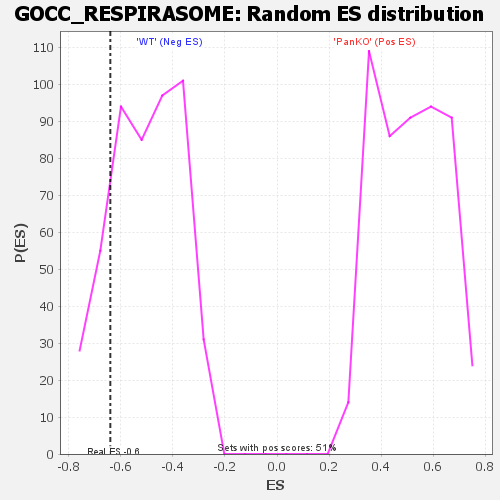
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___PanKO_vs_WT.RPKM_matrix___PanKO_vs_WT.cls#PanKO_versus_WT |
| Phenotype | RPKM_matrix___PanKO_vs_WT.cls#PanKO_versus_WT |
| Upregulated in class | WT |
| GeneSet | GOCC_RESPIRASOME |
| Enrichment Score (ES) | -0.63968945 |
| Normalized Enrichment Score (NES) | -1.2687724 |
| Nominal p-value | 0.16293278 |
| FDR q-value | 0.61727804 |
| FWER p-Value | 0.885 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Ttc19 | na | 3974 | 0.578 | -0.0444 | No |
| 2 | Cox6a2 | na | 4272 | 0.552 | -0.0246 | No |
| 3 | Ndufb4c | na | 5219 | 0.478 | -0.0197 | No |
| 4 | Cox6b2 | na | 5976 | 0.429 | -0.0136 | No |
| 5 | Cox6c2 | na | 6412 | 0.401 | -0.0031 | No |
| 6 | Cox4i2 | na | 6495 | 0.395 | 0.0134 | No |
| 7 | Uqcrh-ps1 | na | 6618 | 0.388 | 0.0288 | No |
| 8 | AA467197 | na | 8464 | 0.287 | 0.0091 | No |
| 9 | Uqcrh | na | 9967 | 0.219 | -0.0076 | No |
| 10 | Ndufa1 | na | 10992 | 0.173 | -0.0180 | No |
| 11 | Cox7a2l | na | 11104 | 0.169 | -0.0123 | No |
| 12 | 1700066M21Rik | na | 11658 | 0.144 | -0.0156 | No |
| 13 | Ndufa4 | na | 11696 | 0.142 | -0.0098 | No |
| 14 | Cox7a1 | na | 11883 | 0.134 | -0.0070 | No |
| 15 | Cox7a2 | na | 12062 | 0.128 | -0.0043 | No |
| 16 | Ndufa11b | na | 12111 | 0.126 | 0.0006 | No |
| 17 | Ndufs6 | na | 13402 | 0.082 | -0.0186 | No |
| 18 | Ndufs4 | na | 13481 | 0.079 | -0.0164 | No |
| 19 | Rab5if | na | 13862 | 0.068 | -0.0201 | No |
| 20 | Ndufs6b | na | 14670 | 0.048 | -0.0323 | No |
| 21 | mt-Co2 | na | 14893 | 0.043 | -0.0343 | No |
| 22 | Ndufb4b | na | 15167 | 0.037 | -0.0374 | No |
| 23 | Ndufs2 | na | 15286 | 0.035 | -0.0379 | No |
| 24 | Dmac1 | na | 15500 | 0.031 | -0.0403 | No |
| 25 | Cox4i1 | na | 15999 | 0.023 | -0.0481 | No |
| 26 | Ndufa8 | na | 17357 | 0.008 | -0.0718 | No |
| 27 | Uqcc4 | na | 17633 | 0.006 | -0.0764 | No |
| 28 | mt-Nd4l | na | 18678 | 0.000 | -0.0950 | No |
| 29 | mt-Co3 | na | 25517 | 0.000 | -0.2165 | No |
| 30 | Cox8c | na | 26692 | 0.000 | -0.2374 | No |
| 31 | Ndufb11b | na | 34760 | 0.000 | -0.3807 | No |
| 32 | Cyct | na | 36838 | 0.000 | -0.4176 | No |
| 33 | Cox7b2 | na | 44947 | 0.000 | -0.5617 | No |
| 34 | Ndufb6 | na | 48156 | -0.023 | -0.6177 | No |
| 35 | Cox7c | na | 48466 | -0.032 | -0.6217 | No |
| 36 | Ndufa2 | na | 48770 | -0.041 | -0.6253 | No |
| 37 | Uqcr10 | na | 48981 | -0.048 | -0.6268 | No |
| 38 | Cox5a | na | 49104 | -0.052 | -0.6266 | No |
| 39 | Ndufa7 | na | 49301 | -0.061 | -0.6273 | No |
| 40 | Sdhc | na | 49325 | -0.061 | -0.6250 | No |
| 41 | Ndufc1 | na | 50155 | -0.097 | -0.6353 | Yes |
| 42 | Higd1a | na | 50356 | -0.105 | -0.6341 | Yes |
| 43 | Ndufv2 | na | 50368 | -0.106 | -0.6295 | Yes |
| 44 | Uqcrc2 | na | 50443 | -0.110 | -0.6258 | Yes |
| 45 | Ndufs3 | na | 50522 | -0.113 | -0.6220 | Yes |
| 46 | Sdha | na | 50534 | -0.114 | -0.6170 | Yes |
| 47 | Ndufv1 | na | 50781 | -0.128 | -0.6156 | Yes |
| 48 | Ndufc2 | na | 50890 | -0.133 | -0.6114 | Yes |
