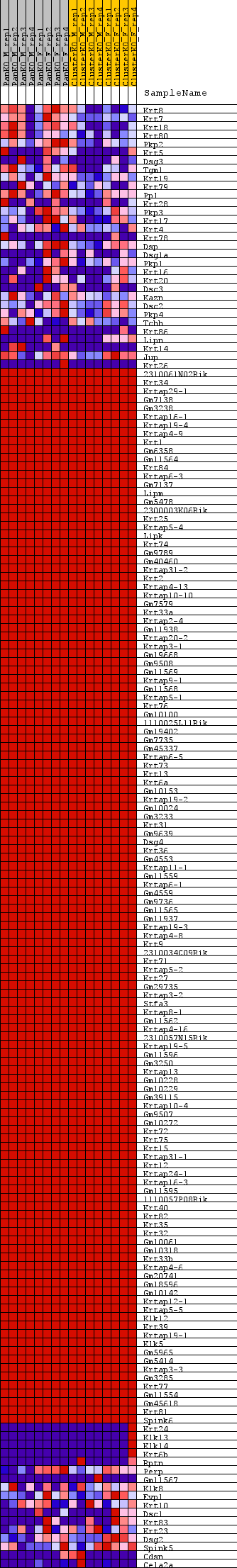
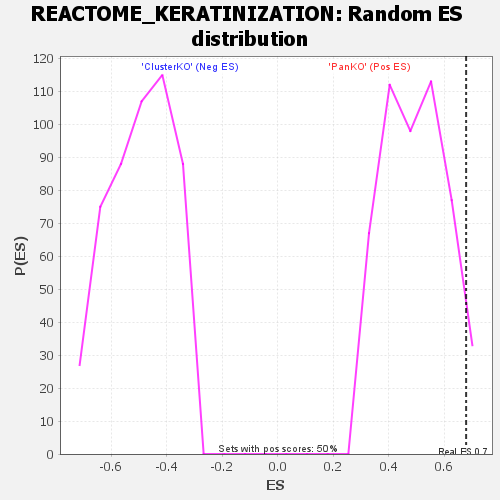
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___PanKO_vs_ClusterKO.RPKM_matrix___PanKO_vs_ClusterKO.cls #PanKO_versus_ClusterKO |
| Phenotype | RPKM_matrix___PanKO_vs_ClusterKO.cls#PanKO_versus_ClusterKO |
| Upregulated in class | PanKO |
| GeneSet | REACTOME_KERATINIZATION |
| Enrichment Score (ES) | 0.67979217 |
| Normalized Enrichment Score (NES) | 1.3674546 |
| Nominal p-value | 0.046 |
| FDR q-value | 0.7200399 |
| FWER p-Value | 0.834 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Krt8 | na | 91 | 0.747 | 0.0688 | Yes |
| 2 | Krt7 | na | 159 | 0.686 | 0.1323 | Yes |
| 3 | Krt18 | na | 233 | 0.647 | 0.1920 | Yes |
| 4 | Krt80 | na | 409 | 0.584 | 0.2439 | Yes |
| 5 | Pkp2 | na | 864 | 0.489 | 0.2820 | Yes |
| 6 | Krt5 | na | 1091 | 0.454 | 0.3208 | Yes |
| 7 | Dsg3 | na | 1340 | 0.427 | 0.3567 | Yes |
| 8 | Tgm1 | na | 1649 | 0.396 | 0.3885 | Yes |
| 9 | Krt19 | na | 2139 | 0.355 | 0.4133 | Yes |
| 10 | Krt79 | na | 2552 | 0.327 | 0.4368 | Yes |
| 11 | Ppl | na | 2802 | 0.310 | 0.4616 | Yes |
| 12 | Krt28 | na | 3025 | 0.297 | 0.4857 | Yes |
| 13 | Pkp3 | na | 3332 | 0.280 | 0.5067 | Yes |
| 14 | Krt17 | na | 3894 | 0.252 | 0.5204 | Yes |
| 15 | Krt4 | na | 4450 | 0.224 | 0.5317 | Yes |
| 16 | Krt78 | na | 5019 | 0.201 | 0.5405 | Yes |
| 17 | Dsp | na | 5022 | 0.200 | 0.5593 | Yes |
| 18 | Dsg1a | na | 5060 | 0.199 | 0.5775 | Yes |
| 19 | Pkp1 | na | 5318 | 0.189 | 0.5907 | Yes |
| 20 | Krt16 | na | 5443 | 0.185 | 0.6060 | Yes |
| 21 | Krt20 | na | 5565 | 0.181 | 0.6209 | Yes |
| 22 | Dsc3 | na | 5779 | 0.173 | 0.6334 | Yes |
| 23 | Kazn | na | 5916 | 0.168 | 0.6468 | Yes |
| 24 | Dsc2 | na | 6745 | 0.137 | 0.6450 | Yes |
| 25 | Pkp4 | na | 6825 | 0.134 | 0.6563 | Yes |
| 26 | Tchh | na | 6868 | 0.133 | 0.6681 | Yes |
| 27 | Krt86 | na | 6908 | 0.132 | 0.6798 | Yes |
| 28 | Lipn | na | 7610 | 0.109 | 0.6776 | No |
| 29 | Krt14 | na | 9041 | 0.063 | 0.6581 | No |
| 30 | Jup | na | 10526 | 0.023 | 0.6338 | No |
| 31 | Krt26 | na | 11936 | 0.002 | 0.6089 | No |
| 32 | 2310061N02Rik | na | 12142 | 0.000 | 0.6053 | No |
| 33 | Krt34 | na | 12488 | 0.000 | 0.5992 | No |
| 34 | Krtap29-1 | na | 12923 | 0.000 | 0.5914 | No |
| 35 | Gm7138 | na | 12972 | 0.000 | 0.5906 | No |
| 36 | Gm3238 | na | 13396 | 0.000 | 0.5831 | No |
