Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___PanKO_vs_WT.RPKM_matrix___PanKO_vs_WT.cls#PanKO_versus_WT |
| Phenotype | RPKM_matrix___PanKO_vs_WT.cls#PanKO_versus_WT |
| Upregulated in class | WT |
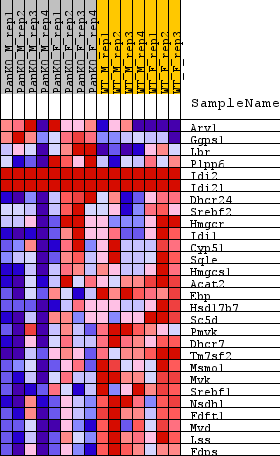
| GeneSet | REACTOME_CHOLESTEROL_BIOSYNTHESIS |
| Enrichment Score (ES) | -0.83174753 |
| Normalized Enrichment Score (NES) | -1.3091948 |
| Nominal p-value | 0.108208954 |
| FDR q-value | 0.5503765 |
| FWER p-Value | 0.895 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Arv1 | na | 2364 | 0.753 | 0.0033 | No |
| 2 | Ggps1 | na | 4085 | 0.567 | 0.0069 | No |
| 3 | Lbr | na | 4496 | 0.532 | 0.0316 | No |
| 4 | Plpp6 | na | 12114 | 0.126 | -0.0960 | No |
| 5 | Idi2 | na | 22007 | 0.000 | -0.2715 | No |
| 6 | Idi2l | na | 42016 | 0.000 | -0.6266 | No |
| 7 | Dhcr24 | na | 47133 | -0.000 | -0.7173 | No |
| 8 | Srebf2 | na | 47496 | -0.008 | -0.7233 | No |
| 9 | Hmgcr | na | 50495 | -0.112 | -0.7697 | No |
| 10 | Idi1 | na | 53991 | -0.327 | -0.8121 | Yes |
| 11 | Cyp51 | na | 54295 | -0.354 | -0.7962 | Yes |
| 12 | Sqle | na | 55143 | -0.467 | -0.7831 | Yes |
| 13 | Hmgcs1 | na | 55302 | -0.493 | -0.7563 | Yes |
| 14 | Acat2 | na | 55487 | -0.532 | -0.7276 | Yes |
| 15 | Ebp | na | 55541 | -0.545 | -0.6957 | Yes |
| 16 | Hsd17b7 | na | 55755 | -0.599 | -0.6635 | Yes |
| 17 | Sc5d | na | 55946 | -0.672 | -0.6265 | Yes |
| 18 | Pmvk | na | 56051 | -0.730 | -0.5845 | Yes |
| 19 | Dhcr7 | na | 56217 | -0.871 | -0.5351 | Yes |
| 20 | Tm7sf2 | na | 56226 | -0.890 | -0.4817 | Yes |
| 21 | Msmo1 | na | 56227 | -0.892 | -0.4281 | Yes |
| 22 | Mvk | na | 56249 | -0.934 | -0.3723 | Yes |
| 23 | Srebf1 | na | 56265 | -0.961 | -0.3147 | Yes |
| 24 | Nsdhl | na | 56288 | -1.016 | -0.2540 | Yes |
| 25 | Fdft1 | na | 56289 | -1.016 | -0.1929 | Yes |
| 26 | Mvd | na | 56292 | -1.024 | -0.1314 | Yes |
| 27 | Lss | na | 56306 | -1.055 | -0.0682 | Yes |
| 28 | Fdps | na | 56334 | -1.155 | 0.0008 | Yes |