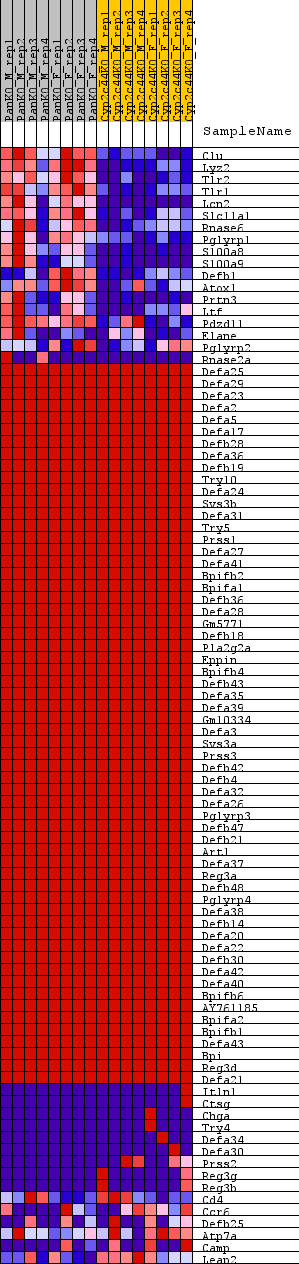
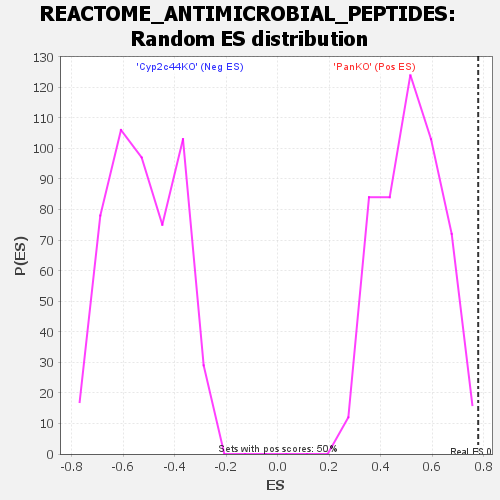
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___PanKO_vs_Cyp2c44KO.RPKM_matrix___PanKO_vs_Cyp2c44KO.cls #PanKO_versus_Cyp2c44KO |
| Phenotype | RPKM_matrix___PanKO_vs_Cyp2c44KO.cls#PanKO_versus_Cyp2c44KO |
| Upregulated in class | PanKO |
| GeneSet | REACTOME_ANTIMICROBIAL_PEPTIDES |
| Enrichment Score (ES) | 0.7787232 |
| Normalized Enrichment Score (NES) | 1.5107188 |
| Nominal p-value | 0.004040404 |
| FDR q-value | 0.37171727 |
| FWER p-Value | 0.222 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Clu | na | 99 | 1.562 | 0.0902 | Yes |
| 2 | Lyz2 | na | 106 | 1.551 | 0.1814 | Yes |
| 3 | Tlr2 | na | 191 | 1.424 | 0.2637 | Yes |
| 4 | Tlr1 | na | 246 | 1.379 | 0.3439 | Yes |
| 5 | Lcn2 | na | 653 | 1.133 | 0.4034 | Yes |
| 6 | Slc11a1 | na | 702 | 1.113 | 0.4681 | Yes |
| 7 | Rnase6 | na | 1322 | 0.905 | 0.5103 | Yes |
| 8 | Pglyrp1 | na | 1550 | 0.856 | 0.5567 | Yes |
| 9 | S100a8 | na | 1960 | 0.775 | 0.5950 | Yes |
| 10 | S100a9 | na | 2063 | 0.762 | 0.6380 | Yes |
| 11 | Defb1 | na | 2533 | 0.698 | 0.6708 | Yes |
| 12 | Atox1 | na | 3043 | 0.628 | 0.6987 | Yes |
| 13 | Prtn3 | na | 3167 | 0.614 | 0.7327 | Yes |
| 14 | Ltf | na | 3260 | 0.604 | 0.7666 | Yes |
| 15 | Pdzd11 | na | 4260 | 0.508 | 0.7787 | Yes |
| 16 | Elane | na | 6852 | 0.337 | 0.7525 | No |
| 17 | Pglyrp2 | na | 9959 | 0.191 | 0.7086 | No |
| 18 | Rnase2a | na | 12643 | 0.078 | 0.6655 | No |
| 19 | Defa25 | na | 16748 | 0.000 | 0.5926 | No |
| 20 | Defa29 | na | 17092 | 0.000 | 0.5865 | No |
| 21 | Defa23 | na | 17560 | 0.000 | 0.5782 | No |
| 22 | Defa2 | na | 18053 | 0.000 | 0.5695 | No |
| 23 | Defa5 | na | 18146 | 0.000 | 0.5679 | No |
| 24 | Defa17 | na | 18876 | 0.000 | 0.5549 | No |
| 25 | Defb28 | na | 19083 | 0.000 | 0.5512 | No |
| 26 | Defa36 | na | 19205 | 0.000 | 0.5491 | No |
| 27 | Defb19 | na | 19907 | 0.000 | 0.5366 | No |
| 28 | Try10 | na | 20668 | 0.000 | 0.5231 | No |
| 29 | Defa24 | na | 21007 | 0.000 | 0.5171 | No |
| 30 | Svs3b | na | 21405 | 0.000 | 0.5101 | No |
| 31 | Defa31 | na | 21726 | 0.000 | 0.5044 | No |
| 32 | Try5 | na | 22057 | 0.000 | 0.4985 | No |
| 33 | Prss1 | na | 22705 | 0.000 | 0.4870 | No |
| 34 | Defa27 | na | 23161 | 0.000 | 0.4790 | No |
| 35 | Defa41 | na | 23597 | 0.000 | 0.4712 | No |
| 36 | Bpifb2 | na | 23725 | 0.000 | 0.4690 | No |
| 37 | Bpifa1 | na | 24317 | 0.000 | 0.4585 | No |
| 38 | Defb36 | na | 24968 | 0.000 | 0.4469 | No |
| 39 | Defa28 | na | 25104 | 0.000 | 0.4445 | No |
| 40 | Gm5771 | na | 25644 | 0.000 | 0.4349 | No |
| 41 | Defb18 | na | 26929 | 0.000 | 0.4121 | No |
