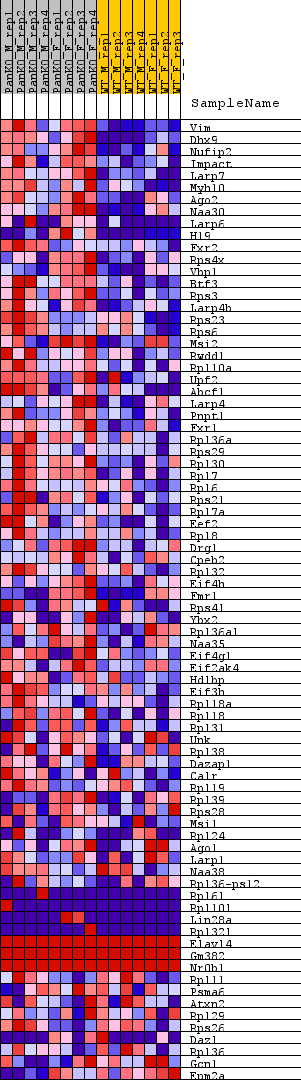
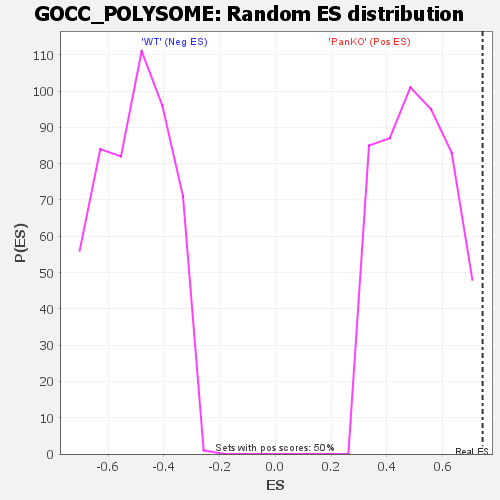
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___PanKO_vs_WT.RPKM_matrix___PanKO_vs_WT.cls#PanKO_versus_WT |
| Phenotype | RPKM_matrix___PanKO_vs_WT.cls#PanKO_versus_WT |
| Upregulated in class | PanKO |
| GeneSet | GOCC_POLYSOME |
| Enrichment Score (ES) | 0.74330735 |
| Normalized Enrichment Score (NES) | 1.4679848 |
| Nominal p-value | 0.002004008 |
| FDR q-value | 0.43293366 |
| FWER p-Value | 0.305 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Vim | na | 141 | 1.472 | 0.0554 | Yes |
| 2 | Dhx9 | na | 1150 | 0.994 | 0.0765 | Yes |
| 3 | Nufip2 | na | 1786 | 0.850 | 0.0987 | Yes |
| 4 | Impact | na | 1806 | 0.846 | 0.1316 | Yes |
| 5 | Larp7 | na | 2059 | 0.799 | 0.1586 | Yes |
| 6 | Myh10 | na | 2300 | 0.762 | 0.1843 | Yes |
| 7 | Ago2 | na | 2615 | 0.723 | 0.2071 | Yes |
| 8 | Naa30 | na | 2910 | 0.686 | 0.2288 | Yes |
| 9 | Larp6 | na | 3315 | 0.641 | 0.2469 | Yes |
| 10 | H19 | na | 3598 | 0.610 | 0.2659 | Yes |
| 11 | Fxr2 | na | 3626 | 0.608 | 0.2893 | Yes |
| 12 | Rps4x | na | 4295 | 0.550 | 0.2990 | Yes |
| 13 | Vbp1 | na | 4615 | 0.523 | 0.3139 | Yes |
| 14 | Btf3 | na | 5367 | 0.469 | 0.3190 | Yes |
| 15 | Rps3 | na | 5512 | 0.459 | 0.3345 | Yes |
| 16 | Larp4b | na | 5569 | 0.455 | 0.3515 | Yes |
| 17 | Rps23 | na | 5973 | 0.429 | 0.3612 | Yes |
| 18 | Rps6 | na | 6068 | 0.423 | 0.3761 | Yes |
| 19 | Msi2 | na | 6086 | 0.422 | 0.3924 | Yes |
| 20 | Rwdd1 | na | 6143 | 0.418 | 0.4078 | Yes |
| 21 | Rpl10a | na | 6266 | 0.411 | 0.4218 | Yes |
| 22 | Upf2 | na | 6554 | 0.391 | 0.4321 | Yes |
| 23 | Abcf1 | na | 6569 | 0.390 | 0.4472 | Yes |
| 24 | Larp4 | na | 6715 | 0.383 | 0.4597 | Yes |
| 25 | Pnpt1 | na | 6734 | 0.382 | 0.4744 | Yes |
| 26 | Fxr1 | na | 6849 | 0.375 | 0.4871 | Yes |
| 27 | Rpl36a | na | 7215 | 0.354 | 0.4945 | Yes |
| 28 | Rps29 | na | 7375 | 0.345 | 0.5053 | Yes |
| 29 | Rpl30 | na | 7591 | 0.334 | 0.5146 | Yes |
| 30 | Rpl7 | na | 7605 | 0.333 | 0.5275 | Yes |
| 31 | Rpl6 | na | 7687 | 0.328 | 0.5389 | Yes |
| 32 | Rps21 | na | 7694 | 0.328 | 0.5517 | Yes |
| 33 | Rpl7a | na | 7723 | 0.327 | 0.5640 | Yes |
| 34 | Eef2 | na | 7770 | 0.324 | 0.5760 | Yes |
| 35 | Rpl8 | na | 7784 | 0.323 | 0.5884 | Yes |