| 37 | Krtap16-1 | na | 13423 | 0.000 | 0.5826 | No |
| 38 | Krtap19-4 | na | 13797 | 0.000 | 0.5760 | No |
| 39 | Krtap4-9 | na | 14195 | 0.000 | 0.5689 | No |
| 40 | Krt1 | na | 14283 | 0.000 | 0.5673 | No |
| 41 | Gm6358 | na | 14560 | 0.000 | 0.5624 | No |
| 42 | Gm11564 | na | 14584 | 0.000 | 0.5620 | No |
| 43 | Krt84 | na | 14906 | 0.000 | 0.5563 | No |
| 44 | Krtap6-3 | na | 15218 | 0.000 | 0.5508 | No |
| 45 | Gm7137 | na | 15421 | 0.000 | 0.5472 | No |
| 46 | Lipm | na | 15521 | 0.000 | 0.5454 | No |
| 47 | Gm5478 | na | 15566 | 0.000 | 0.5446 | No |
| 48 | 2300003K06Rik | na | 16015 | 0.000 | 0.5367 | No |
| 49 | Krt25 | na | 16083 | 0.000 | 0.5355 | No |
| 50 | Krtap5-4 | na | 16242 | 0.000 | 0.5327 | No |
| 51 | Lipk | na | 16512 | 0.000 | 0.5279 | No |
| 52 | Krt74 | na | 16521 | 0.000 | 0.5277 | No |
| 53 | Gm9789 | na | 16600 | 0.000 | 0.5264 | No |
| 54 | Gm40460 | na | 17051 | 0.000 | 0.5183 | No |
| 55 | Krtap31-2 | na | 17297 | 0.000 | 0.5140 | No |
| 56 | Krt2 | na | 17371 | 0.000 | 0.5127 | No |
| 57 | Krtap4-13 | na | 17744 | 0.000 | 0.5061 | No |
| 58 | Krtap10-10 | na | 17839 | 0.000 | 0.5044 | No |
| 59 | Gm7579 | na | 17901 | 0.000 | 0.5033 | No |
| 60 | Krt33a | na | 17909 | 0.000 | 0.5032 | No |
| 61 | Krtap2-4 | na | 18372 | 0.000 | 0.4950 | No |
| 62 | Gm11938 | na | 18538 | 0.000 | 0.4920 | No |
| 63 | Krtap20-2 | na | 19349 | 0.000 | 0.4776 | No |
| 64 | Krtap3-1 | na | 19500 | 0.000 | 0.4750 | No |
| 65 | Gm19668 | na | 19563 | 0.000 | 0.4738 | No |
| 66 | Gm9508 | na | 19767 | 0.000 | 0.4702 | No |
| 67 | Gm11569 | na | 19800 | 0.000 | 0.4697 | No |
| 68 | Krtap9-1 | na | 19818 | 0.000 | 0.4694 | No |
| 69 | Gm11568 | na | 19995 | 0.000 | 0.4662 | No |
| 70 | Krtap5-1 | na | 20174 | 0.000 | 0.4631 | No |
| 71 | Krt76 | na | 20510 | 0.000 | 0.4571 | No |
| 72 | Gm10100 | na | 20770 | 0.000 | 0.4525 | No |
| 73 | 1110025L11Rik | na | 20894 | 0.000 | 0.4503 | No |
| 74 | Gm19402 | na | 21549 | 0.000 | 0.4387 | No |
| 75 | Gm7735 | na | 21961 | 0.000 | 0.4314 | No |
| 76 | Gm45337 | na | 22081 | 0.000 | 0.4292 | No |
| 77 | Krtap6-5 | na | 22171 | 0.000 | 0.4277 | No |
| 78 | Krt73 | na | 22481 | 0.000 | 0.4222 | No |
| 79 | Krt13 | na | 22747 | 0.000 | 0.4174 | No |
| 80 | Krt6a | na | 22757 | 0.000 | 0.4173 | No |
| 81 | Gm10153 | na | 23108 | 0.000 | 0.4111 | No |
| 82 | Krtap19-2 | na | 23206 | 0.000 | 0.4093 | No |
