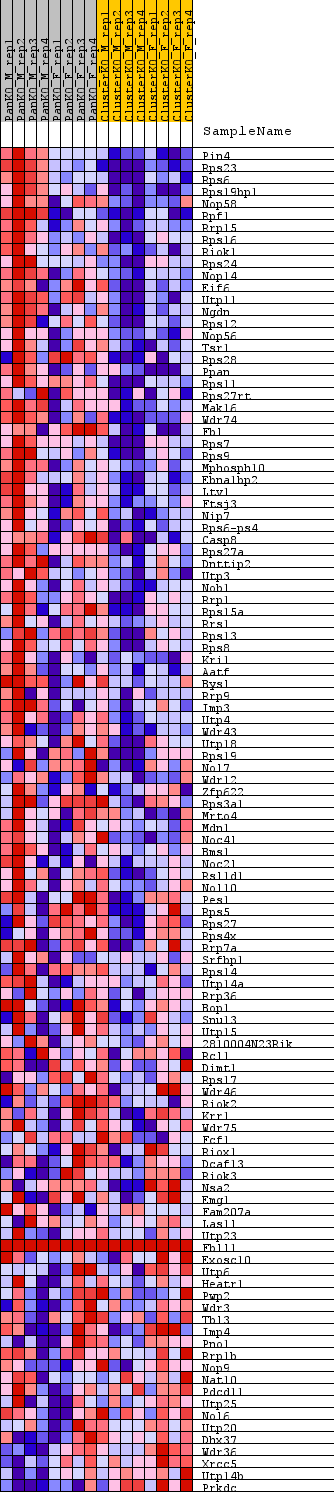
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___PanKO_vs_ClusterKO.RPKM_matrix___PanKO_vs_ClusterKO.cls #PanKO_versus_ClusterKO |
| Phenotype | RPKM_matrix___PanKO_vs_ClusterKO.cls#PanKO_versus_ClusterKO |
| Upregulated in class | PanKO |
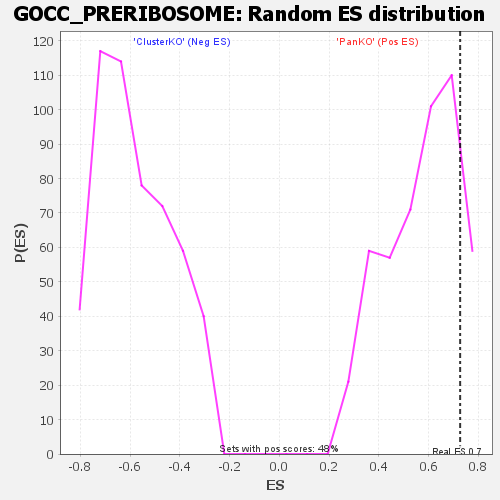
| GeneSet | GOCC_PRERIBOSOME |
| Enrichment Score (ES) | 0.7269975 |
| Normalized Enrichment Score (NES) | 1.2693855 |
| Nominal p-value | 0.14853556 |
| FDR q-value | 0.9489943 |
| FWER p-Value | 0.929 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Pin4 | na | 3 | 1.001 | 0.0330 | Yes |
| 2 | Rps23 | na | 34 | 0.849 | 0.0606 | Yes |
| 3 | Rps6 | na | 138 | 0.699 | 0.0818 | Yes |
| 4 | Rps19bp1 | na | 382 | 0.593 | 0.0971 | Yes |
| 5 | Nop58 | na | 420 | 0.580 | 0.1156 | Yes |
| 6 | Rpf1 | na | 564 | 0.546 | 0.1311 | Yes |
| 7 | Rrp15 | na | 578 | 0.542 | 0.1488 | Yes |
| 8 | Rps16 | na | 640 | 0.530 | 0.1652 | Yes |
| 9 | Riok1 | na | 657 | 0.528 | 0.1824 | Yes |
| 10 | Rps24 | na | 672 | 0.524 | 0.1995 | Yes |
| 11 | Nop14 | na | 690 | 0.520 | 0.2163 | Yes |
| 12 | Eif6 | na | 720 | 0.515 | 0.2328 | Yes |
| 13 | Utp11 | na | 732 | 0.514 | 0.2496 | Yes |
| 14 | Ngdn | na | 740 | 0.513 | 0.2664 | Yes |
| 15 | Rps12 | na | 825 | 0.498 | 0.2814 | Yes |
| 16 | Nop56 | na | 827 | 0.497 | 0.2978 | Yes |
| 17 | Tsr1 | na | 830 | 0.497 | 0.3142 | Yes |
| 18 | Rps28 | na | 952 | 0.474 | 0.3277 | Yes |
| 19 | Ppan | na | 977 | 0.470 | 0.3428 | Yes |
| 20 | Rps11 | na | 1039 | 0.462 | 0.3570 | Yes |
| 21 | Rps27rt | na | 1062 | 0.458 | 0.3717 | Yes |
| 22 | Mak16 | na | 1065 | 0.457 | 0.3868 | Yes |
| 23 | Wdr74 | na | 1158 | 0.447 | 0.3999 | Yes |
| 24 | Fbl | na | 1160 | 0.447 | 0.4147 | Yes |
| 25 | Rps7 | na | 1176 | 0.445 | 0.4291 | Yes |
| 26 | Rps9 | na | 1229 | 0.439 | 0.4427 | Yes |
| 27 | Mphosph10 | na | 1293 | 0.432 | 0.4559 | Yes |
| 28 | Ebna1bp2 | na | 1496 | 0.411 | 0.4659 | Yes |
| 29 | Ltv1 | na | 1698 | 0.392 | 0.4753 | Yes |
| 30 | Ftsj3 | na | 1715 | 0.389 | 0.4879 | Yes |
