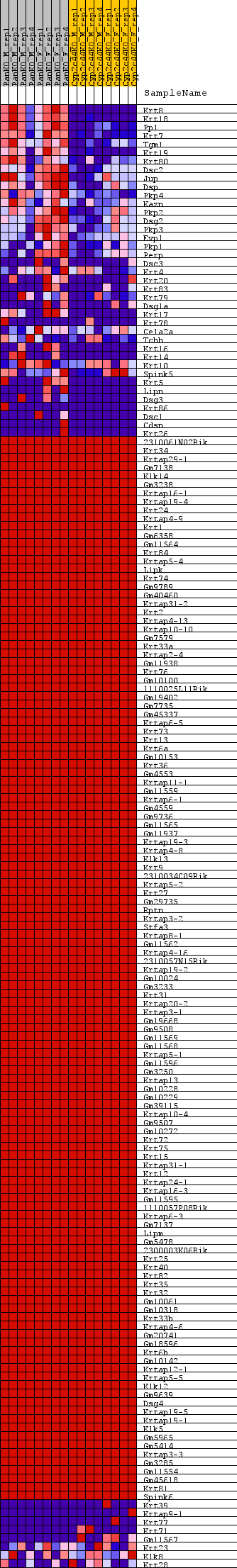
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | RPKM_matrix___PanKO_vs_Cyp2c44KO.RPKM_matrix___PanKO_vs_Cyp2c44KO.cls #PanKO_versus_Cyp2c44KO |
| Phenotype | RPKM_matrix___PanKO_vs_Cyp2c44KO.cls#PanKO_versus_Cyp2c44KO |
| Upregulated in class | PanKO |
| GeneSet | REACTOME_KERATINIZATION |
| Enrichment Score (ES) | 0.7617146 |
| Normalized Enrichment Score (NES) | 1.4952452 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.37834153 |
| FWER p-Value | 0.316 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Krt8 | na | 5 | 1.960 | 0.1021 | Yes |
| 2 | Krt18 | na | 18 | 1.819 | 0.1968 | Yes |
| 3 | Ppl | na | 71 | 1.609 | 0.2797 | Yes |
| 4 | Krt7 | na | 380 | 1.267 | 0.3403 | Yes |
| 5 | Tgm1 | na | 539 | 1.176 | 0.3988 | Yes |
| 6 | Krt19 | na | 772 | 1.085 | 0.4513 | Yes |
| 7 | Krt80 | na | 2060 | 0.762 | 0.4681 | Yes |
| 8 | Dsc2 | na | 2415 | 0.714 | 0.4990 | Yes |
| 9 | Jup | na | 2482 | 0.704 | 0.5346 | Yes |
| 10 | Dsp | na | 2537 | 0.697 | 0.5700 | Yes |
| 11 | Pkp4 | na | 2985 | 0.635 | 0.5951 | Yes |
| 12 | Kazn | na | 3269 | 0.602 | 0.6215 | Yes |
| 13 | Pkp2 | na | 3368 | 0.591 | 0.6506 | Yes |
| 14 | Dsg2 | na | 3413 | 0.586 | 0.6803 | Yes |
| 15 | Pkp3 | na | 4203 | 0.512 | 0.6930 | Yes |
| 16 | Evpl | na | 4540 | 0.487 | 0.7124 | Yes |
| 17 | Pkp1 | na | 5437 | 0.421 | 0.7184 | Yes |
| 18 | Perp | na | 5816 | 0.397 | 0.7324 | Yes |
| 19 | Dsc3 | na | 5834 | 0.396 | 0.7527 | Yes |
| 20 | Krt4 | na | 7588 | 0.297 | 0.7370 | Yes |
| 21 | Krt20 | na | 7590 | 0.297 | 0.7525 | Yes |
| 22 | Krt83 | na | 7898 | 0.281 | 0.7617 | Yes |
| 23 | Krt79 | na | 8875 | 0.237 | 0.7567 | No |
| 24 | Dsg1a | na | 9908 | 0.194 | 0.7484 | No |
| 25 | Krt17 | na | 10140 | 0.183 | 0.7539 | No |
| 26 | Krt78 | na | 10699 | 0.157 | 0.7521 | No |
| 27 | Cela2a | na | 10748 | 0.154 | 0.7593 | No |
| 28 | Tchh | na | 11940 | 0.103 | 0.7435 | No |
| 29 | Krt16 | na | 13075 | 0.063 | 0.7266 | No |
| 30 | Krt14 | na | 13088 | 0.063 | 0.7297 | No |
| 31 | Krt10 | na | 13179 | 0.060 | 0.7312 | No |
| 32 | Spink5 | na | 13391 | 0.053 | 0.7302 | No |
| 33 | Krt5 | na | 13509 | 0.049 | 0.7307 | No |
| 34 | Lipn | na | 13751 | 0.041 | 0.7286 | No |
| 35 | Dsg3 | na | 14933 | 0.012 | 0.7082 | No |
| 36 | Krt86 | na | 14969 | 0.011 | 0.7081 | No |
| 37 | Dsc1 | na | 15521 | 0.004 | 0.6986 | No |