| 83 | Gm10024 | na | 23207 | 0.000 | 0.4093 | No |
| 84 | Gm3233 | na | 23248 | 0.000 | 0.4086 | No |
| 85 | Krt31 | na | 23413 | 0.000 | 0.4057 | No |
| 86 | Gm9639 | na | 23847 | 0.000 | 0.3980 | No |
| 87 | Dsg4 | na | 23947 | 0.000 | 0.3962 | No |
| 88 | Krt36 | na | 24611 | 0.000 | 0.3844 | No |
| 89 | Gm4553 | na | 25198 | 0.000 | 0.3740 | No |
| 90 | Krtap11-1 | na | 25452 | 0.000 | 0.3695 | No |
| 91 | Gm11559 | na | 25539 | 0.000 | 0.3680 | No |
| 92 | Krtap6-1 | na | 25632 | 0.000 | 0.3664 | No |
| 93 | Gm4559 | na | 25691 | 0.000 | 0.3653 | No |
| 94 | Gm9736 | na | 25696 | 0.000 | 0.3652 | No |
| 95 | Gm11565 | na | 26457 | 0.000 | 0.3517 | No |
| 96 | Gm11937 | na | 26913 | 0.000 | 0.3436 | No |
| 97 | Krtap19-3 | na | 27052 | 0.000 | 0.3412 | No |
| 98 | Krtap4-8 | na | 27084 | 0.000 | 0.3406 | No |
| 99 | Krt9 | na | 28052 | 0.000 | 0.3234 | No |
| 100 | 2310034C09Rik | na | 28203 | 0.000 | 0.3208 | No |
| 101 | Krt71 | na | 28297 | 0.000 | 0.3191 | No |
| 102 | Krtap5-2 | na | 28475 | 0.000 | 0.3159 | No |
| 103 | Krt27 | na | 28582 | 0.000 | 0.3141 | No |
| 104 | Gm29735 | na | 28717 | 0.000 | 0.3117 | No |
| 105 | Krtap3-2 | na | 28936 | 0.000 | 0.3078 | No |
| 106 | Stfa3 | na | 29096 | 0.000 | 0.3050 | No |
| 107 | Krtap8-1 | na | 29377 | 0.000 | 0.3000 | No |
| 108 | Gm11562 | na | 29671 | 0.000 | 0.2948 | No |
| 109 | Krtap4-16 | na | 30037 | 0.000 | 0.2883 | No |
| 110 | 2310057N15Rik | na | 30228 | 0.000 | 0.2849 | No |
| 111 | Krtap19-5 | na | 31151 | 0.000 | 0.2685 | No |
| 112 | Gm11596 | na | 31945 | 0.000 | 0.2544 | No |
| 113 | Gm3250 | na | 31951 | 0.000 | 0.2543 | No |
| 114 | Krtap13 | na | 32158 | 0.000 | 0.2506 | No |
| 115 | Gm10228 | na | 32160 | 0.000 | 0.2506 | No |
| 116 | Gm10229 | na | 32278 | 0.000 | 0.2485 | No |
| 117 | Gm39115 | na | 32420 | 0.000 | 0.2460 | No |
| 118 | Krtap10-4 | na | 32555 | 0.000 | 0.2436 | No |
| 119 | Gm9507 | na | 32612 | 0.000 | 0.2426 | No |
| 120 | Gm10272 | na | 32664 | 0.000 | 0.2417 | No |
| 121 | Krt72 | na | 32820 | 0.000 | 0.2390 | No |
| 122 | Krt75 | na | 32898 | 0.000 | 0.2376 | No |
| 123 | Krt15 | na | 33018 | 0.000 | 0.2355 | No |
| 124 | Krtap31-1 | na | 33122 | 0.000 | 0.2337 | No |
| 125 | Krt12 | na | 33178 | 0.000 | 0.2327 | No |
| 126 | Krtap24-1 | na | 33332 | 0.000 | 0.2300 | No |
| 127 | Krtap16-3 | na | 33404 | 0.000 | 0.2287 | No |
