Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___PanKO_vs_Cyp2c44KO.RPKM_matrix___PanKO_vs_Cyp2c44KO.cls #PanKO_versus_Cyp2c44KO |
| Phenotype | RPKM_matrix___PanKO_vs_Cyp2c44KO.cls#PanKO_versus_Cyp2c44KO |
| Upregulated in class | Cyp2c44KO |
| GeneSet | GOCC_MYOFILAMENT |
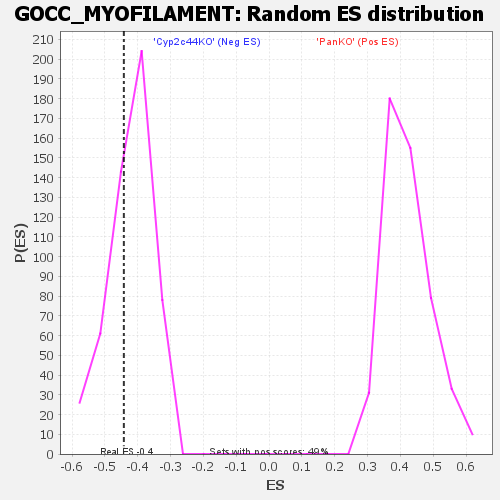
| Enrichment Score (ES) | -0.44212297 |
| Normalized Enrichment Score (NES) | -1.0510885 |
| Nominal p-value | 0.30273438 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.995 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Tpm1 | na | 47 | 1.671 | 0.1315 | No |
| 2 | Tnni2 | na | 1868 | 0.791 | 0.1618 | No |
| 3 | Tmod3 | na | 2975 | 0.636 | 0.1925 | No |
| 4 | Tnnt2 | na | 4167 | 0.516 | 0.2122 | No |
| 5 | Trim32 | na | 5784 | 0.399 | 0.2151 | No |
| 6 | Tpm2 | na | 6587 | 0.351 | 0.2287 | No |
| 7 | Lmod1 | na | 6977 | 0.329 | 0.2478 | No |
| 8 | Acta1 | na | 7987 | 0.278 | 0.2519 | No |
| 9 | Tnnt3 | na | 10261 | 0.177 | 0.2256 | No |
| 10 | Tmod1 | na | 11225 | 0.131 | 0.2189 | No |
| 11 | Tnni1 | na | 15677 | 0.003 | 0.1401 | No |
| 12 | Mybphl | na | 43068 | -0.002 | -0.3458 | No |
| 13 | Tnnc1 | na | 46971 | -0.072 | -0.4093 | No |
| 14 | Actn2 | na | 47392 | -0.092 | -0.4095 | No |
| 15 | Tnnc2 | na | 48580 | -0.143 | -0.4193 | No |
| 16 | Obscn | na | 48741 | -0.151 | -0.4102 | No |
| 17 | Actn3 | na | 49021 | -0.166 | -0.4020 | No |
| 18 | Tnnt1 | na | 51281 | -0.287 | -0.4194 | Yes |
| 19 | Fhod3 | na | 51620 | -0.309 | -0.4010 | Yes |
| 20 | Tmod2 | na | 51632 | -0.309 | -0.3767 | Yes |
| 21 | Ttn | na | 51717 | -0.315 | -0.3532 | Yes |
| 22 | Mybpc3 | na | 52747 | -0.392 | -0.3405 | Yes |
| 23 | Tnni3 | na | 54488 | -0.586 | -0.3249 | Yes |
| 24 | Lmod2 | na | 54507 | -0.589 | -0.2786 | Yes |
| 25 | Lmod3 | na | 54547 | -0.598 | -0.2320 | Yes |
| 26 | Tmod4 | na | 55145 | -0.717 | -0.1859 | Yes |
| 27 | Neb | na | 56371 | -2.624 | 0.0001 | Yes |