Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___PanKO_vs_ClusterKO.RPKM_matrix___PanKO_vs_ClusterKO.cls #PanKO_versus_ClusterKO |
| Phenotype | RPKM_matrix___PanKO_vs_ClusterKO.cls#PanKO_versus_ClusterKO |
| Upregulated in class | PanKO |
| GeneSet | WP_CHOLESTEROL_BIOSYNTHESIS |
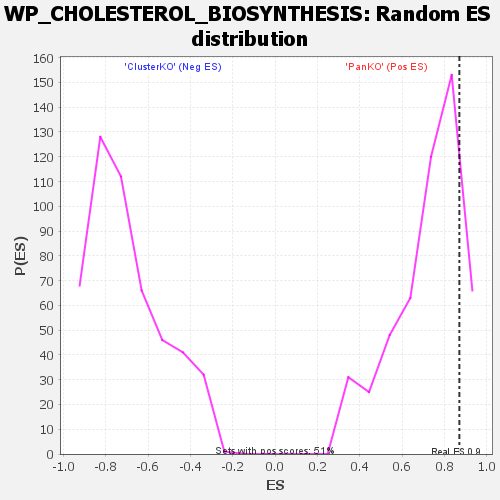
| Enrichment Score (ES) | 0.86992276 |
| Normalized Enrichment Score (NES) | 1.2038145 |
| Nominal p-value | 0.1541502 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.897 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Nsdhl | na | 104 | 0.723 | 0.1267 | Yes |
| 2 | Cyp51 | na | 250 | 0.637 | 0.2374 | Yes |
| 3 | Pmvk | na | 618 | 0.535 | 0.3260 | Yes |
| 4 | Fdps | na | 818 | 0.499 | 0.4111 | Yes |
| 5 | Sqle | na | 880 | 0.485 | 0.4962 | Yes |
| 6 | Lss | na | 1014 | 0.464 | 0.5764 | Yes |
| 7 | Hmgcr | na | 2035 | 0.363 | 0.6228 | Yes |
| 8 | Fdft1 | na | 2262 | 0.346 | 0.6802 | Yes |
| 9 | Mvd | na | 2792 | 0.311 | 0.7261 | Yes |
| 10 | Dhcr7 | na | 3034 | 0.297 | 0.7746 | Yes |
| 11 | Hmgcs1 | na | 3168 | 0.288 | 0.8234 | Yes |
| 12 | Idi1 | na | 4632 | 0.217 | 0.8361 | Yes |
| 13 | Msmo1 | na | 4824 | 0.209 | 0.8699 | Yes |
| 14 | Sc5d | na | 44696 | -0.102 | 0.1806 | No |
| 15 | Mvk | na | 46257 | -0.150 | 0.1796 | No |