| 38 | Cdsn | na | 15595 | 0.003 | 0.6974 | No |
| 39 | Krt26 | na | 15766 | 0.002 | 0.6945 | No |
| 40 | 2310061N02Rik | na | 15999 | 0.000 | 0.6904 | No |
| 41 | Krt34 | na | 16344 | 0.000 | 0.6843 | No |
| 42 | Krtap29-1 | na | 16780 | 0.000 | 0.6765 | No |
| 43 | Gm7138 | na | 16828 | 0.000 | 0.6757 | No |
| 44 | Klk14 | na | 17022 | 0.000 | 0.6722 | No |
| 45 | Gm3238 | na | 17259 | 0.000 | 0.6680 | No |
| 46 | Krtap16-1 | na | 17284 | 0.000 | 0.6676 | No |
| 47 | Krtap19-4 | na | 17658 | 0.000 | 0.6610 | No |
| 48 | Krt24 | na | 17873 | 0.000 | 0.6572 | No |
| 49 | Krtap4-9 | na | 18055 | 0.000 | 0.6540 | No |
| 50 | Krt1 | na | 18145 | 0.000 | 0.6524 | No |
| 51 | Gm6358 | na | 18434 | 0.000 | 0.6472 | No |
| 52 | Gm11564 | na | 18459 | 0.000 | 0.6468 | No |
| 53 | Krt84 | na | 18778 | 0.000 | 0.6412 | No |
| 54 | Krtap5-4 | na | 19180 | 0.000 | 0.6340 | No |
| 55 | Lipk | na | 19447 | 0.000 | 0.6293 | No |
| 56 | Krt74 | na | 19458 | 0.000 | 0.6291 | No |
| 57 | Gm9789 | na | 19535 | 0.000 | 0.6278 | No |
| 58 | Gm40460 | na | 19992 | 0.000 | 0.6197 | No |
| 59 | Krtap31-2 | na | 20241 | 0.000 | 0.6152 | No |
| 60 | Krt2 | na | 20314 | 0.000 | 0.6140 | No |
| 61 | Krtap4-13 | na | 20689 | 0.000 | 0.6073 | No |
| 62 | Krtap10-10 | na | 20781 | 0.000 | 0.6057 | No |
| 63 | Gm7579 | na | 20842 | 0.000 | 0.6046 | No |
| 64 | Krt33a | na | 20850 | 0.000 | 0.6045 | No |
| 65 | Krtap2-4 | na | 21303 | 0.000 | 0.5965 | No |
| 66 | Gm11938 | na | 21467 | 0.000 | 0.5936 | No |
| 67 | Krt76 | na | 22428 | 0.000 | 0.5765 | No |
| 68 | Gm10100 | na | 22691 | 0.000 | 0.5718 | No |
| 69 | 1110025L11Rik | na | 22808 | 0.000 | 0.5697 | No |
| 70 | Gm19402 | na | 23464 | 0.000 | 0.5581 | No |
| 71 | Gm7735 | na | 23887 | 0.000 | 0.5506 | No |
| 72 | Gm45337 | na | 24007 | 0.000 | 0.5485 | No |
| 73 | Krtap6-5 | na | 24100 | 0.000 | 0.5468 | No |
| 74 | Krt73 | na | 24409 | 0.000 | 0.5414 | No |
| 75 | Krt13 | na | 24680 | 0.000 | 0.5365 | No |
| 76 | Krt6a | na | 24689 | 0.000 | 0.5364 | No |
| 77 | Gm10153 | na | 25047 | 0.000 | 0.5301 | No |
| 78 | Krt36 | na | 25588 | 0.000 | 0.5204 | No |
| 79 | Gm4553 | na | 26174 | 0.000 | 0.5100 | No |
| 80 | Krtap11-1 | na | 26427 | 0.000 | 0.5056 | No |
| 81 | Gm11559 | na | 26514 | 0.000 | 0.5040 | No |
| 82 | Krtap6-1 | na | 26607 | 0.000 | 0.5024 | No |