| 128 | Gm11595 | na | 33434 | 0.000 | 0.2282 | No |
| 129 | 1110057P08Rik | na | 33580 | 0.000 | 0.2256 | No |
| 130 | Krt40 | na | 34032 | 0.000 | 0.2176 | No |
| 131 | Krt82 | na | 34805 | 0.000 | 0.2038 | No |
| 132 | Krt35 | na | 34831 | 0.000 | 0.2034 | No |
| 133 | Krt32 | na | 34913 | 0.000 | 0.2020 | No |
| 134 | Gm10061 | na | 35081 | 0.000 | 0.1990 | No |
| 135 | Gm10318 | na | 35346 | 0.000 | 0.1943 | No |
| 136 | Krt33b | na | 35392 | 0.000 | 0.1935 | No |
| 137 | Krtap4-6 | na | 35493 | 0.000 | 0.1917 | No |
| 138 | Gm20741 | na | 35653 | 0.000 | 0.1889 | No |
| 139 | Gm18596 | na | 35698 | 0.000 | 0.1881 | No |
| 140 | Gm10142 | na | 36065 | 0.000 | 0.1816 | No |
| 141 | Krtap12-1 | na | 36113 | 0.000 | 0.1807 | No |
| 142 | Krtap5-5 | na | 36210 | 0.000 | 0.1790 | No |
| 143 | Klk12 | na | 36432 | 0.000 | 0.1751 | No |
| 144 | Krt39 | na | 36644 | 0.000 | 0.1714 | No |
| 145 | Krtap19-1 | na | 36697 | 0.000 | 0.1704 | No |
| 146 | Klk5 | na | 37110 | 0.000 | 0.1631 | No |
| 147 | Gm5965 | na | 37190 | 0.000 | 0.1617 | No |
| 148 | Gm5414 | na | 37289 | 0.000 | 0.1600 | No |
| 149 | Krtap3-3 | na | 37317 | 0.000 | 0.1595 | No |
| 150 | Gm3285 | na | 38022 | 0.000 | 0.1469 | No |
| 151 | Krt77 | na | 38215 | 0.000 | 0.1435 | No |
| 152 | Gm11554 | na | 38571 | 0.000 | 0.1372 | No |
| 153 | Gm45618 | na | 38574 | 0.000 | 0.1372 | No |
| 154 | Krt81 | na | 38602 | 0.000 | 0.1367 | No |
| 155 | Spink6 | na | 38725 | 0.000 | 0.1345 | No |
| 156 | Krt24 | na | 38990 | -0.002 | 0.1300 | No |
| 157 | Klk13 | na | 38997 | -0.002 | 0.1300 | No |
| 158 | Klk14 | na | 39162 | -0.002 | 0.1273 | No |
| 159 | Krt6b | na | 39208 | -0.003 | 0.1268 | No |
| 160 | Rptn | na | 39650 | -0.005 | 0.1194 | No |
| 161 | Perp | na | 39918 | -0.006 | 0.1152 | No |
| 162 | Gm11567 | na | 41001 | -0.017 | 0.0975 | No |
| 163 | Klk8 | na | 42163 | -0.035 | 0.0802 | No |
| 164 | Evpl | na | 44048 | -0.081 | 0.0543 | No |
| 165 | Krt10 | na | 44054 | -0.081 | 0.0619 | No |
| 166 | Dsc1 | na | 45097 | -0.114 | 0.0541 | No |
| 167 | Krt83 | na | 46191 | -0.147 | 0.0485 | No |
| 168 | Krt23 | na | 47304 | -0.182 | 0.0459 | No |
| 169 | Dsg2 | na | 49869 | -0.278 | 0.0265 | No |
| 170 | Spink5 | na | 50233 | -0.293 | 0.0476 | No |
| 171 | Cdsn | na | 50841 | -0.318 | 0.0668 | No |
| 172 | Cela2a | na | 51231 | -0.336 | 0.0916 | No |