| 31 | Nip7 | na | 1746 | 0.387 | 0.5001 | Yes |
| 32 | Rps6-ps4 | na | 1823 | 0.382 | 0.5114 | Yes |
| 33 | Casp8 | na | 1837 | 0.380 | 0.5237 | Yes |
| 34 | Rps27a | na | 1885 | 0.375 | 0.5352 | Yes |
| 35 | Dnttip2 | na | 1901 | 0.374 | 0.5473 | Yes |
| 36 | Utp3 | na | 1983 | 0.367 | 0.5580 | Yes |
| 37 | Nob1 | na | 2347 | 0.340 | 0.5628 | Yes |
| 38 | Rrp1 | na | 2484 | 0.332 | 0.5714 | Yes |
| 39 | Rps15a | na | 2661 | 0.320 | 0.5788 | Yes |
| 40 | Rrs1 | na | 2741 | 0.315 | 0.5878 | Yes |
| 41 | Rps13 | na | 2881 | 0.306 | 0.5955 | Yes |
| 42 | Rps8 | na | 2943 | 0.302 | 0.6044 | Yes |
| 43 | Kri1 | na | 3072 | 0.294 | 0.6118 | Yes |
| 44 | Aatf | na | 3159 | 0.288 | 0.6198 | Yes |
| 45 | Bysl | na | 3172 | 0.288 | 0.6291 | Yes |
| 46 | Rrp9 | na | 3486 | 0.273 | 0.6326 | Yes |
| 47 | Imp3 | na | 3521 | 0.272 | 0.6410 | Yes |
| 48 | Utp4 | na | 3537 | 0.271 | 0.6497 | Yes |
| 49 | Wdr43 | na | 4227 | 0.236 | 0.6452 | Yes |
| 50 | Utp18 | na | 4241 | 0.235 | 0.6527 | Yes |
| 51 | Rps19 | na | 4386 | 0.228 | 0.6577 | Yes |
| 52 | Nol7 | na | 4563 | 0.220 | 0.6618 | Yes |
| 53 | Wdr12 | na | 4850 | 0.208 | 0.6636 | Yes |
| 54 | Zfp622 | na | 4913 | 0.206 | 0.6693 | Yes |
| 55 | Rps3a1 | na | 4915 | 0.206 | 0.6761 | Yes |
| 56 | Mrto4 | na | 5001 | 0.201 | 0.6813 | Yes |
| 57 | Mdn1 | na | 5282 | 0.191 | 0.6826 | Yes |
| 58 | Noc4l | na | 5301 | 0.190 | 0.6886 | Yes |
| 59 | Bms1 | na | 5905 | 0.168 | 0.6834 | Yes |
| 60 | Noc2l | na | 5994 | 0.165 | 0.6873 | Yes |
| 61 | Rsl1d1 | na | 6166 | 0.159 | 0.6895 | Yes |
| 62 | Nol10 | na | 6203 | 0.157 | 0.6940 | Yes |
| 63 | Pes1 | na | 6256 | 0.155 | 0.6982 | Yes |
| 64 | Rps5 | na | 6310 | 0.154 | 0.7024 | Yes |
| 65 | Rps27 | na | 6419 | 0.149 | 0.7054 | Yes |
| 66 | Rps4x | na | 6432 | 0.149 | 0.7101 | Yes |
| 67 | Rrp7a | na | 6478 | 0.147 | 0.7141 | Yes |
| 68 | Srfbp1 | na | 6720 | 0.138 | 0.7144 | Yes |
| 69 | Rps14 | na | 6738 | 0.137 | 0.7186 | Yes |
| 70 | Utp14a | na | 7234 | 0.121 | 0.7138 | Yes |
| 71 | Rrp36 | na | 7237 | 0.121 | 0.7178 | Yes |
