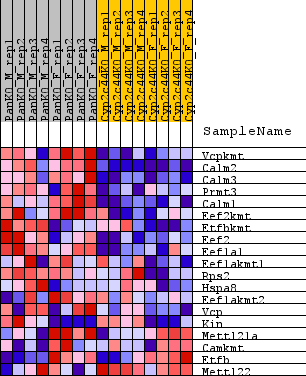
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___PanKO_vs_Cyp2c44KO.RPKM_matrix___PanKO_vs_Cyp2c44KO.cls #PanKO_versus_Cyp2c44KO |
| Phenotype | RPKM_matrix___PanKO_vs_Cyp2c44KO.cls#PanKO_versus_Cyp2c44KO |
| Upregulated in class | PanKO |
| GeneSet | REACTOME_PROTEIN_METHYLATION |
| Enrichment Score (ES) | 0.75597817 |
| Normalized Enrichment Score (NES) | 1.491811 |
| Nominal p-value | 0.008179959 |
| FDR q-value | 0.30579185 |
| FWER p-Value | 0.332 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Vcpkmt | na | 649 | 1.136 | 0.1312 | Yes |
| 2 | Calm2 | na | 1069 | 0.978 | 0.2466 | Yes |
| 3 | Calm3 | na | 1707 | 0.822 | 0.3386 | Yes |
| 4 | Prmt3 | na | 1780 | 0.806 | 0.4387 | Yes |
| 5 | Calm1 | na | 2707 | 0.673 | 0.5067 | Yes |
| 6 | Eef2kmt | na | 3933 | 0.538 | 0.5526 | Yes |
| 7 | Etfbkmt | na | 6055 | 0.384 | 0.5632 | Yes |
| 8 | Eef2 | na | 6110 | 0.381 | 0.6101 | Yes |
| 9 | Eef1a1 | na | 6859 | 0.336 | 0.6391 | Yes |
| 10 | Eef1akmt1 | na | 7984 | 0.279 | 0.6541 | Yes |
| 11 | Rps2 | na | 8106 | 0.272 | 0.6862 | Yes |
| 12 | Hspa8 | na | 8562 | 0.250 | 0.7096 | Yes |
| 13 | Eef1akmt2 | na | 8853 | 0.238 | 0.7343 | Yes |
| 14 | Vcp | na | 9208 | 0.222 | 0.7560 | Yes |
| 15 | Kin | na | 13212 | 0.059 | 0.6924 | No |
| 16 | Mettl21a | na | 14133 | 0.029 | 0.6798 | No |
| 17 | Camkmt | na | 45585 | -0.032 | 0.1257 | No |
| 18 | Etfb | na | 49052 | -0.167 | 0.0851 | No |
| 19 | Mettl22 | na | 52289 | -0.357 | 0.0725 | No |