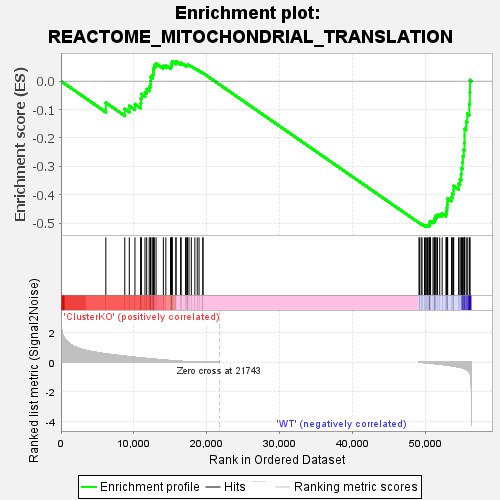
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___ClusterKO_vs_WT.RPKM_matrix___ClusterKO_vs_WT.cls #ClusterKO_versus_WT |
| Phenotype | RPKM_matrix___ClusterKO_vs_WT.cls#ClusterKO_versus_WT |
| Upregulated in class | WT |
| GeneSet | REACTOME_MITOCHONDRIAL_TRANSLATION |
| Enrichment Score (ES) | -0.512153 |
| Normalized Enrichment Score (NES) | -1.0388811 |
| Nominal p-value | 0.4502924 |
| FDR q-value | 0.8400259 |
| FWER p-Value | 1.0 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Mrpl3 | na | 6158 | 0.556 | -0.0759 | No |
| 2 | Mrpl50 | na | 8758 | 0.400 | -0.0979 | No |
| 3 | Dap3 | na | 9380 | 0.367 | -0.0869 | No |
| 4 | Mtrf1l | na | 10157 | 0.326 | -0.0810 | No |
| 5 | Mrpl9 | na | 10944 | 0.284 | -0.0779 | No |
| 6 | Mrpl19 | na | 10979 | 0.283 | -0.0614 | No |
| 7 | Mrpl53 | na | 11039 | 0.280 | -0.0456 | No |
| 8 | Mrpl44 | na | 11534 | 0.256 | -0.0389 | No |
| 9 | Mrps14 | na | 11767 | 0.244 | -0.0284 | No |
| 10 | Mrps6 | na | 12113 | 0.228 | -0.0208 | No |
| 11 | Mrpl35 | na | 12263 | 0.222 | -0.0101 | No |
| 12 | Oxa1l | na | 12318 | 0.220 | 0.0022 | No |
| 13 | Mrpl18 | na | 12340 | 0.219 | 0.0150 | No |
| 14 | Mrpl13 | na | 12607 | 0.205 | 0.0227 | No |
| 15 | Mrps23 | na | 12671 | 0.202 | 0.0337 | No |
| 16 | Mrpl36 | na | 12712 | 0.201 | 0.0451 | No |
| 17 | Mrps31 | na | 12819 | 0.196 | 0.0551 | No |
| 18 | Mrpl37 | na | 13055 | 0.186 | 0.0621 | No |
| 19 | Mrps17 | na | 14056 | 0.145 | 0.0531 | No |
| 20 | Mrpl28 | na | 14400 | 0.131 | 0.0549 | No |
| 21 | Mrpl43 | na | 15040 | 0.104 | 0.0498 | No |
| 22 | Mrpl27 | na | 15150 | 0.100 | 0.0539 | No |
| 23 | Mrps18c | na | 15177 | 0.099 | 0.0594 | No |
| 24 | Mrpl33 | na | 15234 | 0.098 | 0.0643 | No |
| 25 | Mrps18a | na | 15287 | 0.096 | 0.0692 | No |
| 26 | Mrpl17 | na | 15789 | 0.079 | 0.0651 | No |
| 27 | Eral1 | na | 15791 | 0.079 | 0.0698 | No |
| 28 | Mrpl10 | na | 16424 | 0.058 | 0.0621 | No |
| 29 | Ptcd3 | na | 16500 | 0.056 | 0.0642 | No |
| 30 | Mrpl32 | na | 17105 | 0.041 | 0.0559 | No |
| 31 | Mrpl47 | na | 17262 | 0.038 | 0.0554 | No |
| 32 | Mrpl12 | na | 17363 | 0.036 | 0.0558 | No |
| 33 | Mrpl14 | na | 17379 | 0.036 | 0.0577 | No |
| 34 | Mrpl40 | na | 17581 | 0.033 | 0.0561 | No |
| 35 | Mrps33 | na | 17900 | 0.028 | 0.0521 | No |
| 36 | Mrpl30 | na | 18385 | 0.021 | 0.0448 | No |
| 37 | Mrpl4 | na | 18713 | 0.018 | 0.0401 | No |
| 38 | Mrpl16 | na | 18959 | 0.016 | 0.0367 | No |
| 39 | Mrps18b | na | 19491 | 0.011 | 0.0279 | No |
| 40 | Mrpl57 | na | 19493 | 0.011 | 0.0286 | No |
| 41 | Mrpl45 | na | 49193 | -0.003 | -0.4989 | No |
