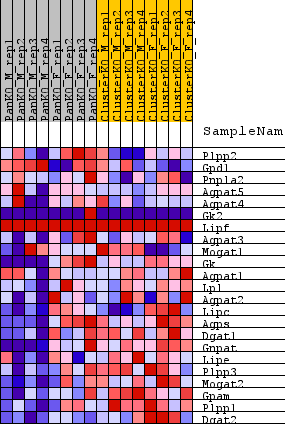
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___PanKO_vs_ClusterKO.RPKM_matrix___PanKO_vs_ClusterKO.cls #PanKO_versus_ClusterKO |
| Phenotype | RPKM_matrix___PanKO_vs_ClusterKO.cls#PanKO_versus_ClusterKO |
| Upregulated in class | ClusterKO |
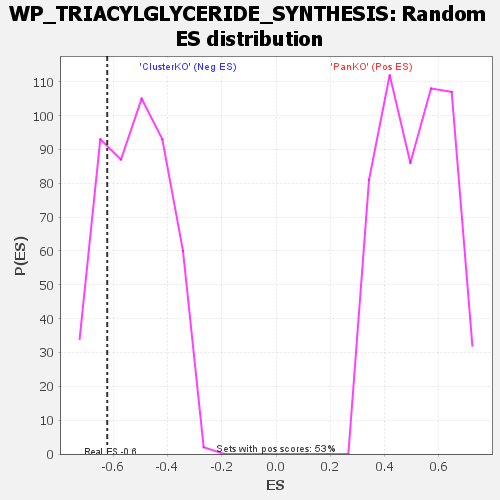
| GeneSet | WP_TRIACYLGLYCERIDE_SYNTHESIS |
| Enrichment Score (ES) | -0.62142855 |
| Normalized Enrichment Score (NES) | -1.1984016 |
| Nominal p-value | 0.23839663 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.888 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Plpp2 | na | 1435 | 0.417 | 0.0395 | No |
| 2 | Gpd1 | na | 2095 | 0.358 | 0.0837 | No |
| 3 | Pnpla2 | na | 4522 | 0.221 | 0.0752 | No |
| 4 | Agpat5 | na | 6359 | 0.151 | 0.0662 | No |
| 5 | Agpat4 | na | 10167 | 0.032 | 0.0037 | No |
| 6 | Gk2 | na | 11891 | 0.002 | -0.0265 | No |
| 7 | Lipf | na | 34351 | 0.000 | -0.4251 | No |
| 8 | Agpat3 | na | 40251 | -0.009 | -0.5283 | No |
| 9 | Mogat1 | na | 44005 | -0.080 | -0.5824 | No |
| 10 | Gk | na | 46204 | -0.147 | -0.5984 | Yes |
| 11 | Agpat1 | na | 46587 | -0.160 | -0.5802 | Yes |
| 12 | Lpl | na | 46984 | -0.173 | -0.5604 | Yes |
| 13 | Agpat2 | na | 47857 | -0.202 | -0.5443 | Yes |
| 14 | Lipc | na | 48239 | -0.215 | -0.5175 | Yes |
| 15 | Agps | na | 49294 | -0.254 | -0.4965 | Yes |
| 16 | Dgat1 | na | 51381 | -0.343 | -0.4801 | Yes |
| 17 | Gnpat | na | 52548 | -0.401 | -0.4382 | Yes |
| 18 | Lipe | na | 52815 | -0.416 | -0.3779 | Yes |
| 19 | Plpp3 | na | 53298 | -0.447 | -0.3167 | Yes |
| 20 | Mogat2 | na | 54365 | -0.527 | -0.2534 | Yes |
| 21 | Gpam | na | 54867 | -0.580 | -0.1718 | Yes |
| 22 | Plpp1 | na | 55005 | -0.598 | -0.0810 | Yes |
| 23 | Dgat2 | na | 55548 | -0.675 | 0.0147 | Yes |