| 83 | Gm4559 | na | 26667 | 0.000 | 0.5013 | No |
| 84 | Gm9736 | na | 26672 | 0.000 | 0.5013 | No |
| 85 | Gm11565 | na | 27438 | 0.000 | 0.4877 | No |
| 86 | Gm11937 | na | 27905 | 0.000 | 0.4794 | No |
| 87 | Krtap19-3 | na | 28052 | 0.000 | 0.4768 | No |
| 88 | Krtap4-8 | na | 28085 | 0.000 | 0.4762 | No |
| 89 | Klk13 | na | 28386 | 0.000 | 0.4709 | No |
| 90 | Krt9 | na | 29066 | 0.000 | 0.4588 | No |
| 91 | 2310034C09Rik | na | 29218 | 0.000 | 0.4561 | No |
| 92 | Krtap5-2 | na | 29483 | 0.000 | 0.4514 | No |
| 93 | Krt27 | na | 29591 | 0.000 | 0.4495 | No |
| 94 | Gm29735 | na | 29724 | 0.000 | 0.4471 | No |
| 95 | Rptn | na | 29740 | 0.000 | 0.4469 | No |
| 96 | Krtap3-2 | na | 29945 | 0.000 | 0.4432 | No |
| 97 | Stfa3 | na | 30112 | 0.000 | 0.4403 | No |
| 98 | Krtap8-1 | na | 30389 | 0.000 | 0.4354 | No |
| 99 | Gm11562 | na | 30687 | 0.000 | 0.4301 | No |
| 100 | Krtap4-16 | na | 31056 | 0.000 | 0.4236 | No |
| 101 | 2310057N15Rik | na | 31246 | 0.000 | 0.4202 | No |
| 102 | Krtap19-2 | na | 31825 | 0.000 | 0.4099 | No |
| 103 | Gm10024 | na | 31826 | 0.000 | 0.4099 | No |
| 104 | Gm3233 | na | 31867 | 0.000 | 0.4092 | No |
| 105 | Krt31 | na | 32027 | 0.000 | 0.4064 | No |
| 106 | Krtap20-2 | na | 32357 | 0.000 | 0.4005 | No |
| 107 | Krtap3-1 | na | 32508 | 0.000 | 0.3978 | No |
| 108 | Gm19668 | na | 32571 | 0.000 | 0.3967 | No |
| 109 | Gm9508 | na | 32774 | 0.000 | 0.3931 | No |
| 110 | Gm11569 | na | 32808 | 0.000 | 0.3926 | No |
| 111 | Gm11568 | na | 33000 | 0.000 | 0.3892 | No |
| 112 | Krtap5-1 | na | 33172 | 0.000 | 0.3861 | No |
| 113 | Gm11596 | na | 33900 | 0.000 | 0.3732 | No |
| 114 | Gm3250 | na | 33906 | 0.000 | 0.3731 | No |
| 115 | Krtap13 | na | 34113 | 0.000 | 0.3694 | No |
| 116 | Gm10228 | na | 34115 | 0.000 | 0.3694 | No |
| 117 | Gm10229 | na | 34228 | 0.000 | 0.3674 | No |
| 118 | Gm39115 | na | 34374 | 0.000 | 0.3648 | No |
| 119 | Krtap10-4 | na | 34506 | 0.000 | 0.3625 | No |
| 120 | Gm9507 | na | 34560 | 0.000 | 0.3616 | No |
| 121 | Gm10272 | na | 34616 | 0.000 | 0.3606 | No |
| 122 | Krt72 | na | 34776 | 0.000 | 0.3578 | No |
| 123 | Krt75 | na | 34853 | 0.000 | 0.3564 | No |
| 124 | Krt15 | na | 34975 | 0.000 | 0.3543 | No |
| 125 | Krtap31-1 | na | 35074 | 0.000 | 0.3525 | No |
| 126 | Krt12 | na | 35130 | 0.000 | 0.3515 | No |
| 127 | Krtap24-1 | na | 35282 | 0.000 | 0.3488 | No |