| 42 | Pla2g2a | na | 27016 | 0.000 | 0.4106 | No |
| 43 | Eppin | na | 27929 | 0.000 | 0.3944 | No |
| 44 | Bpifb4 | na | 28174 | 0.000 | 0.3901 | No |
| 45 | Defb43 | na | 29256 | 0.000 | 0.3709 | No |
| 46 | Defa35 | na | 29341 | 0.000 | 0.3694 | No |
| 47 | Defa39 | na | 29617 | 0.000 | 0.3645 | No |
| 48 | Gm10334 | na | 30417 | 0.000 | 0.3503 | No |
| 49 | Defa3 | na | 30580 | 0.000 | 0.3474 | No |
| 50 | Svs3a | na | 30639 | 0.000 | 0.3464 | No |
| 51 | Prss3 | na | 30853 | 0.000 | 0.3426 | No |
| 52 | Defb42 | na | 30939 | 0.000 | 0.3411 | No |
| 53 | Defb4 | na | 31409 | 0.000 | 0.3328 | No |
| 54 | Defa32 | na | 31673 | 0.000 | 0.3281 | No |
| 55 | Defa26 | na | 32153 | 0.000 | 0.3196 | No |
| 56 | Pglyrp3 | na | 32590 | 0.000 | 0.3118 | No |
| 57 | Defb47 | na | 32629 | 0.000 | 0.3111 | No |
| 58 | Defb21 | na | 33182 | 0.000 | 0.3013 | No |
| 59 | Art1 | na | 33278 | 0.000 | 0.2997 | No |
| 60 | Defa37 | na | 33535 | 0.000 | 0.2951 | No |
| 61 | Reg3a | na | 33591 | 0.000 | 0.2941 | No |
| 62 | Defb48 | na | 34609 | 0.000 | 0.2761 | No |
| 63 | Pglyrp4 | na | 34959 | 0.000 | 0.2699 | No |
| 64 | Defa38 | na | 35521 | 0.000 | 0.2599 | No |
| 65 | Defb14 | na | 35812 | 0.000 | 0.2547 | No |
| 66 | Defa20 | na | 35878 | 0.000 | 0.2536 | No |
| 67 | Defa22 | na | 35905 | 0.000 | 0.2531 | No |
| 68 | Defb30 | na | 36347 | 0.000 | 0.2453 | No |
| 69 | Defa42 | na | 37072 | 0.000 | 0.2324 | No |
| 70 | Defa40 | na | 37253 | 0.000 | 0.2292 | No |
| 71 | Bpifb6 | na | 37560 | 0.000 | 0.2238 | No |
| 72 | AY761185 | na | 37937 | 0.000 | 0.2171 | No |
| 73 | Bpifa2 | na | 38020 | 0.000 | 0.2157 | No |
| 74 | Bpifb1 | na | 38299 | 0.000 | 0.2107 | No |
| 75 | Defa43 | na | 38803 | 0.000 | 0.2018 | No |
| 76 | Bpi | na | 39461 | 0.000 | 0.1901 | No |
| 77 | Reg3d | na | 41761 | 0.000 | 0.1493 | No |
| 78 | Defa21 | na | 42069 | 0.000 | 0.1438 | No |
| 79 | Itln1 | na | 43122 | -0.002 | 0.1252 | No |
| 80 | Ctsg | na | 43207 | -0.003 | 0.1239 | No |
| 81 | Chga | na | 43901 | -0.007 | 0.1120 | No |
| 82 | Try4 | na | 44027 | -0.008 | 0.1102 | No |
| 83 | Defa34 | na | 45470 | -0.029 | 0.0863 | No |
| 84 | Defa30 | na | 45492 | -0.030 | 0.0877 | No |
| 85 | Prss2 | na | 45809 | -0.037 | 0.0843 | No |
| 86 | Reg3g | na | 46850 | -0.068 | 0.0698 | No |
| 87 | Reg3b | na | 46975 | -0.072 | 0.0719 | No |
| 88 | Cd4 | na | 47878 | -0.113 | 0.0625 | No |
| 89 | Ccr6 | na | 49324 | -0.179 | 0.0473 | No |
| 90 | Defb25 | na | 49595 | -0.193 | 0.0539 | No |
| 91 | Atp7a | na | 49867 | -0.206 | 0.0612 | No |
| 92 | Camp | na | 51249 | -0.285 | 0.0535 | No |
| 93 | Leap2 | na | 54812 | -0.639 | 0.0278 | No |