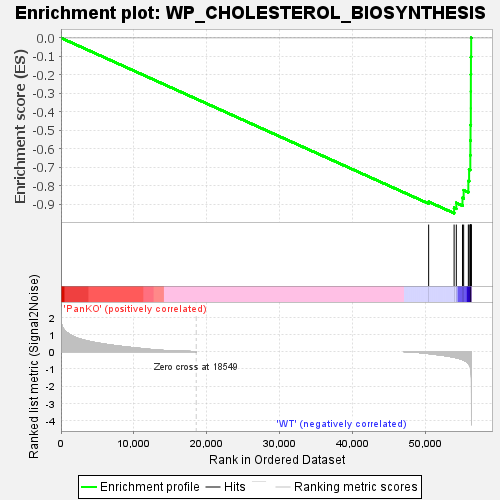
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___PanKO_vs_WT.RPKM_matrix___PanKO_vs_WT.cls#PanKO_versus_WT |
| Phenotype | RPKM_matrix___PanKO_vs_WT.cls#PanKO_versus_WT |
| Upregulated in class | WT |
| GeneSet | WP_CHOLESTEROL_BIOSYNTHESIS |
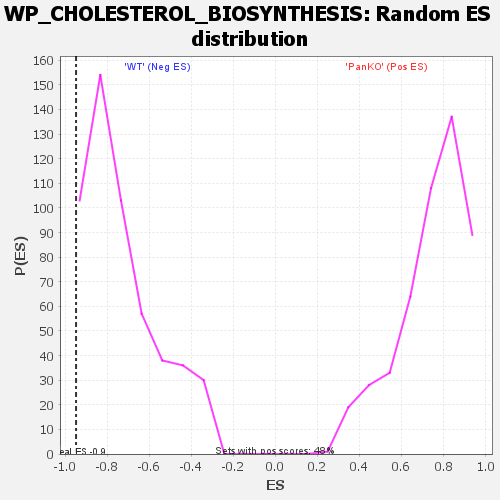
| Enrichment Score (ES) | -0.94779867 |
| Normalized Enrichment Score (NES) | -1.2864174 |
| Nominal p-value | 0.032629557 |
| FDR q-value | 0.5368464 |
| FWER p-Value | 0.775 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Hmgcr | na | 50495 | -0.112 | -0.8858 | No |
| 2 | Idi1 | na | 53991 | -0.327 | -0.9184 | Yes |
| 3 | Cyp51 | na | 54295 | -0.354 | -0.8919 | Yes |
| 4 | Sqle | na | 55143 | -0.467 | -0.8649 | Yes |
| 5 | Hmgcs1 | na | 55302 | -0.493 | -0.8234 | Yes |
| 6 | Sc5d | na | 55946 | -0.672 | -0.7744 | Yes |
| 7 | Pmvk | na | 56051 | -0.730 | -0.7106 | Yes |
| 8 | Dhcr7 | na | 56217 | -0.871 | -0.6352 | Yes |
| 9 | Msmo1 | na | 56227 | -0.892 | -0.5551 | Yes |
| 10 | Mvk | na | 56249 | -0.934 | -0.4714 | Yes |
| 11 | Nsdhl | na | 56288 | -1.016 | -0.3808 | Yes |
| 12 | Fdft1 | na | 56289 | -1.016 | -0.2894 | Yes |
| 13 | Mvd | na | 56292 | -1.024 | -0.1973 | Yes |
| 14 | Lss | na | 56306 | -1.055 | -0.1027 | Yes |
| 15 | Fdps | na | 56334 | -1.155 | 0.0008 | Yes |