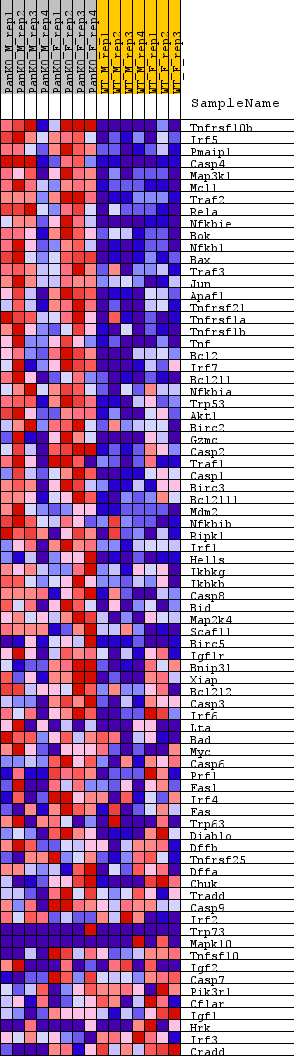
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___PanKO_vs_WT.RPKM_matrix___PanKO_vs_WT.cls#PanKO_versus_WT |
| Phenotype | RPKM_matrix___PanKO_vs_WT.cls#PanKO_versus_WT |
| Upregulated in class | PanKO |
| GeneSet | WP_APOPTOSIS |
| Enrichment Score (ES) | 0.74935335 |
| Normalized Enrichment Score (NES) | 1.3692318 |
| Nominal p-value | 0.021912351 |
| FDR q-value | 0.14340965 |
| FWER p-Value | 0.507 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Tnfrsf10b | na | 81 | 1.561 | 0.0345 | Yes |
| 2 | Irf5 | na | 169 | 1.447 | 0.0663 | Yes |
| 3 | Pmaip1 | na | 340 | 1.302 | 0.0933 | Yes |
| 4 | Casp4 | na | 466 | 1.233 | 0.1195 | Yes |
| 5 | Map3k1 | na | 499 | 1.220 | 0.1470 | Yes |
| 6 | Mcl1 | na | 726 | 1.127 | 0.1690 | Yes |
| 7 | Traf2 | na | 815 | 1.098 | 0.1927 | Yes |
| 8 | Rela | na | 840 | 1.090 | 0.2174 | Yes |
| 9 | Nfkbie | na | 891 | 1.075 | 0.2413 | Yes |
| 10 | Bok | na | 998 | 1.043 | 0.2634 | Yes |
| 11 | Nfkb1 | na | 1104 | 1.005 | 0.2847 | Yes |
| 12 | Bax | na | 1240 | 0.967 | 0.3046 | Yes |
| 13 | Traf3 | na | 1254 | 0.963 | 0.3265 | Yes |
| 14 | Jun | na | 1278 | 0.959 | 0.3482 | Yes |
| 15 | Apaf1 | na | 1341 | 0.945 | 0.3689 | Yes |
| 16 | Tnfrsf21 | na | 1509 | 0.908 | 0.3868 | Yes |
| 17 | Tnfrsf1a | na | 1583 | 0.893 | 0.4061 | Yes |
| 18 | Tnfrsf1b | na | 1715 | 0.862 | 0.4236 | Yes |
| 19 | Tnf | na | 1764 | 0.854 | 0.4425 | Yes |
| 20 | Bcl2 | na | 1788 | 0.850 | 0.4616 | Yes |
| 21 | Irf7 | na | 1827 | 0.843 | 0.4804 | Yes |
| 22 | Bcl2l1 | na | 2161 | 0.783 | 0.4925 | Yes |
| 23 | Nfkbia | na | 2180 | 0.779 | 0.5101 | Yes |
| 24 | Trp53 | na | 2262 | 0.769 | 0.5264 | Yes |
| 25 | Akt1 | na | 2391 | 0.749 | 0.5414 | Yes |
| 26 | Birc2 | na | 2857 | 0.692 | 0.5491 | Yes |
| 27 | Gzmc | na | 2958 | 0.679 | 0.5629 | Yes |
| 28 | Casp2 | na | 3037 | 0.672 | 0.5770 | Yes |
| 29 | Traf1 | na | 3360 | 0.637 | 0.5860 | Yes |
| 30 | Casp1 | na | 3401 | 0.631 | 0.5998 | Yes |
| 31 | Birc3 | na | 3524 | 0.617 | 0.6118 | Yes |
| 32 | Bcl2l11 | na | 3531 | 0.617 | 0.6259 | Yes |
| 33 | Mdm2 | na | 3696 | 0.603 | 0.6369 | Yes |
| 34 | Nfkbib | na | 3989 | 0.576 | 0.6450 | Yes |
