| Rank | Motif | Name | P-value | log P-pvalue | q-value (Benjamini) | # Target Sequences with Motif | % of Targets Sequences with Motif | # Background Sequences with Motif | % of Background Sequences with Motif | Motif File |
PDF |
| 1 |  | Tcfcp2l1(CP2)/mES-Tcfcp2l1-ChIP-Seq/Homer | 1e-6 | -1.554e+01 | 0.0000 | 395.0 | 32.94% | 12705.4 | 26.28% | motif file (matrix) |
pdf |
| 2 |  | Stat3(Stat)/mES-Stat3-ChIP-Seq/Homer | 1e-5 | -1.377e+01 | 0.0001 | 904.0 | 75.40% | 33431.3 | 69.15% | motif file (matrix) |
pdf |
| 3 |  | STAT6/Macrophage-Stat6-ChIP-Seq/Homer | 1e-4 | -1.068e+01 | 0.0018 | 899.0 | 74.98% | 33656.1 | 69.62% | motif file (matrix) |
pdf |
| 4 |  | CTCF(Zf)/CD4+-CTCF-ChIP-Seq/Homer | 1e-4 | -1.010e+01 | 0.0024 | 334.0 | 27.86% | 11086.9 | 22.93% | motif file (matrix) |
pdf |
| 5 |  | HOXD13(Homeobox)/Chicken-Hoxd13-ChIP-Seq(GSE38910)/Homer | 1e-4 | -9.908e+00 | 0.0024 | 1103.0 | 91.99% | 42799.9 | 88.53% | motif file (matrix) |
pdf |
| 6 |  | Elk4(ETS)/Hela-Elk4-ChIP-Seq(GSE31477)/Homer | 1e-4 | -9.884e+00 | 0.0024 | 862.0 | 71.89% | 32212.9 | 66.63% | motif file (matrix) |
pdf |
| 7 |  | STAT5(Stat)/mCD4+-Stat5a|b-ChIP-Seq/Homer | 1e-4 | -9.797e+00 | 0.0024 | 716.0 | 59.72% | 26171.7 | 54.14% | motif file (matrix) |
pdf |
| 8 |  | NRF1/Promoter/Homer | 1e-4 | -9.581e+00 | 0.0024 | 304.0 | 25.35% | 10029.7 | 20.75% | motif file (matrix) |
pdf |
| 9 |  | PRDM9(Zf)/Testis-DMC1-ChIP-Seq(GSE35498)/Homer | 1e-3 | -9.138e+00 | 0.0027 | 834.0 | 69.56% | 31161.8 | 64.46% | motif file (matrix) |
pdf |
| 10 |  | Myf5(bHLH)/GM-Myf5-ChIP-Seq(GSE24852)/Homer | 1e-3 | -8.892e+00 | 0.0031 | 1010.0 | 84.24% | 38728.4 | 80.11% | motif file (matrix) |
pdf |
| 11 |  | Elk1(ETS)/Hela-Elk1-ChIP-Seq(GSE31477)/Homer | 1e-3 | -8.423e+00 | 0.0046 | 858.0 | 71.56% | 32291.6 | 66.79% | motif file (matrix) |
pdf |
| 12 |  | STAT4(Stat)/CD4-Stat4-ChIP-Seq/Homer | 1e-3 | -7.911e+00 | 0.0070 | 1116.0 | 93.08% | 43639.3 | 90.27% | motif file (matrix) |
pdf |
| 13 |  | STAT6(Stat)/CD4-Stat6-ChIP-Seq/Homer | 1e-3 | -7.589e+00 | 0.0089 | 893.0 | 74.48% | 33914.7 | 70.15% | motif file (matrix) |
pdf |
| 14 |  | CEBP:AP1(bZIP)/ThioMac-CEBPb-ChIP-Seq(GSE21512)/Homer | 1e-3 | -7.534e+00 | 0.0089 | 1019.0 | 84.99% | 39330.2 | 81.35% | motif file (matrix) |
pdf |
| 15 | 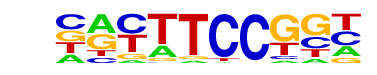 | Fli1(ETS)/CD8-FLI-ChIP-Seq(GSE20898)/Homer | 1e-3 | -7.377e+00 | 0.0095 | 1134.0 | 94.58% | 44547.9 | 92.15% | motif file (matrix) |
pdf |
| 16 |  | HIF-1a(HLH)/MCF7-HIF1a-ChIP-Seq/Homer | 1e-3 | -7.279e+00 | 0.0099 | 597.0 | 49.79% | 21824.7 | 45.14% | motif file (matrix) |
pdf |
| 17 |  | Bcl6(Zf)/Liver-Bcl6-ChIP-Seq(GSE31578)/Homer | 1e-3 | -7.272e+00 | 0.0099 | 1183.0 | 98.67% | 47011.3 | 97.24% | motif file (matrix) |
pdf |
| 18 |  | HIF2a(HLH)/O785-HIF2a-ChIP-Seq(GSE34871)/Homer | 1e-2 | -6.649e+00 | 0.0165 | 705.0 | 58.80% | 26317.3 | 54.44% | motif file (matrix) |
pdf |
| 19 |  | Foxa2(Forkhead)/Liver-Foxa2-ChIP-Seq/Homer | 1e-2 | -6.565e+00 | 0.0170 | 1032.0 | 86.07% | 40055.8 | 82.85% | motif file (matrix) |
