Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___PanKO_vs_WT.RPKM_matrix___PanKO_vs_WT.cls#PanKO_versus_WT |
| Phenotype | RPKM_matrix___PanKO_vs_WT.cls#PanKO_versus_WT |
| Upregulated in class | PanKO |
| GeneSet | GOCC_AGGRESOME |
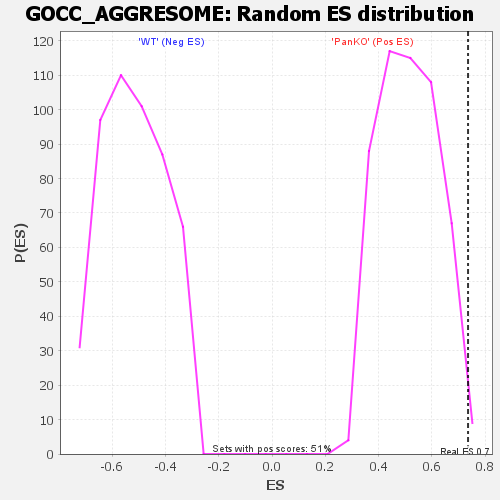
| Enrichment Score (ES) | 0.73611265 |
| Normalized Enrichment Score (NES) | 1.4286925 |
| Nominal p-value | 0.001968504 |
| FDR q-value | 0.2427959 |
| FWER p-Value | 0.511 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Fgr | na | 36 | 1.666 | 0.0971 | Yes |
| 2 | Clu | na | 283 | 1.345 | 0.1716 | Yes |
| 3 | Sqstm1 | na | 862 | 1.083 | 0.2248 | Yes |
| 4 | Prdm16 | na | 1049 | 1.024 | 0.2816 | Yes |
| 5 | Slc2a3 | na | 1614 | 0.887 | 0.3236 | Yes |
| 6 | Eps15 | na | 1758 | 0.855 | 0.3712 | Yes |
| 7 | Ubd | na | 1785 | 0.850 | 0.4207 | Yes |
| 8 | Pold1 | na | 3011 | 0.674 | 0.4385 | Yes |
| 9 | Xrn2 | na | 3053 | 0.670 | 0.4770 | Yes |
| 10 | Rangap1 | na | 3427 | 0.629 | 0.5073 | Yes |
| 11 | Urb2 | na | 3597 | 0.610 | 0.5400 | Yes |
| 12 | Psap | na | 3603 | 0.609 | 0.5757 | Yes |
| 13 | Hspa1a | na | 3863 | 0.587 | 0.6055 | Yes |
| 14 | Cabin1 | na | 4304 | 0.550 | 0.6299 | Yes |
| 15 | Trim50 | na | 5252 | 0.476 | 0.6410 | Yes |
| 16 | Hspb7 | na | 5278 | 0.475 | 0.6684 | Yes |
| 17 | Trim37 | na | 5526 | 0.458 | 0.6909 | Yes |
| 18 | Dvl2 | na | 6026 | 0.426 | 0.7071 | Yes |
| 19 | Psen1 | na | 6337 | 0.405 | 0.7253 | Yes |
| 20 | Eef2 | na | 7770 | 0.324 | 0.7189 | Yes |
| 21 | Zbtb14 | na | 7857 | 0.319 | 0.7361 | Yes |
| 22 | Card14 | na | 9400 | 0.243 | 0.7230 | No |
| 23 | Stradb | na | 13333 | 0.084 | 0.6582 | No |
| 24 | Klf8 | na | 13641 | 0.075 | 0.6571 | No |
| 25 | Hspa1b | na | 13775 | 0.071 | 0.6589 | No |
| 26 | Klhl14 | na | 17012 | 0.011 | 0.6021 | No |
| 27 | Tdp2 | na | 17527 | 0.007 | 0.5934 | No |
| 28 | Hoxd3 | na | 23478 | 0.000 | 0.4878 | No |
| 29 | Hoxc9 | na | 30176 | 0.000 | 0.3689 | No |
| 30 | Ubqln1 | na | 48016 | -0.020 | 0.0534 | No |
| 31 | Sfmbt2 | na | 49384 | -0.064 | 0.0330 | No |
| 32 | Rnf32 | na | 49712 | -0.077 | 0.0317 | No |
| 33 | Edem1 | na | 50855 | -0.132 | 0.0191 | No |
| 34 | Hdac6 | na | 52333 | -0.210 | 0.0052 | No |
| 35 | Prkcq | na | 53164 | -0.263 | 0.0059 | No |
| 36 | Prkn | na | 56220 | -0.871 | 0.0028 | No |