| 35 | Ripk1 | na | 4046 | 0.570 | 0.6571 | Yes |
| 36 | Irf1 | na | 4551 | 0.528 | 0.6603 | Yes |
| 37 | Hells | na | 4579 | 0.526 | 0.6720 | Yes |
| 38 | Ikbkg | na | 4743 | 0.514 | 0.6809 | Yes |
| 39 | Ikbkb | na | 4887 | 0.502 | 0.6900 | Yes |
| 40 | Casp8 | na | 5050 | 0.491 | 0.6984 | Yes |
| 41 | Bid | na | 5144 | 0.484 | 0.7079 | Yes |
| 42 | Map2k4 | na | 5148 | 0.484 | 0.7190 | Yes |
| 43 | Scaf11 | na | 5398 | 0.467 | 0.7253 | Yes |
| 44 | Birc5 | na | 5467 | 0.462 | 0.7348 | Yes |
| 45 | Igf1r | na | 5859 | 0.436 | 0.7379 | Yes |
| 46 | Bnip3l | na | 6155 | 0.418 | 0.7423 | Yes |
| 47 | Xiap | na | 6550 | 0.391 | 0.7443 | Yes |
| 48 | Bcl2l2 | na | 7424 | 0.343 | 0.7367 | Yes |
| 49 | Casp3 | na | 8031 | 0.310 | 0.7330 | Yes |
| 50 | Irf6 | na | 8123 | 0.306 | 0.7385 | Yes |
| 51 | Lta | na | 8139 | 0.305 | 0.7452 | Yes |
| 52 | Bad | na | 8695 | 0.277 | 0.7417 | Yes |
| 53 | Myc | na | 8734 | 0.274 | 0.7474 | Yes |
| 54 | Casp6 | na | 9770 | 0.227 | 0.7342 | Yes |
| 55 | Prf1 | na | 9846 | 0.224 | 0.7381 | Yes |
| 56 | Fasl | na | 9980 | 0.218 | 0.7407 | Yes |
| 57 | Irf4 | na | 10148 | 0.211 | 0.7426 | Yes |
| 58 | Fas | na | 10328 | 0.202 | 0.7441 | Yes |
| 59 | Trp63 | na | 10503 | 0.194 | 0.7455 | Yes |
| 60 | Diablo | na | 10537 | 0.193 | 0.7494 | Yes |
| 61 | Dffb | na | 10837 | 0.180 | 0.7482 | No |
| 62 | Tnfrsf25 | na | 11340 | 0.157 | 0.7429 | No |
| 63 | Dffa | na | 11465 | 0.152 | 0.7442 | No |
| 64 | Chuk | na | 14191 | 0.059 | 0.6971 | No |
| 65 | Tradd | na | 14355 | 0.055 | 0.6955 | No |
| 66 | Casp9 | na | 15400 | 0.033 | 0.6777 | No |
| 67 | Irf2 | na | 15846 | 0.025 | 0.6704 | No |
| 68 | Trp73 | na | 18532 | 0.000 | 0.6227 | No |
| 69 | Mapk10 | na | 47153 | -0.001 | 0.1144 | No |
| 70 | Tnfsf10 | na | 47899 | -0.017 | 0.1016 | No |
| 71 | Igf2 | na | 48153 | -0.023 | 0.0976 | No |
| 72 | Casp7 | na | 51549 | -0.168 | 0.0412 | No |
| 73 | Pik3r1 | na | 52729 | -0.233 | 0.0256 | No |
| 74 | Cflar | na | 52859 | -0.242 | 0.0289 | No |
| 75 | Igf1 | na | 53121 | -0.261 | 0.0303 | No |
| 76 | Hrk | na | 53172 | -0.264 | 0.0355 | No |
| 77 | Irf3 | na | 53255 | -0.270 | 0.0402 | No |
| 78 | Cradd | na | 55918 | -0.661 | 0.0082 | No |