pdf |
| 20 |  | CRX(Homeobox)/Retina-Crx-ChIP-Seq/Homer | 1e-2 | -6.503e+00 | 0.0172 | 1198.0 | 99.92% | 47991.6 | 99.27% | motif file (matrix) |
pdf |
| 21 |  | RUNX1(Runt)/Jurkat-RUNX1-ChIP-Seq/Homer | 1e-2 | -6.305e+00 | 0.0199 | 1141.0 | 95.16% | 45001.4 | 93.08% | motif file (matrix) |
pdf |
| 22 |  | EWS:FLI1-fusion(ETS)/SK_N_MC-EWS:FLI1-ChIP-Seq/Homer | 1e-2 | -6.242e+00 | 0.0202 | 921.0 | 76.81% | 35352.3 | 73.13% | motif file (matrix) |
pdf |
| 23 |  | Foxo1(Forkhead)/RAW-Foxo1-ChIP-Seq/Homer | 1e-2 | -6.212e+00 | 0.0202 | 1198.0 | 99.92% | 48004.4 | 99.30% | motif file (matrix) |
pdf |
| 24 |  | Oct4:Sox17/F9-Sox17-ChIP-Seq(GSE44553)/Homer | 1e-2 | -6.093e+00 | 0.0216 | 397.0 | 33.11% | 14164.7 | 29.30% | motif file (matrix) |
pdf |
| 25 |  | MyoG(HLH)/C2C12-MyoG-ChIP-Seq(GSE36024)/Homer | 1e-2 | -5.837e+00 | 0.0267 | 1113.0 | 92.83% | 43767.6 | 90.53% | motif file (matrix) |
pdf |
| 26 |  | X-box(HTH)/NPC-H3K4me1-ChIP-Seq/Homer | 1e-2 | -5.727e+00 | 0.0287 | 268.0 | 22.35% | 9265.5 | 19.17% | motif file (matrix) |
pdf |
| 27 |  | FOXA1(Forkhead)/LNCAP-FOXA1-ChIP-Seq(GSE27824)/Homer | 1e-2 | -5.543e+00 | 0.0332 | 1125.0 | 93.83% | 44354.6 | 91.75% | motif file (matrix) |
pdf |
| 28 |  | BORIS(Zf)/K562-CTCFL-ChIP-Seq/Homer | 1e-2 | -5.511e+00 | 0.0332 | 492.0 | 41.03% | 18020.9 | 37.28% | motif file (matrix) |
pdf |
| 29 |  | YY1(Zf)/Promoter/Homer | 1e-2 | -5.438e+00 | 0.0343 | 166.0 | 13.84% | 5482.1 | 11.34% | motif file (matrix) |
pdf |
| 30 |  | MYB(HTH)/ERMYB-Myb-ChIPSeq(GSE22095)/Homer | 1e-2 | -5.388e+00 | 0.0349 | 1189.0 | 99.17% | 47479.6 | 98.21% | motif file (matrix) |
pdf |
| 31 |  | FOXA1(Forkhead)/MCF7-FOXA1-ChIP-Seq/Homer | 1e-2 | -5.328e+00 | 0.0358 | 1090.0 | 90.91% | 42810.1 | 88.55% | motif file (matrix) |
pdf |
| 32 |  | Nr5a2(NR)/mES-Nr5a2-ChIP-Seq/Homer | 1e-2 | -5.117e+00 | 0.0429 | 906.0 | 75.56% | 34957.7 | 72.31% | motif file (matrix) |
pdf |
| 33 |  | Egr2/Thymocytes-Egr2-ChIP-Seq(GSE34254)/Homer | 1e-2 | -5.068e+00 | 0.0437 | 422.0 | 35.20% | 15361.1 | 31.77% | motif file (matrix) |
pdf |
| 34 |  | CArG(MADS)/PUER-Srf-ChIP-Seq/Homer | 1e-2 | -4.988e+00 | 0.0459 | 667.0 | 55.63% | 25155.0 | 52.03% | motif file (matrix) |
pdf |
| 35 |  | Stat3+il23(Stat)/CD4-Stat3-ChIP-Seq/Homer | 1e-2 | -4.931e+00 | 0.0472 | 1022.0 | 85.24% | 39914.6 | 82.56% | motif file (matrix) |
pdf |
| 36 |  | NFY(CCAAT)/Promoter/Homer | 1e-2 | -4.859e+00 | 0.0494 | 1013.0 | 84.49% | 39542.3 | 81.79% | motif file (matrix) |
pdf |
| 37 |  | E2F7(E2F)/Hela-E2F7-ChIP-Seq(GSE32673)/Homer | 1e-2 | -4.801e+00 | 0.0509 | 278.0 | 23.19% | 9823.2 | 20.32% | motif file (matrix) |
pdf |
| 38 |  | EBF(EBF)/proBcell-EBF-ChIP-Seq/Homer | 1e-2 | -4.665e+00 | 0.0567 | 570.0 | 47.54% | 21331.9 | 44.12% | motif file (matrix) |
pdf |
| 39 |  | Pbx3(Homeobox)/GM12878-PBX3-ChIP-Seq/Homer | 1e-2 | -4.636e+00 | 0.0569 | 532.0 | 44.37% | 19820.5 | 41.00% | motif file (matrix) |
pdf |













































