| 128 | Krtap16-3 | na | 35353 | 0.000 | 0.3476 | No |
| 129 | Gm11595 | na | 35384 | 0.000 | 0.3471 | No |
| 130 | 1110057P08Rik | na | 35533 | 0.000 | 0.3444 | No |
| 131 | Krtap6-3 | na | 35794 | 0.000 | 0.3398 | No |
| 132 | Gm7137 | na | 35996 | 0.000 | 0.3362 | No |
| 133 | Lipm | na | 36100 | 0.000 | 0.3344 | No |
| 134 | Gm5478 | na | 36145 | 0.000 | 0.3336 | No |
| 135 | 2300003K06Rik | na | 36597 | 0.000 | 0.3256 | No |
| 136 | Krt25 | na | 36666 | 0.000 | 0.3244 | No |
| 137 | Krt40 | na | 36914 | 0.000 | 0.3200 | No |
| 138 | Krt82 | na | 37684 | 0.000 | 0.3063 | No |
| 139 | Krt35 | na | 37712 | 0.000 | 0.3058 | No |
| 140 | Krt32 | na | 37794 | 0.000 | 0.3044 | No |
| 141 | Gm10061 | na | 37965 | 0.000 | 0.3014 | No |
| 142 | Gm10318 | na | 38223 | 0.000 | 0.2968 | No |
| 143 | Krt33b | na | 38268 | 0.000 | 0.2960 | No |
| 144 | Krtap4-6 | na | 38367 | 0.000 | 0.2943 | No |
| 145 | Gm20741 | na | 38534 | 0.000 | 0.2913 | No |
| 146 | Gm18596 | na | 38580 | 0.000 | 0.2905 | No |
| 147 | Krt6b | na | 38742 | 0.000 | 0.2876 | No |
| 148 | Gm10142 | na | 38953 | 0.000 | 0.2839 | No |
| 149 | Krtap12-1 | na | 38999 | 0.000 | 0.2831 | No |
| 150 | Krtap5-5 | na | 39092 | 0.000 | 0.2815 | No |
| 151 | Klk12 | na | 39318 | 0.000 | 0.2775 | No |
| 152 | Gm9639 | na | 39651 | 0.000 | 0.2716 | No |
| 153 | Dsg4 | na | 39753 | 0.000 | 0.2698 | No |
| 154 | Krtap19-5 | na | 40335 | 0.000 | 0.2594 | No |
| 155 | Krtap19-1 | na | 40622 | 0.000 | 0.2543 | No |
| 156 | Klk5 | na | 41039 | 0.000 | 0.2469 | No |
| 157 | Gm5965 | na | 41119 | 0.000 | 0.2455 | No |
| 158 | Gm5414 | na | 41221 | 0.000 | 0.2437 | No |
| 159 | Krtap3-3 | na | 41255 | 0.000 | 0.2431 | No |
| 160 | Gm3285 | na | 41957 | 0.000 | 0.2307 | No |
| 161 | Gm11554 | na | 42521 | 0.000 | 0.2207 | No |
| 162 | Gm45618 | na | 42524 | 0.000 | 0.2206 | No |
| 163 | Krt81 | na | 42550 | 0.000 | 0.2202 | No |
| 164 | Spink6 | na | 42673 | 0.000 | 0.2180 | No |
| 165 | Krt39 | na | 43076 | -0.002 | 0.2110 | No |
| 166 | Krtap9-1 | na | 43176 | -0.003 | 0.2093 | No |
| 167 | Krt77 | na | 44389 | -0.012 | 0.1884 | No |
| 168 | Krt71 | na | 44839 | -0.018 | 0.1813 | No |
| 169 | Gm11567 | na | 45606 | -0.032 | 0.1694 | No |
| 170 | Krt23 | na | 47534 | -0.097 | 0.1401 | No |
| 171 | Klk8 | na | 47723 | -0.105 | 0.1423 | No |
| 172 | Krt28 | na | 50198 | -0.224 | 0.1099 | No |