| 49 | Ndufa12 | na | 50892 | -0.133 | -0.6054 | Yes |
| 50 | Sdhd | na | 50965 | -0.137 | -0.6004 | Yes |
| 51 | Ndufa3 | na | 50972 | -0.137 | -0.5943 | Yes |
| 52 | Ndufs1 | na | 50976 | -0.138 | -0.5881 | Yes |
| 53 | Uqcc3 | na | 51034 | -0.140 | -0.5827 | Yes |
| 54 | Uqcrq | na | 51144 | -0.145 | -0.5781 | Yes |
| 55 | Ndufab1 | na | 51157 | -0.146 | -0.5716 | Yes |
| 56 | Uqcrb | na | 51173 | -0.147 | -0.5652 | Yes |
| 57 | Ndufb7 | na | 51551 | -0.168 | -0.5643 | Yes |
| 58 | Cox7b | na | 51569 | -0.169 | -0.5569 | Yes |
| 59 | Cox5b | na | 51643 | -0.173 | -0.5503 | Yes |
| 60 | Stmp1 | na | 51672 | -0.175 | -0.5429 | Yes |
| 61 | Ndufb3 | na | 51845 | -0.183 | -0.5376 | Yes |
| 62 | Ndufa9 | na | 51916 | -0.187 | -0.5304 | Yes |
| 63 | Ndufa13 | na | 51947 | -0.188 | -0.5224 | Yes |
| 64 | Ndufab1-ps | na | 52035 | -0.193 | -0.5151 | Yes |
| 65 | Cox6b1 | na | 52132 | -0.197 | -0.5079 | Yes |
| 66 | Uqcrfs1 | na | 52138 | -0.197 | -0.4990 | Yes |
| 67 | Ndufa11 | na | 52164 | -0.199 | -0.4905 | Yes |
| 68 | Uqcrc1 | na | 52316 | -0.208 | -0.4837 | Yes |
| 69 | Uqcr11 | na | 52397 | -0.214 | -0.4754 | Yes |
| 70 | Ndufb2 | na | 52451 | -0.217 | -0.4665 | Yes |
| 71 | Cox8a | na | 52452 | -0.217 | -0.4566 | Yes |
| 72 | Ndufb1 | na | 52458 | -0.217 | -0.4468 | Yes |
| 73 | Foxred1 | na | 52484 | -0.219 | -0.4374 | Yes |
| 74 | mt-Nd6 | na | 52499 | -0.220 | -0.4276 | Yes |
| 75 | Ndufs5 | na | 52616 | -0.226 | -0.4194 | Yes |
| 76 | Ndufb8 | na | 52782 | -0.237 | -0.4116 | Yes |
| 77 | Cox6a1 | na | 52916 | -0.247 | -0.4027 | Yes |
| 78 | mt-Nd2 | na | 52917 | -0.247 | -0.3915 | Yes |
| 79 | Ndufs8 | na | 52980 | -0.252 | -0.3811 | Yes |
| 80 | Ndufb11 | na | 53142 | -0.262 | -0.3721 | Yes |
| 81 | Cyc1 | na | 53209 | -0.266 | -0.3611 | Yes |
| 82 | Coa6 | na | 53239 | -0.268 | -0.3495 | Yes |
| 83 | Ndufv3 | na | 53251 | -0.269 | -0.3375 | Yes |
| 84 | mt-Nd3 | na | 53284 | -0.272 | -0.3257 | Yes |
| 85 | Ndufb5 | na | 53360 | -0.278 | -0.3144 | Yes |
| 86 | Cox6c | na | 53510 | -0.289 | -0.3039 | Yes |
| 87 | mt-Nd5 | na | 53975 | -0.326 | -0.2973 | Yes |
| 88 | Ndufb4 | na | 54053 | -0.333 | -0.2836 | Yes |
| 89 | Ndufa6 | na | 54058 | -0.333 | -0.2685 | Yes |
| 90 | Ndufb9 | na | 54149 | -0.340 | -0.2547 | Yes |
| 91 | mt-Nd1 | na | 54381 | -0.363 | -0.2423 | Yes |
| 92 | Ndufa10 | na | 54517 | -0.376 | -0.2276 | Yes |
| 93 | mt-Co1 | na | 54792 | -0.413 | -0.2137 | Yes |
| 94 | Ndufa5 | na | 55204 | -0.477 | -0.1994 | Yes |
| 95 | Ndufs7 | na | 55223 | -0.479 | -0.1779 | Yes |
| 96 | Cox8b | na | 55249 | -0.483 | -0.1564 | Yes |
| 97 | Ndufb10 | na | 55357 | -0.504 | -0.1354 | Yes |
| 98 | mt-Cytb | na | 55358 | -0.504 | -0.1125 | Yes |
| 99 | Sdhb | na | 55446 | -0.522 | -0.0903 | Yes |
| 100 | Ndufa4l2 | na | 55471 | -0.529 | -0.0667 | Yes |
| 101 | mt-Nd4 | na | 55568 | -0.554 | -0.0433 | Yes |
| 102 | Dmac2 | na | 55830 | -0.628 | -0.0194 | Yes |
| 103 | Wdr93 | na | 55860 | -0.640 | 0.0092 | Yes |