| 36 | Drg1 | na | 7878 | 0.318 | 0.5993 | Yes |
| 37 | Cpeb2 | na | 7931 | 0.315 | 0.6107 | Yes |
| 38 | Rpl32 | na | 8192 | 0.302 | 0.6180 | Yes |
| 39 | Eif4h | na | 8560 | 0.283 | 0.6226 | Yes |
| 40 | Fmr1 | na | 8605 | 0.281 | 0.6329 | Yes |
| 41 | Rps4l | na | 8611 | 0.281 | 0.6438 | Yes |
| 42 | Ybx2 | na | 8795 | 0.272 | 0.6513 | Yes |
| 43 | Rpl36al | na | 8843 | 0.270 | 0.6610 | Yes |
| 44 | Naa35 | na | 9193 | 0.253 | 0.6648 | Yes |
| 45 | Eif4g1 | na | 9250 | 0.251 | 0.6737 | Yes |
| 46 | Eif2ak4 | na | 9277 | 0.249 | 0.6830 | Yes |
| 47 | Hdlbp | na | 9366 | 0.245 | 0.6911 | Yes |
| 48 | Eif3h | na | 9607 | 0.234 | 0.6960 | Yes |
| 49 | Rpl18a | na | 9748 | 0.228 | 0.7025 | Yes |
| 50 | Rpl18 | na | 9764 | 0.227 | 0.7111 | Yes |
| 51 | Rpl31 | na | 9937 | 0.220 | 0.7168 | Yes |
| 52 | Unk | na | 10099 | 0.213 | 0.7223 | Yes |
| 53 | Rpl38 | na | 10102 | 0.213 | 0.7306 | Yes |
| 54 | Dazap1 | na | 10524 | 0.193 | 0.7308 | Yes |
| 55 | Calr | na | 10672 | 0.187 | 0.7355 | Yes |
| 56 | Rpl19 | na | 11086 | 0.169 | 0.7348 | Yes |
| 57 | Rpl39 | na | 11139 | 0.167 | 0.7405 | Yes |
| 58 | Rps28 | na | 11331 | 0.158 | 0.7433 | Yes |
| 59 | Msi1 | na | 11830 | 0.137 | 0.7398 | No |
| 60 | Rpl24 | na | 14073 | 0.062 | 0.7024 | No |
| 61 | Ago1 | na | 14942 | 0.042 | 0.6887 | No |
| 62 | Larp1 | na | 15089 | 0.039 | 0.6876 | No |
| 63 | Naa38 | na | 15557 | 0.030 | 0.6805 | No |
| 64 | Rpl36-ps12 | na | 15605 | 0.029 | 0.6808 | No |
| 65 | Rpl6l | na | 16120 | 0.021 | 0.6725 | No |
| 66 | Rpl10l | na | 16170 | 0.021 | 0.6725 | No |
| 67 | Lin28a | na | 16847 | 0.012 | 0.6609 | No |
| 68 | Rpl32l | na | 16992 | 0.011 | 0.6588 | No |
| 69 | Elavl4 | na | 19114 | 0.000 | 0.6211 | No |
| 70 | Gm382 | na | 35470 | 0.000 | 0.3306 | No |
| 71 | Nr0b1 | na | 38268 | 0.000 | 0.2810 | No |
| 72 | Rpl11 | na | 47613 | -0.011 | 0.1154 | No |
| 73 | Psma6 | na | 47843 | -0.016 | 0.1119 | No |
| 74 | Atxn2 | na | 48018 | -0.020 | 0.1096 | No |
| 75 | Rpl29 | na | 48051 | -0.021 | 0.1099 | No |
| 76 | Rps26 | na | 48237 | -0.025 | 0.1076 | No |
| 77 | Dazl | na | 49327 | -0.061 | 0.0906 | No |
| 78 | Rpl36 | na | 49342 | -0.062 | 0.0928 | No |
| 79 | Gcn1 | na | 51703 | -0.176 | 0.0579 | No |
| 80 | Epm2a | na | 55857 | -0.639 | 0.0092 | No |