| 72 | Bop1 | na | 7394 | 0.116 | 0.7189 | Yes |
| 73 | Snu13 | na | 7418 | 0.115 | 0.7222 | Yes |
| 74 | Utp15 | na | 7554 | 0.110 | 0.7235 | Yes |
| 75 | 2810004N23Rik | na | 7563 | 0.110 | 0.7270 | Yes |
| 76 | Rcl1 | na | 8049 | 0.093 | 0.7214 | No |
| 77 | Dimt1 | na | 8295 | 0.084 | 0.7199 | No |
| 78 | Rps17 | na | 8382 | 0.081 | 0.7210 | No |
| 79 | Wdr46 | na | 9012 | 0.064 | 0.7120 | No |
| 80 | Riok2 | na | 9175 | 0.059 | 0.7110 | No |
| 81 | Krr1 | na | 9337 | 0.054 | 0.7100 | No |
| 82 | Wdr75 | na | 9408 | 0.052 | 0.7105 | No |
| 83 | Fcf1 | na | 10161 | 0.032 | 0.6982 | No |
| 84 | Riox1 | na | 10205 | 0.031 | 0.6984 | No |
| 85 | Dcaf13 | na | 10435 | 0.025 | 0.6952 | No |
| 86 | Riok3 | na | 10958 | 0.014 | 0.6864 | No |
| 87 | Nsa2 | na | 10977 | 0.014 | 0.6865 | No |
| 88 | Emg1 | na | 11116 | 0.011 | 0.6844 | No |
| 89 | Fam207a | na | 11444 | 0.007 | 0.6788 | No |
| 90 | Las1l | na | 11546 | 0.006 | 0.6772 | No |
| 91 | Utp23 | na | 11834 | 0.003 | 0.6722 | No |
| 92 | Fbll1 | na | 31934 | 0.000 | 0.3150 | No |
| 93 | Exosc10 | na | 39289 | -0.003 | 0.1844 | No |
| 94 | Utp6 | na | 41052 | -0.017 | 0.1537 | No |
| 95 | Heatr1 | na | 41632 | -0.026 | 0.1442 | No |
| 96 | Pwp2 | na | 41664 | -0.026 | 0.1446 | No |
| 97 | Wdr3 | na | 41719 | -0.027 | 0.1445 | No |
| 98 | Tbl3 | na | 42033 | -0.032 | 0.1400 | No |
| 99 | Imp4 | na | 43180 | -0.058 | 0.1215 | No |
| 100 | Pno1 | na | 43521 | -0.067 | 0.1177 | No |
| 101 | Rrp1b | na | 44557 | -0.097 | 0.1025 | No |
| 102 | Nop9 | na | 44715 | -0.103 | 0.1031 | No |
| 103 | Nat10 | na | 45149 | -0.115 | 0.0992 | No |
| 104 | Pdcd11 | na | 45595 | -0.128 | 0.0956 | No |
| 105 | Utp25 | na | 45607 | -0.128 | 0.0996 | No |
| 106 | Nol6 | na | 46462 | -0.156 | 0.0896 | No |
| 107 | Utp20 | na | 47958 | -0.205 | 0.0698 | No |
| 108 | Dhx37 | na | 49362 | -0.257 | 0.0533 | No |
| 109 | Wdr36 | na | 51141 | -0.332 | 0.0327 | No |
| 110 | Xrcc5 | na | 52309 | -0.388 | 0.0248 | No |
| 111 | Utp14b | na | 55010 | -0.598 | -0.0034 | No |
| 112 | Prkdc | na | 56112 | -0.839 | 0.0047 | No |