| 42 | Mrpl20 | na | 49204 | -0.004 | -0.4988 | No |
| 43 | Mrpl48 | na | 49460 | -0.011 | -0.5027 | No |
| 44 | Mrpl46 | na | 49587 | -0.015 | -0.5041 | No |
| 45 | Mrpl11 | na | 49923 | -0.027 | -0.5084 | No |
| 46 | Mrps2 | na | 50048 | -0.032 | -0.5087 | No |
| 47 | Mrps12 | na | 50224 | -0.038 | -0.5095 | Yes |
| 48 | Mrps24 | na | 50334 | -0.042 | -0.5089 | Yes |
| 49 | Mrpl24 | na | 50519 | -0.050 | -0.5091 | Yes |
| 50 | Mrrf | na | 50591 | -0.054 | -0.5071 | Yes |
| 51 | Mrps30 | na | 50642 | -0.056 | -0.5046 | Yes |
| 52 | Mrps16 | na | 50658 | -0.057 | -0.5015 | Yes |
| 53 | Mrpl54 | na | 50662 | -0.057 | -0.4981 | Yes |
| 54 | Gfm2 | na | 50676 | -0.057 | -0.4949 | Yes |
| 55 | Mrps9 | na | 50757 | -0.062 | -0.4926 | Yes |
| 56 | Mrpl55 | na | 51094 | -0.078 | -0.4939 | Yes |
| 57 | Mrps7 | na | 51240 | -0.084 | -0.4914 | Yes |
| 58 | Mrpl1 | na | 51326 | -0.088 | -0.4876 | Yes |
| 59 | Chchd1 | na | 51327 | -0.088 | -0.4823 | Yes |
| 60 | Mrps35 | na | 51497 | -0.097 | -0.4794 | Yes |
| 61 | Mrpl58 | na | 51498 | -0.097 | -0.4736 | Yes |
| 62 | Aurkaip1 | na | 51713 | -0.108 | -0.4709 | Yes |
| 63 | Mrpl15 | na | 52017 | -0.126 | -0.4687 | Yes |
| 64 | Mrps26 | na | 52349 | -0.145 | -0.4659 | Yes |
| 65 | Mrpl2 | na | 52873 | -0.175 | -0.4646 | Yes |
| 66 | Mrps28 | na | 52952 | -0.180 | -0.4551 | Yes |
| 67 | Mrpl49 | na | 53004 | -0.184 | -0.4449 | Yes |
| 68 | Mrps11 | na | 53056 | -0.187 | -0.4346 | Yes |
| 69 | Mrps22 | na | 53093 | -0.189 | -0.4238 | Yes |
| 70 | Mrpl23 | na | 53101 | -0.189 | -0.4126 | Yes |
| 71 | Mrps25 | na | 53640 | -0.230 | -0.4082 | Yes |
| 72 | Mrpl21 | na | 53748 | -0.241 | -0.3956 | Yes |
| 73 | Gfm1 | na | 53886 | -0.249 | -0.3831 | Yes |
| 74 | Mrps34 | na | 53935 | -0.254 | -0.3686 | Yes |
| 75 | Mrpl39 | na | 54612 | -0.312 | -0.3618 | Yes |
| 76 | Mrps10 | na | 54792 | -0.329 | -0.3452 | Yes |
| 77 | Mrps15 | na | 54956 | -0.348 | -0.3271 | Yes |
| 78 | Mrpl22 | na | 54994 | -0.354 | -0.3064 | Yes |
| 79 | Mrpl38 | na | 55142 | -0.377 | -0.2863 | Yes |
| 80 | Mrpl41 | na | 55190 | -0.385 | -0.2640 | Yes |
| 81 | Mrpl34 | na | 55317 | -0.404 | -0.2419 | Yes |
| 82 | Mrpl51 | na | 55411 | -0.420 | -0.2182 | Yes |
| 83 | Mrps36 | na | 55422 | -0.422 | -0.1930 | Yes |
| 84 | Gadd45gip1 | na | 55423 | -0.422 | -0.1676 | Yes |
| 85 | Mrps5 | na | 55644 | -0.472 | -0.1431 | Yes |
| 86 | Mrpl52 | na | 55799 | -0.513 | -0.1149 | Yes |
| 87 | Mrpl42 | na | 56065 | -0.642 | -0.0809 | Yes |
| 88 | Mrps21 | na | 56148 | -0.704 | -0.0399 | Yes |
| 89 | Mrps27 | na | 56168 | -0.730 | 0.0037 | Yes |

